-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praktické
Top novinky
ReklamaFunctional Antagonism between Sas3 and Gcn5 Acetyltransferases and ISWI Chromatin Remodelers
Chromatin-modifying enzymes and ATP-dependent remodeling complexes have been intensely studied individually, yet how these activities are coordinated to ensure essential cell functions such as transcription, replication, and repair of damage is not well understood. In this study, we show that the critical loss of Sas3 and Gcn5 acetyltransferases in yeast can be functionally rescued by inactivation of ISWI remodelers. This genetic interaction depends on the ATPase activities of Isw1 and Isw2, suggesting that it involves chromatin remodeling activities driven by the enzymes. Genetic dissection of the Isw1 complexes reveals that the antagonistic effects are mediated specifically by the Isw1a complex. Loss of Sas3 and Gcn5 correlates with defective RNA polymerase II (RNAPII) occupancy at actively transcribed genes, as well as a significant loss of H3K14 acetylation. Inactivation of the Isw1a complex in the acetyltransferase mutants restores RNAPII recruitment at active genes, indicating that transcriptional regulation may be the mechanism underlying suppression. Dosage studies and further genetic dissection reveal that the Isw1b complex may act in suppression through down-regulation of Isw1a. These studies highlight the importance of balanced chromatin modifying and remodeling activities for optimal transcription and cell growth.
Published in the journal: . PLoS Genet 8(10): e32767. doi:10.1371/journal.pgen.1002994
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1002994Summary
Chromatin-modifying enzymes and ATP-dependent remodeling complexes have been intensely studied individually, yet how these activities are coordinated to ensure essential cell functions such as transcription, replication, and repair of damage is not well understood. In this study, we show that the critical loss of Sas3 and Gcn5 acetyltransferases in yeast can be functionally rescued by inactivation of ISWI remodelers. This genetic interaction depends on the ATPase activities of Isw1 and Isw2, suggesting that it involves chromatin remodeling activities driven by the enzymes. Genetic dissection of the Isw1 complexes reveals that the antagonistic effects are mediated specifically by the Isw1a complex. Loss of Sas3 and Gcn5 correlates with defective RNA polymerase II (RNAPII) occupancy at actively transcribed genes, as well as a significant loss of H3K14 acetylation. Inactivation of the Isw1a complex in the acetyltransferase mutants restores RNAPII recruitment at active genes, indicating that transcriptional regulation may be the mechanism underlying suppression. Dosage studies and further genetic dissection reveal that the Isw1b complex may act in suppression through down-regulation of Isw1a. These studies highlight the importance of balanced chromatin modifying and remodeling activities for optimal transcription and cell growth.
Introduction
Two major classes of enzymes regulate the architecture of chromatin and are thereby critical for DNA-templated processes such as transcription, replication, and repair of damage. Remodeling enzymes use the energy of ATP hydrolysis to alter the structure or position of nucleosomes (reviewed in [1], [2]), whereas chromatin modifying enzymes act post-translationally on multiple nuclear substrates. Prominent among these are the nucleosomal histones that are extensively modified on their N - and C-terminal tails (reviewed in [3]). Covalent modifications of histones and other chromatin proteins are diverse, including at least six specific types of reversible and dynamic modifications that are catalyzed by multimeric enzyme complexes. Among the consequences resulting from histone modifications, two have been especially well characterized. The first is disruption of contacts between DNA and histones or between nucleosomes. In this case, lysine ε-acetylation can destabilize nucleosomal interactions since it neutralizes this otherwise charged residue. The second consequence involves recruitment of effector proteins that bind via conserved recognition domains. For example, histone acetylation can be recognized by bromodomains, whereas histone methylation is recognized by chromo-like-domains and PHD domains (reviewed in [4]). These domains are found in many nuclear proteins, including chromatin modifying enzymes and remodeling complexes.
The simultaneous existence of multiple different marks on histones has led to the recognition that crosstalk among modifications can be a critical determinant for regulation of gene expression [5]. In addition, cooperation between histone modifiers and ATP-dependent remodeling complexes can contribute to transcriptional regulation (reviewed in [6]–[8]) and DNA damage repair (reviewed in [9]). For example, the histone acetyltransferase (HAT) Gcn5 and the SWI/SNF chromatin remodeling complex were proposed early on to have cooperative functions in transcriptional activation by working in concert to modify chromatin structure [10]–[12]. Indeed, histone acetylation mediated by the SAGA complex containing Gcn5 stabilizes the anchoring of SWI/SNF to nucleosomes at promoters, and thus is important for SWI/SNF-dependent nucleosome remodeling and transcriptional activation in vitro and in vivo [13], [14]. The SAGA complex interacts with another chromatin remodeling factor, the chromodomain protein Chd1 [15], [16], which is a component of Gcn5-containing SAGA and SLIK/SALSA HAT complexes [17]. In addition, Gcn5 is functionally linked to the essential RSC chromatin remodeling complex: H3K14 acetylation is recognized by one essential tandem bromodomain of Rsc4 and contributes to RSC complex-dependent gene activation [18], [19].
Acetylation of histone H3 at lysines 9 and 14 strongly correlates with transcriptional activity and peaks over start sites of active genes [20]. Gcn5 is the HAT responsible for the majority of this acetylation in vivo [21]–[23], consistent with the observation that Gcn5 is generally recruited to promoters of active genes, as described for H3K9 and K14 acetylation marks [20], [24]. Two other HATs also specifically target histone H3 at K9 and K14 in vivo: the MYST family HAT Sas3 (reviewed in [25]), and Elp3 [21], [23]. Their functions appear most critical in the absence of Gcn5. In particular, although inactivation of Sas3 in otherwise wild-type cells does not elicit obvious phenotypes, diminished Sas3 activity in a gcn5Δ null mutant results in defects in cell cycle progression, and complete loss of activity leads to cell death [22]. Genome-wide mapping established that Sas3 and Gcn5 are recruited to many of the same actively transcribed genes [26]. Furthermore, the binding sites of these HATs correlates with the H3K14 acetylation mark [26]. These observations strongly suggest that Sas3 and Gcn5 acetyltransferases are critical for active transcription, although the molecular mechanisms underlying their regulation have not been fully elucidated. In particular, since mutations of the established target lysines in histone H3 result in only mild phenotypes [27], [28], the essential function revealed in gcn5Δ sas3 mutants most likely extends beyond acetylation of H3.
Functional links have not yet been established between Sas3 and chromatin remodeling, or the loss of viability that results when both Sas3 and Gcn5 activities are compromised. We describe here a critical interaction between Sas3 and Gcn5 acetyltransferases and ISWI chromatin remodelers. Strikingly, and in contrast with nucleosome disrupting remodelers such as SWI/SNF and Chd1, inactivation of Isw1 or Isw2 relieved conditional lethality in a gcn5Δ sas3 mutant. Genetic dissection of the complexes through which Isw1 acts clearly reveals that the antagonistic effects are mediated through the Isw1a complex, and furthermore, that elimination of non-catalytic subunits of Isw1b can overcome this antagonism. At a molecular level, the effects on cell viability tightly correlate with the recruitment of RNA polymerase II (RNAPII) at active genes. Together these studies provide new evidence for functional distinctions between the families of chromatin remodeling activities and point to critical interactions between the Sas3 and Gcn5 acetyltransferases, ISWI remodeling machines, RNAPII recruitment, and chromatin compaction.
Results
Functional antagonism between Sas3 and Gcn5 acetyltransferases and ISWI chromatin remodeling enzymes
Among the major H3 acetyltransferases, either Gcn5 or Sas3 is required for cell viability: loss of both enzymes leads to death. To understand this loss of viability, we asked if other chromatin-modulating activities contributed to it, and in particular if there was a role for ATP-dependent chromatin remodeling activities.
