-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praktické
Top novinky
ReklamaThe Regulation of Trypanosome Gene Expression by RNA-Binding Proteins
article has not abstract
Published in the journal: . PLoS Pathog 9(11): e32767. doi:10.1371/journal.ppat.1003680
Category: Pearls
doi: https://doi.org/10.1371/journal.ppat.1003680Summary
article has not abstract
Trypanosomes Depend on Post-transcriptional Mechanisms to Regulate Gene Expression
The genome organization of trypanosomes, leishmanias, and related kinetoplastid organisms is highly unusual: Each RNA polymerase II promoter precedes multiple open reading frames, and the primary transcript is cut into individual mRNAs by 5′ trans splicing and 3′ polyadenylation [1]. Dedicated control of transcription at the level of individual mRNAs is therefore not possible. Nevertheless, mRNA levels are highly controlled: Although some gene sequences are represented by several hundred mRNAs, most mRNAs are present at just one or two copies per cell [2]. Constitutively high mRNA levels can be obtained by having many identical genes, but for most genes, more flexible control points are used: mRNA processing, export from the nucleus, translation, and mRNA decay.
Kinetoplastid mRNAs are probably bound by multiple proteins possessing varying degrees of sequence specificity. The poly(A) tail is masked by poly(A) binding proteins (PABP1 and 2). During translation the 5′ cap is bound by initiation factors [3], [4]; it is expected that, as in other eukaryotes, these interact with PABP, increasing translation efficiency and shielding the mRNA from degradation. Multiple other proteins could bind both translated and untranslated regions, with potential competition between proteins for particular binding sites. Interactions between RNA-binding proteins and proteins of the processing, translation, and degradation machineries will combine to determine RNA fate.
This review focuses on Trypanosoma brucei. T. brucei has two life-cycle stages that grow in culture in the laboratory: the bloodstream form (similar to that in mammalian blood) and the procyclic form (similar to that in the midgut of Tsetse flies). Differentiation of bloodstream forms to procyclics occurs via an intermediate, non-dividing stage, the stumpy form. In Tsetse, the procyclic form in the midgut differentiates to epimastigotes, then to mammalian-infective metacyclic forms in the salivary glands. Methods used for the investigation of RNA-binding proteins are listed in Table 1.
Tab. 1. Methods that can be used to study the functions of mRNA-binding proteins. 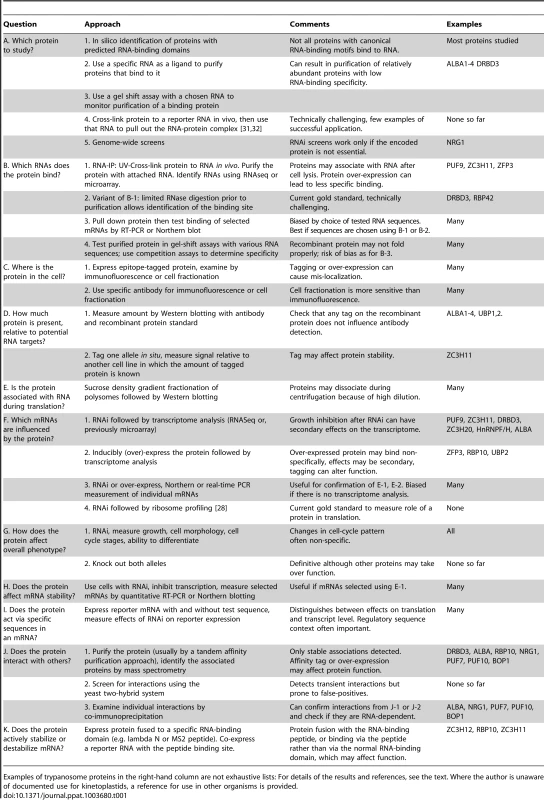
Examples of trypanosome proteins in the right-hand column are not exhaustive lists: For details of the results and references, see the text. Where the author is unaware of documented use for kinetoplastids, a reference for use in other organisms is provided. The Kinetics and Enzymes of mRNA Decay
Most bloodstream-form T. brucei mRNAs have half-lives of five to 20 minutes, but some higher abundance mRNAs have half-lives exceeding two hours; results for procyclics are quite similar ([2] and our unpublished results). The first step in degradation of most mRNAs is removal of the poly(A) tail by the CAF1/NOT complex [2], [5]. This will result in loss of PABP, exposing the mRNA to further degradation—removal of the cap, then degradation by the 5′-exoribonuclease XRNA [2] and/or from the 3′-end by the exosome complex. Depletion of XRNA by RNAi inhibited degradation of some unstable mRNAs; in contrast, a decrease in exosome abundance caused slight delays in degradation of some mRNAs with intermediate half-lives (20 to 60 minutes) [2], [5]. Although most mRNAs show simple exponential decay, others show biphasic kinetics, suggesting involvement of two different rate-limiting processes (our unpublished results).
An RNA-binding protein that interacts with XRNA could enhance 5′-3′ degradation, while one that recruits the CAF1/NOT complex should accelerate deadenylation. Other proteins could stabilize RNAs or enhance translation. They could do this via interactions with translation factors, or simply by preventing the binding of a protein that promotes degradation. This review focuses on selected RNA-binding proteins that influence mRNA levels or translation.
Alba and Pumilio Domain Proteins
Four small proteins with an Alba domain, ALBA1, 2, 3, and 4, form homo - and heterodimers. ALBA2 and ALBA3 are associated with polysomes, and their depletion affects translation of a reporter in a sequence-specific fashion; ALBA3 also interacts with a translation initiation factor. Each ALBA protein is present at 10,000–20,000 molecules per procyclic trypanosome [6]. For comparison: Each procyclic trypanosome has about 40,000 mRNAs, with half as many in bloodstream forms [7]. The lower the protein-to-mRNA ratio, the higher the probability of sequence-specific interactions and function.
Of the ten T. brucei pumilio-domain proteins, PUF9 is the only one with a clearly defined role in mRNA degradation: It binds to, and is required for stability of, a small number of mRNAs that increase in the late G1 phase of the cell cycle [8]. A putative recognition motif, UUGUAC, was identified and shown to be required for PUF9-mediated regulation.
PUF7 and PUF10 are in the nucleolus and required for rRNA maturation, as are two other nucleolar proteins, BOP1 and NRG1 [9]. These four proteins show several mutual interactions. Intriguingly, depletion of any of them results in an increase in the level of the mRNA that encodes the GPEET procyclin surface protein at times when it is normally suppressed. This is especially interesting because the GPEET genes are transcribed by RNA polymerase I, in the nucleolus. NRG1 is also associated, either directly or indirectly, with the GPEET mRNA [9].
RRM Domain Proteins
T. brucei has about 70 different proteins with at least one RRM (RNA Recognition Motif) [10]; of these, about half are essential for normal growth in at least one life-cycle stage [11]. The two poly(A) binding proteins, PABP1 and PABP2, are both associated with polysomes, but may nevertheless have different functions [12]. UBP1 and UBP2, which are similar proteins with a single RRM, are present in a 100-fold excess over mRNA [13], so are unlikely to have sequence-specific functions in vivo, although results from Trypanosoma cruzi indicate that they do show preference for some sequences over others [14]. UBP1 and UBP2 are essential, but a microarray analysis detected little effect on the transcriptome when UBP1 and UBP2 were targeted by RNAi. Over-expression caused transcriptome changes [13], but all results from over-expression must be viewed with extreme caution due to the likelihood of artifacts (Table 1, E2).
RBP42 has a single RRM and is partially associated with polysomes [15]. Its RNA target sites have been mapped accurately (Table 1, B2) [15]. Two thousand different mRNAs showed some binding; interestingly, the 188 that showed most enrichment in the RBP42-associated fraction included many that are involved in energy metabolism. RBP42 bound preferentially to coding regions rather than to untranslated regions [15]; how this is consistent with translation is not yet known. RBP42 is essential in all stages tested, but the effect of depletion on the transcriptome has not yet been described.
DRBD3 (also called PTB1), which has two RRMs, has roles in both splicing and mRNA stability. RNA interference caused moderate (1.5–3×) decreases in 21 different mRNAs, with enrichment for membrane proteins [16]. For a few targets, the decrease in mRNA was shown to be due to destabilization [16], [17], mediated via the 3′ untranslated region of the mRNA (3′-UTR) [16]; the latter observation correlates with the preference of DRBD3 for binding to 3′-UTR sequences [15]. Oddly for a stabilizing protein, DRBD3 was not detected in polysomes, although dissociation during gradient centrifugation has not been ruled out. Affinity purification revealed that DRBD3 interacts with two splicing factors, which correlates with its second likely function in splicing: DRBD3 is in both nucleus and cytoplasm, and RNAi targeting DRBD3 caused clear increases in the levels of some mRNA precursors [17].