There are four distinct families of biochemically defined chromatin remodeling complexes: SWI-SNF, ISWI, CHD, and INO80 [1]. The RSC and INO80 catalytic ATPases Sth1 and Ino80 are essential, although the ino80Δ lethality appears restricted to the W303 genetic background [29], [30]. Individual inactivation of the other catalytic ATPases, Snf2, Isw1, Isw2 or Chd1 does not trigger marked growth defects in otherwise wild-type cells [15], [31], [32]. However, earlier studies reported synthetic lethality between SWI-SNF components and members of the Gcn5-SAGA complex [11].
We began by evaluating the effects of ATP-dependent chromatin remodeling activities when Sas3 and Gcn5 activities were compromised, using the temperature-sensitive gcn5Δ sas3 conditional mutant described earlier [22]. We observed two types of functional interactions between chromatin remodelers and the Sas3 and Gcn5 acetyltransferases.
First, the temperature-sensitive phenotype of the gcn5Δ sas3 double mutant was exacerbated upon deletion of CHD1 (Figure 1A and Figure S1A). This suggested parallel functions, likely through overlap in transcriptional regulation, in agreement with previous studies [12], [17]. Second, and in distinct contrast, deletion of ISW1 and to a lesser extent ISW2, improved growth of the gcn5Δ sas3 cells (Figure 1A). Deletion of ISW2 does not further restore growth of the gcn5Δ sas3 isw1Δ mutant, indicating that rescue is maximal upon inactivation of Isw1 (Figure S1B).
Fig. 1. Sas3 and Gcn5 acetyltransferases and ISWI chromatin remodeling enzymes have antagonistic functions. 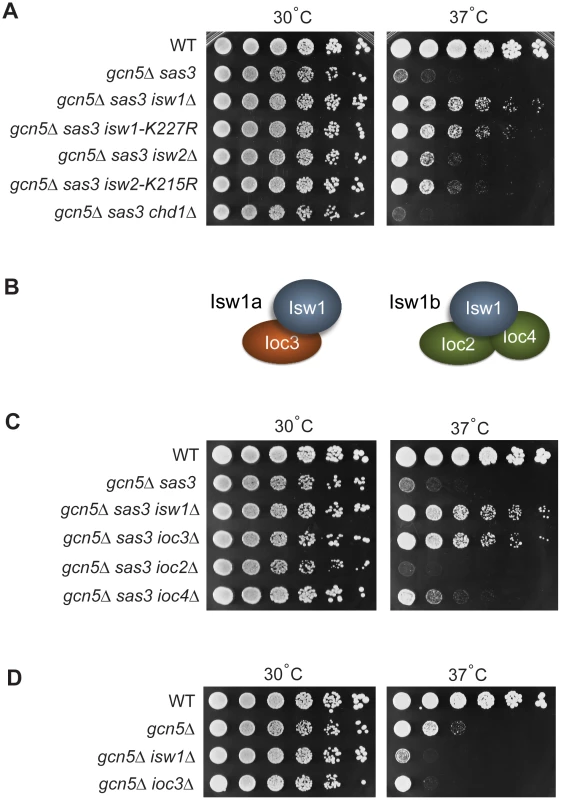
(A) Inactivation of the chromatin remodeling ATPases Isw1 and Isw2 suppresses the growth defects of the gcn5Δ sas3 mutant. Fivefold serial dilutions of cells were plated onto SC medium supplemented with 1 M sorbitol and grown for 4 days at the indicated temperatures (see also Figure S1A). (B) Isw1 is the catalytic subunit of two structurally distinct complexes. (C) Inactivation of the Isw1a complex (Isw1/Ioc3) suppresses the gcn5Δ sas3 mutant. Strains were grown and plated as in panel A. (D) Inactivation of the Isw1a complex (Isw1/Ioc3) does not rescue the temperature sensitivity of the gcn5Δ mutant. Strains were plated on SC and grown for 4 days. The Isw1 and Isw2 chromatin remodeling complexes use the energy of ATP hydrolysis to alter nucleosome positioning [32]–[35]. To determine if the ATP-dependent catalytic activity, and not some other property of the enzymes was responsible for suppression, we analyzed the isw1-K227R and isw2-K215R mutants that affect ATP-binding sites to inactivate the enzymes [32]. Figure 1A shows that both catalytic mutations rescued the temperature sensitivity of the gcn5Δ sas3 mutant, with isw1-K227R having the stronger effect. Thus, suppression is dependent on the catalytic activities of the ISWI ATPases.
The suppression observed was unexpected since most reported interactions between chromatin modifying enzymes and chromatin remodelers describe parallel functions, often through recognition of modified nucleosomes by the remodelers [12], [17], [18], [36]–[38]. Because inactivation of the ATPase function of Isw1 consistently resulted in a stronger rescue phenotype than that of Isw2, we focused on dissecting the mechanism underlying the potential antagonism between ISW1 remodeling and acetyltransferase activities mediated by SAS3 and GCN5.
The Sas3 and Gcn5 acetyltransferases counteract Isw1 function through the Isw1a complex
The Isw1 ATPase is the catalytic subunit of two distinct complexes (Figure 1B and reviewed in [1], [39]). The Isw1a complex includes the non-catalytic subunit Ioc3, whereas the non-catalytic subunits of Isw1b are Ioc2 and Ioc4 [32], [35]. The in vivo functions of these two complexes are not yet fully established. Microarray studies reveal that the Isw1a and Isw1b complexes have overlapping roles in transcriptional regulation at some genes, but distinct functions at others [35]. To determine whether one or both Isw1 complexes are involved in antagonizing Sas3 and Gcn5 function, we inactivated the complexes individually by deleting genes encoding their non-catalytic subunits. The ioc3Δ mutant, but neither ioc2Δ nor ioc4Δ strains, strongly rescued the gcn5Δ sas3 phenotype, demonstrating that inactivation of the Isw1a complex is primarily responsible for the suppression mediated by loss of Isw1 (Figure 1C). Furthermore, deletion of IOC2 exacerbated loss of viability, whereas ioc4Δ had no significant reproducible effects. This suggests an additional relationship between the Isw1b complex and the acetyltransferases Sas3 and Gcn5, and further supports the existence of distinct functions for the two Isw1 complexes. Since combined inactivation of both complexes through deletion of ISW1 rescued the temperature sensitivity, Isw1a appears to have a prominent role in antagonizing Sas3 and Gcn5 activities.
Previous studies demonstrated interactions between the SWI-SNF remodelers and Gcn5 alone, independent of Sas3 [11], [12]. To determine if the antagonism between Sas3 and Gcn5 and the Isw1a complex is Gcn5-specific or acts through shared Sas3 and Gcn5 functions, we asked if inactivation of the Isw1a complex rescued temperature sensitivity associated with the single gcn5Δ mutation. Deleting IOC3 or ISW1 did not rescue, demonstrating that these acetyltransferases counteract Isw1a through shared functions of both acetyltransferases (Figure 1D).
Loss of Sas3 and Gcn5 acetyltransferases modestly alters Isw1a recruitment at some active genes
Isw1 contains a SANT domain (reviewed in [40]) that is critical for binding to chromatin at regulated genes in vivo [41]. Biochemical studies indicate that the SANT domains from Ada2 and SMRT preferentially bind unacetylated histone H3 tails [42], [43]. To determine whether H3 acetylation antagonizes Isw1 recruitment to chromatin, we evaluated Ioc3-Myc occupancy in gcn5Δ sas3 cells by chromatin immunoprecipitation (ChIP) at transcriptionally active target genes (Figure 2). Indeed, Sas3 and Gcn5 acetyltransferases are recruited to a similar set of actively transcribed genes, which correlate with H3K14 acetylation [26]. We selected the PYK1 gene for analysis since H3 acetyltransferases and Isw1 are enriched at this locus [24], [26], [44]. For other candidate genes, a recent genome-wide study revealed that Sas3 is enriched at RPL10 whereas UBP7 and CDC25 are impaired for H3K14 acetylation in a sas3Δ strain [26].