Two RRM-domain proteins have been implicated in life-cycle-stage-specific gene expression control. The RBP6 mRNA is most abundant in epimastigotes. Remarkably, ectopic expression of RBP6 in procyclic forms promotes differentiation not only to epimastigotes, but also to the mammalian-infective metacyclic form [18]. RBP6 target mRNAs are not yet known. Meanwhile, the presence of RBP10 correlates with a bloodstream-form expression pattern, especially for genes of energy metabolism [19]. RBP10 RNAi in bloodstream forms decreases their expression, while artificial expression in procyclics increases the mRNA levels. The obvious interpretation is that RBP10 stabilizes energy metabolism mRNAs, but three other observations contradict this idea. Even using a sensitive assay, no binding of mRNA by RBP10 was detected, and it was not associated with polysomes. Moreover, artificial attachment of RBP10 to mRNAs inhibited their translation [19].
Finally, another RRM domain protein, HnRNPF/H, is able to influence both mRNA stability and splicing, with binding sites in 5′ - or 3′-UTRs [20]. This provides a potential link between nuclear processing events and cytoplasmic mRNA regulation.
Zinc-Finger Domain Proteins
There are about 40 T. brucei proteins with C-X8-C-X5-C-X3-H zinc finger domains. ZFP1 is procyclic-specific, while ZFP2 is expressed in bloodstream forms as well; both are required for differentiation of the bloodstream form to the procyclic form [21], [22]. A global analysis of mRNAs associated with ZFP3 in procyclics [23] revealed a bias towards mRNAs that are increased in the stumpy form. ZFP3 also binds specifically to some mRNAs encoding procyclic surface coat proteins, with over-expression promoting translation [24] and RNAi decreasing target mRNA levels.
ZC3H20 is required for growth of procyclic forms [25]. After depletion by RNAi, 12 RNAs were decreased; the two tested were, as expected, destabilized and the effect could be assigned to their 3′-UTRs. Binding of ZC3H20 to two target mRNAs was also demonstrated.
ZC3H11 is essential in bloodstream forms. It is not required for procyclic growth at 27°C, but is required at 37°C, and for the parasites to survive a transient 41°C heat shock [26]. Correspondingly, ZC3H11 binds to, and stabilizes, mRNAs that encode the complete set of chaperones needed for protein refolding after heat denaturation. The N-terminal zinc finger binds specifically to (UAU) repeats in the mRNA 3′-UTRs, while the C-terminal domain has the stabilizing activity [26]. At the normal growth temperature in both forms, ZC3H11 is almost undetectable, but the amount of protein increases dramatically during heat shock. ZC3H11 is the only RNA-binding protein for which a mechanism is known. It interacts with two other proteins, MKT1 and PBP1; PBP1 in turn interacts with PABP (unpublished results). Recruitment of PABP to the 3′-UTR is already known to greatly increase mRNA stability [27]. Thus ZC3H11 could act as a platform to recruit PABP to the 3′-UTR.
Conclusions and Perspectives
Our knowledge of the functions of mRNA-binding proteins in T. brucei is still very fragmented and almost entirely restricted to those with predicted RNA-binding domains. Only a minority of such proteins has been assessed at a transcriptome-wide level for roles in determining mRNA abundance, and global analysis of translation [28] has not yet been applied. Moreover, by restricting investigation to proteins with known RNA binding domains, many important regulators are likely to be missed, since surveys of mRNA-bound proteins in other organisms have revealed that many lack such domains [29], [30].
By comparison with yeast and mammalian cells [29], [30], we can expect 100–200 different proteins to be associated with trypanosome mRNA. Different sets of proteins, with varying degrees of specificity, will be associated with different sequences, competing and cooperating with each other and changing over the mRNA lifetime. In the future, it will be essential to move from studies of individual proteins towards understanding the complex networks of interactions that determine the fates of individual mRNAs.