Fig. 2. Loss of the Sas3 and Gcn5 H3 acetyltransferases modestly alters Isw1a recruitment to chromatin. 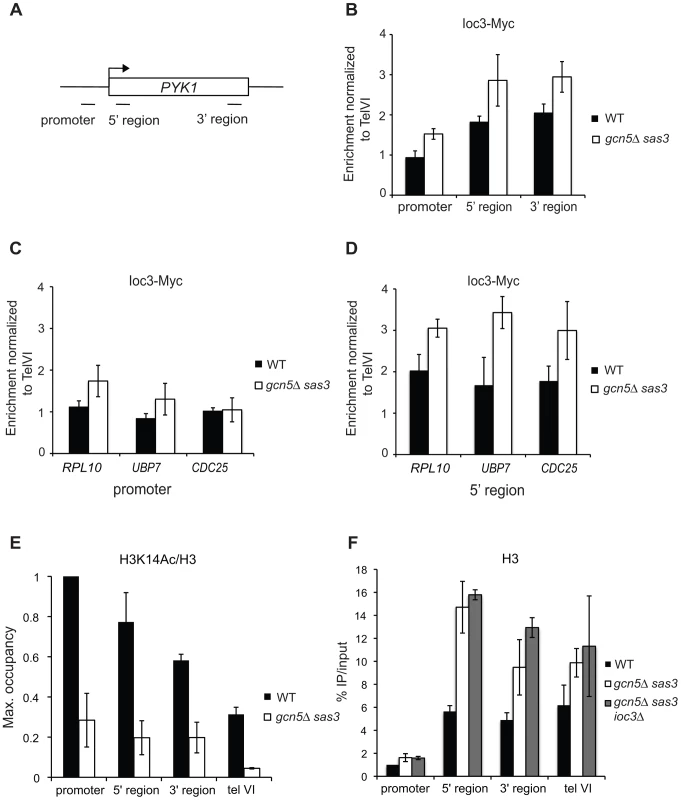
(A) Schematic representation of the PYK1 locus and positions of primer pairs used for ChIP analysis in (B), (E) and (F). Ioc3-Myc occupancy was assayed by ChIP in wild-type and gcn5Δ sas3 cells grown in SC medium at 34°C over the PYK1 gene (B), at promoter (C) and 5′ coding regions (D) of RPL10, CDC25 and UBP7 genes. (E) Histone H3K14 acetylation occupancy was analyzed by ChIP over the PYK1 gene in wild-type and gcn5Δ sas3 cells grown in SC medium at 34°C. (F) Histone H3 occupancy over the PYK1 gene was assayed by ChIP in wild-type, gcn5Δ sas3, and gcn5Δ sas3 ioc3Δ cells grown in SC medium at 34°C. Ioc3-Myc occupancy was normalized to the telVI control region. H3K14Ac and H3 levels were normalized to the promoter region of PYK1. The values represent the means from two or more independent experiments, with error bars reflecting standard deviations. We observed that Ioc3 occupancy is enriched in the coding region compared to the promoter at the PYK1, RPL10, UBP7 and CDC25 genes (Figure 2B, Figure 2C and Figure 2E), as previously described for Isw1 at regulated genes [41]. The inactivation of Sas3 and Gcn5 modestly increased the recruitment of Ioc3 at promoter and coding regions of PYK1 (Figure 2B), although the levels of H3K14 acetylation were severely decreased at this locus (Figure 2E). Similarly, Ioc3 occupancy was moderately affected by loss of Sas3 and Gcn5 at the promoter of RPL10 and at coding regions of RPL10, UBP7 and CDC25 (Figure 2C and Figure 2D). Of note, nucleosome density increased in the coding region of PYK1 upon inactivation of Sas3 and Gcn5, as revealed by H3 occupancy (Figure 2F).
Inactivation of the Isw1a complex does not rescue defects in H3 acetylation, nor nucleosome occupancy
As the combined loss of Sas3 and Gcn5 resulted in a dramatic loss of H3 acetylation, we asked if suppression mediated by inactivation of the Isw1a complex might rescue this defect. Although the chromatin remodeling ATPase Isw1 has not been reported to regulate histone H3 acetylation, we hypothesized that nucleosome repositioning resulting from Isw1a inactivation might rescue gcn5Δ sas3 defects. Since K14 is the major and common target of Sas3 and Gcn5 acetyltransferases in vitro and in vivo [22], [26], [45], we assayed global levels of H3K14 acetylation in wild-type, gcn5Δ sas3 and gcn5Δ sas3 ioc3Δ cells by protein immunoblotting. As previously described [22], H3K14 acetylation decreased in the gcn5Δ sas3 strain, however there was no significant difference in acetylation in the gcn5Δ sas3 ioc3Δ strain (Figure S3).
Because global restoration of H3K14 acetylation did not occur, we tested the idea that locus-specific changes might be responsible for rescue of the gcn5Δ sas3 mutant by assaying the local levels of H3K14 acetylation at the Isw1-responsive PYK1 gene under suppressing conditions. H3K14 acetylation levels were impaired over the whole PYK1 gene (promoter and coding regions) in the gcn5Δ strain, and more dramatically in the gcn5Δ sas3 strain (Figure 3B), demonstrating that Sas3 and Gcn5 are responsible for H3K14 acetylation at this locus. Further, as revealed by the levels of H3K14 acetylation in the gcn5Δ strain, Sas3 markedly contributed to H3K14 acetylation in the promoter and 3′ regions of the PYK1 gene (Figure 3B). Yet, no further difference in H3K14 acetylation levels was observed upon deletion of IOC3 or ISW1 (Figure 3B). Similarly, the elevated H3 occupancy observed in the gcn5Δ sas3 strain at the PYK1 gene remained unaffected by inactivation of Ioc3 (Figure 2F). Because inactivation of Isw1a does not suppress Sas3 and Gcn5 defects by directly restoring K14 acetylation either globally or locally, or by decreasing nucleosome occupancy, suppression must occur through some other mechanism.
Fig. 3. Sas3 and Gcn5 acetyltransferases and Isw1a antagonistically regulate RNAPII recruitment to active genes. 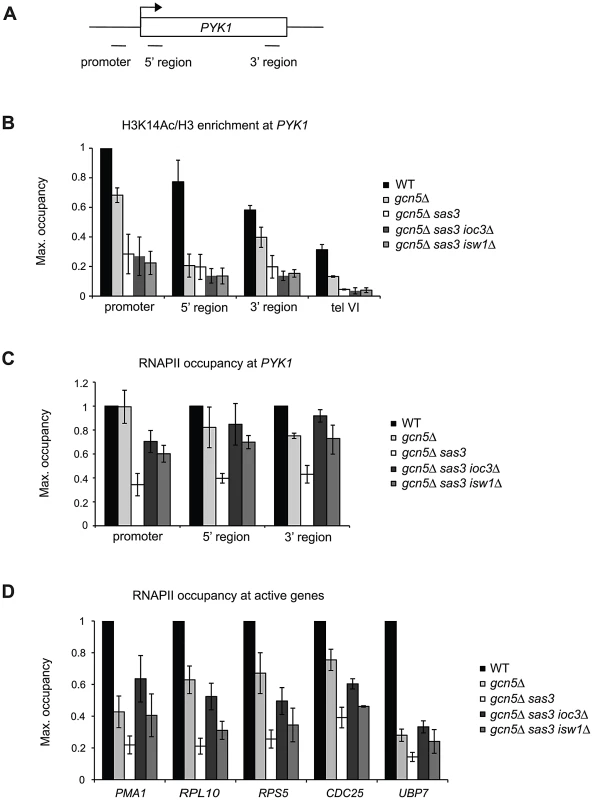
(A) Schematic representation of the PYK1 locus and positions of primer pairs used for ChIP analysis in (B) and (C). (B) Histone H3K14 acetylation (Ac) levels and (C) RNAPII occupancy over the PYK1 gene were assayed by ChIP analysis of cells grown in SC medium at 34°C. H3K14Ac ChIP samples were normalized to total H3. (D) RNAPII occupancy was analyzed by ChIP as in (C), but at the 5′ regions of PMA1, RPL10, RPS5, CDC25 and UBP7 genes. H3K14Ac levels were further normalized to the promoter region of PYK1, and RNAPII occupancies were normalized to WT for each locus, arbitrarily set to 1. The values represent the means from two or more independent experiments, with error bars reflecting standard deviations. Isw1a and the acetyltransferases Sas3 and Gcn5 regulate the recruitment of RNAPII to active genes
We asked if the interaction between Isw1a and H3 HATs is related to transcription as reflected by RNAPII occupancy. Indeed, genome-wide analyses revealed that Sas3 and Gcn5 are recruited to actively transcribed genes and that their occupancies correlate with transcriptional rates [20], [24], [26]. Based on the fact that Isw1 also regulates transcriptional activation [35], [44], we assayed RNAPII occupancy at the PYK1 gene.
In agreement with the proposed roles of Sas3 and Gcn5 in transcriptional activation, we observed that loss of these HATs resulted in defective RNAPII recruitment at PYK1 (Figure 3C and Figure S4A). Loss of Gcn5 slightly impaired occupancy at the 3′ region yet did not affect the promoter and 5′ regions, highlighting a role for Sas3 in RNAPII recruitment and in transcriptional activation. Significantly, deletion of IOC3 and ISW1 in a gcn5Δ sas3 mutant partially rescued RNAPII occupancy at the promoter and coding region of PYK1 (Figure 3C and Figure S4A).
We evaluated RNAPII at the other active genes described above to determine how general the suppressive effects were on occupancy. For this study, we also included the additional Isw1 and Sas3 target genes PMA1 and RPS5, respectively [26], [44]. As shown for PYK1, we found that Sas3 and Gcn5 contributed to the recruitment of RNAPII at PMA1, RPL10, RPS5, CDC25 and UBP7 coding regions (Figure 3D and Figure S4B). Furthermore, deletion of IOC3 improved RNAPII occupancy at these genes, more than the modest effects observed with isw1Δ. We asked if these differences in the recruitment of RNAPII influenced trancription (Figure S4C). We assayed the steady state levels of PYK1, PMA1 and RPL10 mRNAs by RT-qPCR, and observed no significant changes in the gcn5Δ sas3 and gcn5Δ sas3 ioc3Δ mutants when compared to the wild-type strain (Figure S4C). Together these results demonstrate that Sas3 and Gcn5 acetyltransferases and the Isw1a complex have antagonistic roles in chromatin as reflected by recruitment of RNAPII.
Isw1-dependent rescue of RNAPII at PYK1 is independent of nucleosome repositioning
RNAPII function and regulated recruitment during transcription are critically dependent on chromatin architecture. Given that Sas3 and Gcn5 acetyltransferases and the Isw1a complex have opposing effects on RNAPII recruitment to transcriptional target genes, we asked whether they also antagonistically regulate nucleosomal occupancy at the PYK1 locus. It has been suggested that PYK1 chromatin structure is dependent on Isw1 since the ATPase is associated with this coding region [44].
We examined nucleosomal organization at PYK1 by comparing micrococcal nuclease (MNase) cleavage patterns of chromatin prepared from wild-type and mutant strains (Figure S5). We observed no significant differences in MNase cleavage patterns in gcn5Δ sas3 ioc3Δ cells when compared to gcn5Δ sas3 or wild-type cells (Figure S5). One possible explanation was that the MNase assay might not detect the Isw1 remodeling activities at the PYK1 gene. For example, the ISWI-dependent rescue may involve nucleosome repositioning at a level too subtle to be detected using the MNase mapping assay. Indeed, Isw1 and Isw2 remodeling activities increase genome-wide nucleosome occupancy at mid-coding regions and intergenic regions, respectively, to prevent cryptic transcription [46], [47]. In order to map nucleosome location with high resolution at the PYK1 gene, we took advantage of nucleosome-scanning analysis [48]. This method couples isolation of mononucleosomal DNA by MNase digestion with qPCR analysis using a set of overlapping primer pairs spanning the region of interest [48]. Nucleosome scanning analysis of the PYK1 region revealed the presence of four positioned nucleosomes, one located in the promoter region and three others positioned in the 5′ coding region (Figure 4). Further, a 150 bp region highly sensitive to MNase digestion has been identified in the promoter region (−350 to −200 from the start codon), indicating the presence of a nucleosome depleted region (NDR) (Figure 4). This organization is characteristic of “open” promoters which favor the binding of transcription factors at the expense of nucleosomes [49]. Open promoters are a common property of constitutive genes, such as the conditionally essential gene PYK1. No major changes in nucleosome positioning were observed upon inactivation of Gcn5 and Sas3, or further depletion of Ioc3 (Figure 4).
Fig. 4. Inactivation of the Isw1a complex in the gcn5Δ sas3 mutant does not significantly alter nucleosome positioning at the PYK1 gene. 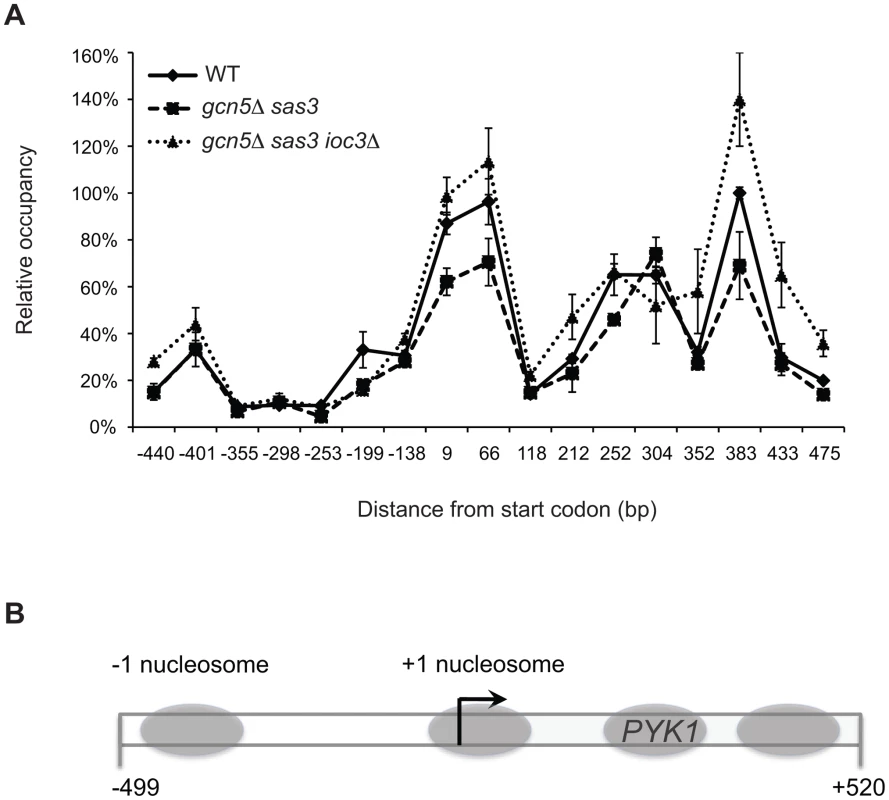
(A) A nucleosome scanning assay was performed on chromatin from WT, gcn5Δ sas3, and gcn5Δ sas3 ioc3Δ cells grown in SC medium at 34°C. Mononucleosomal DNA was purified and analyzed by real-time qPCR using seventeen overlapping primer pairs spanning the promoter and 5′ coding region of PYK1. Values are expressed as percentage of input, and normalized to the PHO5 TATA region [77]. The values represent the means from three independent experiments, with error bars reflecting standard deviations. (B) Diagram of the PYK1 locus indicates the positions of nucleosomes (gray ovals) extrapolated from the MNase protection assay. A dual role for the ATPase Isw1 revealed by functional interactions between the Isw1a and Isw1b complexes and the acetyltransferases Sas3 and Gcn5
Multiprotein complexes can be altered both by mutation and by changing subunit abundance through gene dosage. We took advantage of gene overexpression (reviewed in [50]–[52]) as an independent approach to probe the relationship of ISW1 to SAS3 and GCN5.