Zdroje
1. MichaeliS (2011) Trans-splicing in trypanosomes: machinery and its impact on the parasite transcriptome. Future Microbiol 6 : 459–474.
2. ManfulT, FaddaA, ClaytonC (2011) The role of the 5′-3′ exoribonuclease XRNA in transcriptome-wide mRNA degradation. RNA 17 : 2039–2047.
3. FreireE, DhaliaR, MouraaD, da Costa LimaT, LimaR, et al. (2011) The four trypanosomatid eIF4E homologues fall into two separate groups, with distinct features in primary sequence and biological properties. Mol Biochem Parasitol 176 : 25–36.
4. YoffeY, LegerM, ZinovievA, ZuberekJ, DarzynkiewiczE, et al. (2009) Evolutionary changes in the Leishmania eIF4F complex involve variations in the eIF4E-eIF4G interactions. Nucleic Acids Res 37 : 3243–3253.
5. FaddaA, FärberV, DrollD, ClaytonC (2013) The roles of 3′-exoribonucleases and the exosome in trypanosome mRNA degradation. RNA 19 : 937–947.
6. ManiJ, GüttingerA, SchimanskiB, HellerM, Acosta-SerranoA, et al. (2011) Alba-domain proteins of Trypanosoma brucei are cytoplasmic RNA-binding proteins that interact with the translation machinery. PLoS ONE 6: e22463 doi: 10.1371/journal.pone.0022463
7. HaanstraJ, StewartM, LuuV-D, van TuijlA, WesterhoffH, et al. (2008) Control and regulation of gene expression: quantitative analysis of the expression of phosphoglycerate kinase in bloodstream form Trypanosoma brucei. J Biol Chem 283 : 2495–2507.
8. ArcherSK, van LuuD, de QueirozR, BremsS, ClaytonCE (2009) Trypanosoma brucei PUF9 regulates mRNAs for proteins involved in replicative processes over the cell cycle. PLoS Pathog 5: e1000565 doi: 10.1371/journal.ppat.1000565
9. Schumann BurkardG, KaserS, de AraujoPR, SchimanskiB, NaguleswaranA, et al. (2013) Nucleolar proteins regulate stage-specific gene expression and ribosomal RNA maturation in Trypanosoma brucei. Mol Microbiol 88 : 827–840.
10. WurstM, RoblesA, PoJ, LuuV, BremsS, et al. (2009) An RNAi screen of the RRM-domain proteins of Trypanosoma brucei. Mol Biochem Parasitol 163 : 61–65.
11. AlsfordS, TurnerD, ObadoS, Sanchez-FloresA, GloverL, et al. (2011) High throughput phenotyping using parallel sequencing of RNA interference targets in the African trypanosome. Genome Res 21 : 915–924.
12. KramerS, Bannerman-ChukualimB, EllisL, BouldenE, KellyS, et al. (2013) Differential localization of the two T. brucei poly(A) binding proteins to the nucleus and RNP granules suggests binding to distinct mRNA pools. PLoS ONE 8: e54004 doi: 10.1371/journal.pone.0054004
13. HartmannC, BenzC, BremsS, EllisL, LuuV-D, et al. (2007) The small trypanosome RNA-binding proteins TbUBP1 and TbUBP2 influence expression of F box protein mRNAs in bloodstream trypanosomes. Eukaryotic Cell 6 : 1964–1978.
14. NoeG, De GaudenziJ, FraschA (2008) Functionally related transcripts have common RNA motifs for specific RNA-binding proteins in trypanosomes. BMC Mol Biol 9 : 107.
15. DasA, MoralesR, BandayM, GarciaS, HaoL, et al. (2012) The essential polysome-associated RNA-binding protein RBP42 targets mRNAs involved in Trypanosoma brucei energy metabolism. RNA 18 : 1968–1983.
16. EstévezA (2008) The RNA-binding protein TbDRBD3 regulates the stability of a specific subset of mRNAs in trypanosomes. Nucleic Acids Res 36 : 4573–4586.
17. SternM, GuptaS, Salmon-DivonM, HahamT, BardaO, et al. (2009) Multiple roles for polypyrimidine tract binding (PTB) proteins in trypanosome RNA metabolism. RNA 15 : 648–665.