Increased gene dosage of IOC2 or IOC4 restored viability at elevated temperature, whereas overexpression of IOC3 exacerbated sickness of the gcn5Δ sas3 mutant (Figure 5A). Increased ISW1 also interfered with growth, confirming its generally antagonistic function. Furthermore, increased gene dosage of IOC2 did not rescue thermosensitivity of the gcn5Δ single mutant (Figure S6A), nor did it serve as a bypass suppressor of the gcn5Δ sas3Δ strain. Together these results suggest that increasing the stoichiometry of the Isw1b complex can ameliorate the negative effects of the Isw1a complex in the gcn5Δ sas3 mutant.
Fig. 5. Functional interactions between Isw1a/b complexes and acetyltransferases reveal a dual role for the ATPase Isw1.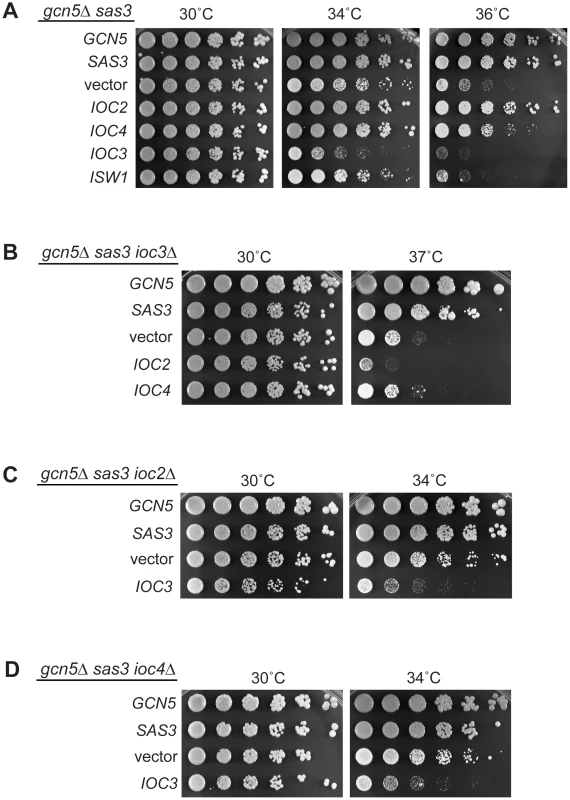 
(A) Overexpression of the Isw1b complex (Isw1/Ioc2/Ioc4) rescues the temperature sensitivity of the gcn5Δ sas3 mutant. In contrast, overexpression of the Isw1a complex (Isw1/Ioc3) exacerbates the growth defects of the HAT mutants. The gcn5Δ sas3 mutant was transformed with 2μ-LEU2 plasmids containing GCN5, SAS3, vector, IOC2, IOC4, IOC3 and ISW1. Transformants were plated onto SC–Leu medium and grown for 4 days. (B) Suppression of temperature sensitivity mediated by overexpression of the Isw1a complex (Isw1/Ioc2/Ioc4) requires a functional Isw1b complex (Isw1/Ioc3). The gcn5Δ sas3 ioc3Δ mutant was transformed with the indicated plasmids. Suppression was assayed by growth on SC–Leu medium. Note that the growth difference between the gcn5Δ sas3 ioc3Δ mutant in Figure 1C and the vector control shown here is due to the presence of sorbitol in the medium in Figure 1C, which partially relieves cell growth defects at elevated temperatures. Plating is shown here without sorbitol to increase the dynamic range of the assay. (C) and (D) Exacerbation of the temperature sensitivity mediated by overexpression of the Isw1b complex (Isw1/Ioc3) is not dependent on a functional Isw1a complex (Isw1/Ioc2/Ioc4). Transformants of the gcn5Δ sas3 ioc2Δ (C) and gcn5Δ sas3 ioc4Δ (D) mutants were assayed on SC–Leu medium. To determine if the Isw1a and Isw1b complexes act in parallel or in the same pathway, we performed a series of analyses to dissect the relative contribution of each complex. We first evaluated increased gene dosage of the Isw1b complex components in a strain depleted for the Isw1a complex. Of note, deletion of IOC3 partially rescues the gcn5Δ sas3 temperature sensitivity. Therefore, although the gcn5Δ sas3 ioc3Δ strain is less sensitive than the gcn5Δ sas3 (Figure 1C), this mutant is still somewhat temperature sensitive (Figure 5B), and provides the possibility for a dynamic range in which growth enhancement or inhibition could be observed. We found that overexpression of IOC2 or IOC4 did not rescue the residual gcn5Δ sas3 temperature sensitivity in an ioc3Δ background, demonstrating that the Isw1a complex is required for Isw1b-mediated dosage suppression (Figure 5B).
We next evaluated whether Isw1a overexpression enhanced the gcn5Δ sas3 phenotype in a strain depleted for Isw1b by deleting IOC2 and IOC4. Overexpression of IOC3 still exacerbated the gcn5Δ sas3 temperature sensitivity in a strain depleted for Isw1b (Figure 5C and Figure 5D). These results support the idea that increasing the relative balance of the Isw1b complex suppresses the gcn5Δ sas3 phenotype by counteracting Isw1a function. In addition, we observed that deletion of ISW1 interfered with the phenotypes resulting from overexpression of the IOC genes in the gcn5Δ sas3 strain, underscoring the critical role for the Isw1 catalytic ATPase itself (Figure S6B).
Discussion
Since the acetyltransferase activities of Sas3 and Gcn5 were initially reported to be essential [22], understanding the molecular mechanisms underlying this synthetic lethality has remained incomplete. Indeed, whereas disruption of these H3-specific acetyltransferases results in cell death, mutation of lysine residues in histone H3 that are known to be targeted by Sas3 and Gcn5 has only modest phenotypes [27], [28]. This discrepancy and the growing number of validated non-histone acetylation targets [53], [54] strongly suggest that the Sas3-Gcn5 essential function may reside in acetylation of histone and/or non-histone targets.
Our previous work demonstrated that the Sas3 and Gcn5 acetyltransferases are jointly required for viability and are responsible for the majority of histone H3 acetylation at K9 and K14 in vivo [22]. Here we show that both HATs contribute to in vivo acetylation of H3K14 at an actively transcribed gene, and furthermore we correlate the dramatic decrease in H3K14 acetylation with defective recruitment of RNAPII to promoter and coding regions. This reinforces the view that H3K14 acetylation is an epigenetic mark associated with transcriptional activation [20]. Moreover, the joint contributions of Sas3 and Gcn5 provide a molecular explanation for previous results showing that loss of Gcn5 only modestly affects acetylation of H3K14 at various genes [23], [55].
We found that inactivation of ISWI family remodelers alleviates gcn5Δ sas3 cell death. In deep contrast, inactivation of the Chd1 remodeler exacerbates the sickness of the gcn5Δ sas3 strain. Unlike other chromatin remodeler families, most ISWI complexes are required for formation of repressed structures [32], [44], [46], [56], [57] and chromosome compaction in vivo [58]–[60]. Furthermore, inactivation of the linker histone H1, another critical player in chromatin condensation, also rescues the growth defect of a gcn5Δ sas3 mutant [28]. Thus, destabilizing chromatin compaction, repressed structures, or higher-order chromatin structures through inactivation of ISWI complexes or histone H1 partially relieves growth defects in gcn5Δ sas3 cells. This supports the view that Sas3 and Gcn5 acetyltransferase activities counterbalance negative effects of repressed structures and condensed chromatin. In agreement with this model, we observed that inactivation of the Isw1a complex rescues RNAPII recruitment at active genes in a gcn5Δ sas3 mutant, to levels similar to those observed in the gcn5Δ mutant. This correlates with the degree of rescue observed for cell viability, strongly suggesting that Isw1-dependent rescue is mediated at least partially through the recruitment of RNAPII at actively transcribed genes. Similarly, the H4 HAT Esa1 can overcome the repressing function of Isw1 on transcription [61]. Conditional inactivation of Esa1 leads to defects in RNAPII recruitment at the MET16 gene upon induction, as well as impaired accumulation of MET16 transcript. Deletion of ISW1 was also seen to restore MET16 RNA levels and RNAPII distribution, which is very similar to our observations with RNAPII recruitment at active genes in the gcn5Δ sas3 mutant. However, the outcomes of these genetic interactions are clearly different: whereas inactivation of Isw1 rescues the sickness of gcn5Δ sas3 cells, it exacerbates cell growth defects of esa1 mutants [62], [63]. These differences are likely to reflect the distinct biological functions and substrates of the Sas3/Gcn5 and Esa1 acetyltransferases.
In contrast with Sas3 and Gcn5 acetyltransferases, functional characterization of ISWI enzymes in transcriptional regulation reveals significant roles in gene repression [35], [46], [47], [56]. Future studies should determine if Sas3 and Gcn5 acetyltransferases and ISWI family remodelers act in the same pathway.
In Drosophila, H4K16 acetylation regulates chromatin compaction by reducing the ability of ISWI to bind chromatin [64], [65]. Such a mechanism has not been described for H4 acetylation in yeast, and we show here that loss of the main H3 acetyltrasferases Sas3 and Gcn5 only modestly affects Isw1a recruitment to chromatin at some active genes (Figure 2). Thus it appears that Sas3 and Gcn5 acetyltransferase activities may counteract Isw1 function independently of chromatin binding. Such regulation has been reported for the Chd1 and Isw2 remodeling enzymes: H4 acetylation antagonizes nucleosome remodeling by lowering the catalytic turnover of ATP hydrolysis without affecting nucleosome binding [38]. Further, Sas3 and Gcn5 acetyltransferases might control Isw1 function directly through acetylation. In Drosophila, Gcn5 acetylates Isw1 at a single lysine in vitro and in vivo [66]. This acetylation occurs in a region similar to the N-terminal tail of H3, at a residue corresponding to K14. We also detected low levels of Isw1 acetylation in wild-type cells, with a two-fold decrease in the gcn5Δ sas3 mutant (data not shown).
Although loss of the ATP-dependent remodeling activity of Isw1 is required to restore cell viability, we observed no significant effects on nucleosome positioning upon inactivation of Isw1a at the PYK1 gene. Of note, nucleosome occupancy appears slightly reduced in the gcn5Δ sas3 mutant, at the predicted nucleosomes +1 and +3 (Figure 4). This appears in contrast with the increased H3 occupancy observed at the PYK1 promoter and coding region by ChIP (Figure 2F). However, it should be noted that nucleosome occupancy assayed by the MNase scanning method was normalized to the PHO5 promoter while the ChIPs for H3 were not, which is likely to account for this discrepancy. Alternatively, these results may suggest a local change in nucleosome assembly or disassembly in gcn5Δ sas3 mutants that results in accumulation of incomplete nucleosomes. Addition and removal of the H3/H4 tetramer is the first and last step of nucleosome assembly and disassembly, respectively (reviewed in [67]). Tetrasomes protect around 80 base pairs of DNA, below the resolution of our MNase-qPCR study, which may contribute to the loss of nucleosome signal in the MNase assay. This possibility can be explored in future studies by evaluating H2B occupancy at the 5′ region of PYK1 and by testing genetic interactions between gcn5Δ sas3 and histone chaperones.
An additional possibility for the ISWI-dependent chromatin remodeling rescue in gcn5Δ sas3 mutants is that it may be mediated through alteration of higher order chromatin structures. Indeed, the Drosophila ISWI-containing remodeler ACF can assemble regularly spaced arrays of H1-containing nucleosomes and can further catalyze repositioning of chromatosomes (nucleosome+H1) in chromatin fibers [68], [69]. Furthermore, inactivation of histone H1 also rescues the growth defect of a gcn5Δ sas3 mutant [28]. Therefore, restoration of RNAPII upon inactivation of Isw1a complex might rely on different states of chromatin that differ in the periodicity of chromatosome arrays.
An unexpected finding uncovered by our genetic analysis is the opposing functions of Isw1a and Isw1b complexes in relation to the H3 HATs. Specifically, we observed that rescue mediated by Ioc2 and Ioc4 requires Ioc3, but this requirement is not reciprocal. This suggests that, at least in the context of gcn5Δ sas3, the Isw1b complex antagonizes the function of the Isw1a complex. Early on, functional characterization of these two complexes revealed distinct roles [35]. Notably they differ in their biochemical activities to bind and space nucleosomes [35], [70], [71], their nucleosome remodeling properties in vivo [46], [47], as well as their roles in transcriptional regulation [35], [44]. Similarly in other eukaryotes, ISWI complexes that share the same catalytic subunit have distinct biological functions, specified by their associated proteins (reviewed in [1], [39]). The observations reported here bring new understanding toward defining their differences by showing that one ISWI complex may counteract the function of another ISWI complex.
Together, the results from this study deepen understanding of the essential roles for H3 HATs. Not only do the HATs positively promote gene-specific transcriptional activation, as has been well established, they also have a critical role in balancing the activities of ATP-driven chromatin remodelers. The functional antagonism between Isw1a and the Sas3 and Gcn5 acetyltransferases further defines biological distinctions between the Isw1 enzyme's catalytic activities in its two structurally distinct complexes. Thus, in addition to interactions between histone modifications defining transcriptional output (reviewed in [6]–[8]), it is likely that future studies will reveal increasingly diverse interactions between the modifying machines and the complexes that dynamically define chromatin architecture through its remodeling.
Materials and Methods
Yeast strains and plasmids
Strains used are listed in Table S1 and are in the W303 background. Gene deletions and other standard procedures were performed as described [72]. The gcn5Δ sas3 conditional mutant was constructed with a chromosomal allele of the sas3-C357Y, P375A double point mutation. As described for the plasmid conditional mutant [22], the chromosomal version of the mutant grows well at 30°C, but dies at 37°C. All strains carrying isw1Δ::kanMX, ioc2Δ::kanMX, ioc3Δ::natMX, ioc4Δ::hphMX, isw1-K227R and isw2-K215R-3FLAG-kanMX alleles were derived from the strains YTT441, YTT823, YTT825, YTT855 [35], YTT1223 and YTT1996 respectively, generous gifts from T. Tsukiyama. The gcn5Δ::natMX allele was obtained by marker swapping using p4339 (generous gift from C. Boone) on gcn5Δ::kanMX. The strains expressing Ioc3-13Myc and Isw1-13Myc from their chromosomal loci were constructed as described in [73]. All plasmids were derived from Yep351 (2μ LEU2). pLP645 was constructed by inserting a BamHI-SalI fragment containing SAS3 into Yep351 opened with SalI and BamHI (J. Lowell). pLP1524 was obtained from a genomic library generously provided by S. Roeder [74]. It contains a 3.9 kb fragment encompassing GCN5 gene (Chr.VII 995,188 to 998,784 bp). To construct Yep351-IOC2 (pLP2234), a HindIII fragment (4.6 kb) containing IOC2 was subcloned from pLP2170 (containing genomic fragment from Chr.XII 328,038 to 332,847 bp) into Yep351. To create Yep351-ISW1 (pLP2256), a BamHI-PstI fragment was subcloned from pRS416-ISW1 (a generous gift from T. Tsukiyama) into Yep351. Plasmid Yep351-IOC4 (pLP2260) was constructed by PCR amplification of IOC4 (−395 bp from start codon to +412 bp from stop codon), and cloned into Yep351 using HindIII-PstI restriction sites. Plasmid Yep351-IOC3 (pLP2266) was constructed by PCR amplification of IOC3 (−700 bp from start codon to +400 bp from stop codon), and cloned into Yep351 using HindIII-PstI restriction sites. Integrity of the constructs was confirmed by DNA sequencing.