18. KolevNG, Ramey-ButlerK, CrossGA, UlluE, TschudiC (2012) Developmental progression to infectivity in Trypanosoma brucei triggered by an RNA-binding protein. Science 338 : 1352–1353.
19. WurstM, SelingerB, JhaB, KleinC, QueirozR, et al. (2012) Expression of the RNA Recognition Motif protein RBP10 promotes a bloodstream-form transcript pattern in Trypanosoma brucei. Mol Microbiol 83 : 1048–1063.
20. GuptaSK, KostiI, PlautG, PivkoA, TkaczID, et al. (2013) The hnRNP F/H homologue of Trypanosoma brucei is differentially expressed in the two life cycle stages of the parasite and regulates splicing and mRNA stability. Nucleic Acids Res
21. HendriksEF, RobinsonDR, HinkinsM, MatthewsKR (2001) A novel CCCH protein which modulates differentiation of Trypanosoma brucei to its procyclic form. EMBO J 20 : 6700–6711.
22. HendriksEF, MatthewsKR (2005) Disruption of the developmental programme of Trypanosoma brucei by genetic ablation of TbZFP1, a differentiation-enriched CCCH protein. Mol Microbiol 57 : 706–716.
23. WalradP, CapewellP, FennK, MatthewsK (2011) The post-transcriptional trans-acting regulator, TbZFP3, co-ordinates transmission-stage enriched mRNAs in Trypanosoma brucei. Nucleic Acids Res 40 : 2869–2883.
24. WalradP, PaterouA, Acosta-SerranoA, MatthewsK (2009) Differential trypanosome surface coat regulation by a CCCH protein that co-associates with procyclin mRNA cis-elements. PLoS Pathog 5: e1000317 doi: 10.1371/journal.ppat.1000317
25. LingA, TrotterJ, HendriksE (2011) A zinc finger protein, TbZC3H20, stabilises two developmentally regulated mRNAs in trypanosomes. J Biol Chem 286 : 20152–20162.
26. DrollD, MiniaI, FaddaA, SinghA, StewartM, et al. (2013) Post-transcriptional regulation of the trypanosome heat shock response by a zinc finger protein. PLoS Pathog 9: e1003286 doi: 10.1371/journal.ppat.1003286
27. DelhiP, QueirozR, InchausteguiD, CarringtonM, ClaytonC (2011) Is there a classical nonsense-mediated decay pathway in trypanosomes? PLoSOne 6: e25112 doi: 10.1371/journal.pone.0025112
28. IngoliaN, GhaemmaghamiS, NewmanJ, WeissmanJ (2009) Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324 : 218–223.
29. MitchellS, JainS, SheM, ParkerR (2013) Global analysis of yeast mRNPs. Nat Struct Mol Biol 20 : 127–135.
30. BaltzAG, MunschauerM, SchwanhausserB, VasileA, MurakawaY, et al. (2012) The mRNA-bound proteome and its global occupancy profile on protein-coding transcripts. Mol Cell 46 : 674–690.
31. IiokaH, LoiselleD, HaysteadTA, MacaraIG (2011) Efficient detection of RNA-protein interactions using tethered RNAs. Nucleic Acids Res 39: e53.
32. SlobodinB, GerstJ (2010) A novel mRNA affinity purification technique for the identification of interacting proteins and transcripts in ribonucleoprotein complexes. RNA 16 : 2277–2290.
Štítky
Hygiena a epidemiologie Infekční lékařství Laboratoř
Článek vyšel v časopisePLOS Pathogens
Nejčtenější tento týden
2013 Číslo 11- Farmakovigilanční studie perorálních antivirotik indikovaných v léčbě COVID-19
- Jak souvisí postcovidový syndrom s poškozením mozku?
- Měli bychom postcovidový syndrom léčit antidepresivy?
- 10 bodů k očkování proti COVID-19: stanovisko České společnosti alergologie a klinické imunologie ČLS JEP
-
Všechny články tohoto čísla
- Baculoviruses: Sophisticated Pathogens of Insects
- How Do Viruses Avoid Inhibition by Endogenous Cellular MicroRNAs?