Temperature sensitivity assays
Cultures were grown for 2 days in SC or appropriate selective medium at permissive temperature. Cells were diluted to an A600 of 1 and plated in fivefold serial dilutions onto SC or selective medium, supplemented with 1 M sorbitol where indicated, and incubated for 4 days at the indicated temperatures prior to data collection.
ChIP
Chromatin immunoprecipitation assays were performed as described previously [75] with minor modifications. Cultures were grown in SC medium at 34°C to A600 of 0.7–0.9 and cross-linked with 0.86% formaldehyde for 40 min. Immunoprecipitations (IP) were either pre-cleared with CL4B Sepharose beads (Sigma) for 1 hour at 4°C, then incubated overnight at 4°C with anti-RNAPII (8GW16, Covance) or directly incubated overnight at 4°C with antibodies against H3 (07-690, Upstate/Millipore), acetylated H3K14 (07-353, Upstate/Millipore), or the myc epitope (9E10). DNA was purified using PCR purification columns (Qiagen) and analyzed by real-time PCR (MJ Research Opticon2 system). Primer sequences are listed in Table S2. For quantification of ChIP samples, standard curves were generated for each set of primers, and DNA from IP and input samples was assayed for each strain in triplicate real-time PCR reactions. Each IP sample was normalized to the control IP (i.e. no epitope or no antibody) by subtraction, divided by the input sample, and expressed as percent of input (% IP/input). The % IP/input values for H3K14Ac were further normalized to % IP/input values for total H3. The % IP/input values for Ioc3-Myc and RNAPII were normalized to telVI and rDNA 5S control regions, respectively. Data represent averages from two or more independent experiments.
Nucleosome scanning analysis
Extracts from MNase digestions were prepared as described [76], [77]. Briefly, cultures were grown in SC medium at 34°C to an A600 of 0.7–0.9. Then ∼2×109 cells were harvested, washed in 1 ml sorbitol 1 M, resuspended in 1 ml of zymolyase solution (sorbitol 1.1 M, 20 mM KPO4, pH7, 0.5 mM CaCl2, β-mercaptoethanol 0.5 mM, zymolyase 100T 1 mg/ml) and incubated for 1.5 min at room temperature. Spheroplasts were then washed twice in 1 M sorbitol and gently resuspended in 1.6 ml of cold buffer A (1 M sorbitol, 50 mM NaCl, 10 mM Tris-HCl, pH 7.4, 5 mM MgCl2, 0.5 mM spermidine, 0.075% NP40 and 1 mM β-mercaptoethanol). The cell slurry was divided into 400 µl aliquots, and each was added to a microfuge tube containing the MNase (0, 60, 150 and 400 U/ml final concentrations) and incubated at 37°C for 4 minutes. The reaction was stopped by addition of 40 µl of stop buffer (250 mM EDTA, 5% SDS). DNA purification was performed as described in [76]. MNase digested DNA was run out on a 1.5% agarose gel and mononucleosome sized fragments were excised and purified using Qiagen's Gel Extraction kit. Purified mononucleosomes were analyzed by real-time PCR using the MJ Research Opticon2 system. Primer sequences are listed in Table S2. For quantification of MNase digested samples, standard curves were generated for each set of primers. Digested and input DNAs were assayed for each strain with each primer set in triplicate PCR reactions. MNase digested samples were divided by the input value for each primer set to generate percent of input. This was further normalized to the PHO5 TATA region [78].
The protein immunoblotting, mRNA quantification, and chromatin analysis techniques used in Figures S2, S3, S4, S5 are described in Supporting Methods (Text S1).
Supporting Information
Zdroje
1. ClapierCR, CairnsBR (2009) The biology of chromatin remodeling complexes. Annu Rev Biochem 78 : 273–304.
2. KorberP, BeckerPB (2010) Nucleosome dynamics and epigenetic stability. Essays Biochem 48 : 63–74.
3. KouzaridesT (2007) Chromatin modifications and their function. Cell 128 : 693–705.
4. TavernaSD, LiH, RuthenburgAJ, AllisCD, PatelDJ (2007) How chromatin-binding modules interpret histone modifications: lessons from professional pocket pickers. Nat Struct Mol Biol 14 : 1025–1040.
5. SuganumaT, WorkmanJL (2008) Crosstalk among Histone Modifications. Cell 135 : 604–607.
6. NarlikarGJ, FanHY, KingstonRE (2002) Cooperation between complexes that regulate chromatin structure and transcription. Cell 108 : 475–487.
7. BergerSL (2007) The complex language of chromatin regulation during transcription. Nature 447 : 407–412.
8. LiB, CareyM, WorkmanJL (2007) The role of chromatin during transcription. Cell 128 : 707–719.
9. van AttikumH, GasserSM (2009) Crosstalk between histone modifications during the DNA damage response. Trends Cell Biol 19 : 207–217.
10. PollardKJ, PetersonCL (1997) Role for ADA/GCN5 products in antagonizing chromatin-mediated transcriptional repression. Mol Cell Biol 17 : 6212–6222.
11. RobertsSM, WinstonF (1997) Essential functional interactions of SAGA, a Saccharomyces cerevisiae complex of Spt, Ada, and Gcn5 proteins, with the Snf/Swi and Srb/mediator complexes. Genetics 147 : 451–465.
12. SudarsanamP, CaoY, WuL, LaurentBC, WinstonF (1999) The nucleosome remodeling complex, Snf/Swi, is required for the maintenance of transcription in vivo and is partially redundant with the histone acetyltransferase, Gcn5. EMBO J 18 : 3101–3106.
13. SyntichakiP, TopalidouI, ThireosG (2000) The Gcn5 bromodomain co-ordinates nucleosome remodelling. Nature 404 : 414–417.
14. HassanAH, NeelyKE, WorkmanJL (2001) Histone acetyltransferase complexes stabilize swi/snf binding to promoter nucleosomes. Cell 104 : 817–827.
15. WoodageT, BasraiMA, BaxevanisAD, HieterP, CollinsFS (1997) Characterization of the CHD family of proteins. Proc Natl Acad Sci U S A 94 : 11472–11477.
16. TranHG, StegerDJ, IyerVR, JohnsonAD (2000) The chromo domain protein chd1p from budding yeast is an ATP-dependent chromatin-modifying factor. EMBO J 19 : 2323–2331.
17. Pray-GrantMG, DanielJA, SchieltzD, YatesJR3rd, GrantPA (2005) Chd1 chromodomain links histone H3 methylation with SAGA - and SLIK-dependent acetylation. Nature 433 : 434–438.
18. KastenM, SzerlongH, Erdjument-BromageH, TempstP, WernerM, et al. (2004) Tandem bromodomains in the chromatin remodeler RSC recognize acetylated histone H3 Lys14. EMBO J 23 : 1348–1359.
19. VanDemarkAP, KastenMM, FerrisE, HerouxA, HillCP, et al. (2007) Autoregulation of the rsc4 tandem bromodomain by gcn5 acetylation. Mol Cell 27 : 817–828.
20. PokholokDK, HarbisonCT, LevineS, ColeM, HannettNM, et al. (2005) Genome-wide map of nucleosome acetylation and methylation in yeast. Cell 122 : 517–527.
21. DurantM, PughBF (2006) Genome-wide relationships between TAF1 and histone acetyltransferases in Saccharomyces cerevisiae. Mol Cell Biol 26 : 2791–2802.
22. HoweL, AustonD, GrantP, JohnS, CookRG, et al. (2001) Histone H3 specific acetyltransferases are essential for cell cycle progression. Genes Dev 15 : 3144–3154.