- The Regulation of Trypanosome Gene Expression by RNA-Binding Proteins
- DNA Damage Repair and Bacterial Pathogens
- Disease to Dirt: The Biology of Microbial Amyloids
- Fungal Immune Evasion in a Model Host–Pathogen Interaction: Versus Macrophages
- Infectious Prions Accumulate to High Levels in Non Proliferative C2C12 Myotubes
- The Biology and Taxonomy of Head and Body Lice—Implications for Louse-Borne Disease Prevention
- Antibodies Trap Tissue Migrating Helminth Larvae and Prevent Tissue Damage by Driving IL-4Rα-Independent Alternative Differentiation of Macrophages
- The Effects of Somatic Hypermutation on Neutralization and Binding in the PGT121 Family of Broadly Neutralizing HIV Antibodies
- Natural Selection Promotes Antigenic Evolvability
- Type I Interferon Protects against Pneumococcal Invasive Disease by Inhibiting Bacterial Transmigration across the Lung
- Mode of Parainfluenza Virus Transmission Determines the Dynamics of Primary Infection and Protection from Reinfection
- Type I and Type III Interferons Drive Redundant Amplification Loops to Induce a Transcriptional Signature in Influenza-Infected Airway Epithelia
- Unraveling a Three-Step Spatiotemporal Mechanism of Triggering of Receptor-Induced Nipah Virus Fusion and Cell Entry
- A Novel Membrane Sensor Controls the Localization and ArfGEF Activity of Bacterial RalF
- Macrophage and T Cell Produced IL-10 Promotes Viral Chronicity
- Global Rescue of Defects in HIV-1 Envelope Glycoprotein Incorporation: Implications for Matrix Structure
- Turning Defense into Offense: Defensin Mimetics as Novel Antibiotics Targeting Lipid II
- The Neonatal Fc Receptor (FcRn) Enhances Human Immunodeficiency Virus Type 1 (HIV-1) Transcytosis across Epithelial Cells
- Brd4 Is Displaced from HPV Replication Factories as They Expand and Amplify Viral DNA
- A Viral Genome Landscape of RNA Polyadenylation from KSHV Latent to Lytic Infection
- The Cytotoxic Necrotizing Factor of (CNF) Enhances Inflammation and Yop Delivery during Infection by Activation of Rho GTPases
- The Inflammatory Kinase MAP4K4 Promotes Reactivation of Kaposi's Sarcoma Herpesvirus and Enhances the Invasiveness of Infected Endothelial Cells
- Conservative Sex and the Benefits of Transformation in
- Microbial Endocrinology in the Microbiome-Gut-Brain Axis: How Bacterial Production and Utilization of Neurochemicals Influence Behavior
- Colonization Resistance: Battle of the Bugs or Ménage à Trois with the Host?
- Intracellular Interferons in Fish: A Unique Means to Combat Viral Infection
- SPOC1-Mediated Antiviral Host Cell Response Is Antagonized Early in Human Adenovirus Type 5 Infection
- Involvement of the Cellular Phosphatase DUSP1 in Vaccinia Virus Infection
- Killer Bee Molecules: Antimicrobial Peptides as Effector Molecules to Target Sporogonic Stages of
- A Unique SUMO-2-Interacting Motif within LANA Is Essential for KSHV Latency
- A Role for Host Activation-Induced Cytidine Deaminase in Innate Immune Defense against KSHV
- Haploid Genetic Screens Identify an Essential Role for PLP2 in the Downregulation of Novel Plasma Membrane Targets by Viral E3 Ubiquitin Ligases
- A Small Molecule Glycosaminoglycan Mimetic Blocks Invasion of the Mosquito Midgut
- Identification of the Adenovirus E4orf4 Protein Binding Site on the B55α and Cdc55 Regulatory Subunits of PP2A: Implications for PP2A Function, Tumor Cell Killing and Viral Replication
- Can Non-lytic CD8+ T Cells Drive HIV-1 Escape?
- Deletion of the α-(1,3)-Glucan Synthase Genes Induces a Restructuring of the Conidial Cell Wall Responsible for the Avirulence of
- PLOS Pathogens
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Baculoviruses: Sophisticated Pathogens of Insects
- Identification of the Adenovirus E4orf4 Protein Binding Site on the B55α and Cdc55 Regulatory Subunits of PP2A: Implications for PP2A Function, Tumor Cell Killing and Viral Replication
- A Unique SUMO-2-Interacting Motif within LANA Is Essential for KSHV Latency
- Natural Selection Promotes Antigenic Evolvability
Přihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