23. KristjuhanA, WalkerJ, SukaN, GrunsteinM, RobertsD, et al. (2002) Transcriptional inhibition of genes with severe histone h3 hypoacetylation in the coding region. Mol Cell 10 : 925–933.
24. RobertF, PokholokDK, HannettNM, RinaldiNJ, ChandyM, et al. (2004) Global position and recruitment of HATs and HDACs in the yeast genome. Mol Cell 16 : 199–209.
25. LafonA, ChangCS, ScottEM, JacobsonSJ, PillusL (2007) MYST opportunities for growth control: yeast genes illuminate human cancer gene functions. Oncogene 26 : 5373–5384.
26. RosalenyLE, Ruiz-GarciaAB, Garcia-MartinezJ, Perez-OrtinJE, TorderaV (2007) The Sas3p and Gcn5p histone acetyltransferases are recruited to similar genes. Genome Biol 8: R119.
27. ZhangW, BoneJR, EdmondsonDG, TurnerBM, RothSY (1998) Essential and redundant functions of histone acetylation revealed by mutation of target lysines and loss of the Gcn5p acetyltransferase. EMBO J 17 : 3155–3167.
28. ChoiJK, GrimesDE, RoweKM, HoweLJ (2008) Acetylation of Rsc4p by Gcn5p is essential in the absence of histone H3 acetylation. Mol Cell Biol 28 : 6967–6972.
29. LaurentBC, YangX, CarlsonM (1992) An essential Saccharomyces cerevisiae gene homologous to SNF2 encodes a helicase-related protein in a new family. Mol Cell Biol 12 : 1893–1902.
30. ShenX, MizuguchiG, HamicheA, WuC (2000) A chromatin remodelling complex involved in transcription and DNA processing. Nature 406 : 541–544.
31. NeigebornL, CarlsonM (1984) Genes affecting the regulation of SUC2 gene expression by glucose repression in Saccharomyces cerevisiae. Genetics 108 : 845–858.
32. TsukiyamaT, PalmerJ, LandelCC, ShiloachJ, WuC (1999) Characterization of the imitation switch subfamily of ATP-dependent chromatin-remodeling factors in Saccharomyces cerevisiae. Genes Dev 13 : 686–697.
33. GelbartME, RechsteinerT, RichmondTJ, TsukiyamaT (2001) Interactions of Isw2 chromatin remodeling complex with nucleosomal arrays: analyses using recombinant yeast histones and immobilized templates. Mol Cell Biol 21 : 2098–2106.
34. FazzioTG, TsukiyamaT (2003) Chromatin remodeling in vivo: evidence for a nucleosome sliding mechanism. Mol Cell 12 : 1333–1340.
35. VaryJCJr, GangarajuVK, QinJ, LandelCC, KooperbergC, et al. (2003) Yeast Isw1p forms two separable complexes in vivo. Mol Cell Biol 23 : 80–91.
36. FlanaganJF, MiLZ, ChruszczM, CymborowskiM, ClinesKL, et al. (2005) Double chromodomains cooperate to recognize the methylated histone H3 tail. Nature 438 : 1181–1185.
37. CareyM, LiB, WorkmanJL (2006) RSC exploits histone acetylation to abrogate the nucleosomal block to RNA polymerase II elongation. Mol Cell 24 : 481–487.
38. FerreiraH, FlausA, Owen-HughesT (2007) Histone modifications influence the action of Snf2 family remodelling enzymes by different mechanisms. J Mol Biol 374 : 563–579.
39. YadonAN, TsukiyamaT (2011) SnapShot: Chromatin remodeling: ISWI. Cell 144 : 453–453 e451.
40. BoyerLA, LatekRR, PetersonCL (2004) The SANT domain: a unique histone-tail-binding module? Nat Rev Mol Cell Biol 5 : 158–163.
41. PinskayaM, NairA, ClynesD, MorillonA, MellorJ (2009) Nucleosome remodeling and transcriptional repression are distinct functions of Isw1 in Saccharomyces cerevisiae. Mol Cell Biol 29 : 2419–2430.
42. BoyerLA, LangerMR, CrowleyKA, TanS, DenuJM, et al. (2002) Essential role for the SANT domain in the functioning of multiple chromatin remodeling enzymes. Mol Cell 10 : 935–942.
43. YuJ, LiY, IshizukaT, GuentherMG, LazarMA (2003) A SANT motif in the SMRT corepressor interprets the histone code and promotes histone deacetylation. EMBO J 22 : 3403–3410.
44. MorillonA, KarabetsouN, O'SullivanJ, KentN, ProudfootN, et al. (2003) Isw1 chromatin remodeling ATPase coordinates transcription elongation and termination by RNA polymerase II. Cell 115 : 425–435.
45. GrantPA, EberharterA, JohnS, CookRG, TurnerBM, et al. (1999) Expanded lysine acetylation specificity of Gcn5 in native complexes. J Biol Chem 274 : 5895–5900.
46. WhitehouseI, RandoOJ, DelrowJ, TsukiyamaT (2007) Chromatin remodelling at promoters suppresses antisense transcription. Nature 450 : 1031–1035.
47. TiroshI, SigalN, BarkaiN (2010) Widespread remodeling of mid-coding sequence nucleosomes by Isw1. Genome Biol 11: R49.
48. SekingerEA, MoqtaderiZ, StruhlK (2005) Intrinsic histone-DNA interactions and low nucleosome density are important for preferential accessibility of promoter regions in yeast. Mol Cell 18 : 735–748.
49. CairnsBR (2009) The logic of chromatin architecture and remodelling at promoters. Nature 461 : 193–198.
50. RineJ (1991) Gene overexpression in studies of Saccharomyces cerevisiae. Methods Enzymol 194 : 239–251.
51. SopkoR, HuangD, PrestonN, ChuaG, PappB, et al. (2006) Mapping pathways and phenotypes by systematic gene overexpression. Mol Cell 21 : 319–330.
52. PrelichG (2012) Gene overexpression: uses, mechanisms, and interpretation. Genetics 190 : 841–854.
53. YangXJ, SetoE (2008) Lysine acetylation: codified crosstalk with other posttranslational modifications. Mol Cell 31 : 449–461.
54. SpangeS, WagnerT, HeinzelT, KramerOH (2009) Acetylation of non-histone proteins modulates cellular signalling at multiple levels. Int J Biochem Cell Biol 41 : 185–198.
55. SukaN, SukaY, CarmenAA, WuJ, GrunsteinM (2001) Highly specific antibodies determine histone acetylation site usage in yeast heterochromatin and euchromatin. Mol Cell 8 : 473–479.
56. GoldmarkJP, FazzioTG, EstepPW, ChurchGM, TsukiyamaT (2000) The Isw2 chromatin remodeling complex represses early meiotic genes upon recruitment by Ume6p. Cell 103 : 423–433.
57. KentNA, KarabetsouN, PolitisPK, MellorJ (2001) In vivo chromatin remodeling by yeast ISWI homologs Isw1p and Isw2p. Genes Dev 15 : 619–626.
58. DeuringR, FantiL, ArmstrongJA, SarteM, PapoulasO, et al. (2000) The ISWI chromatin-remodeling protein is required for gene expression and the maintenance of higher order chromatin structure in vivo. Mol Cell 5 : 355–365.
59. FyodorovDV, BlowerMD, KarpenGH, KadonagaJT (2004) Acf1 confers unique activities to ACF/CHRAC and promotes the formation rather than disruption of chromatin in vivo. Genes Dev 18 : 170–183.
60. CoronaDF, SiriacoG, ArmstrongJA, SnarskayaN, McClymontSA, et al. (2007) ISWI regulates higher-order chromatin structure and histone H1 assembly in vivo. PLoS Biol 5: e232 doi:10.1371/journal.pbio.0050232.
61. MorillonA, KarabetsouN, NairA, MellorJ (2005) Dynamic lysine methylation on histone H3 defines the regulatory phase of gene transcription. Mol Cell 18 : 723–734.
62. LindstromKC, VaryJCJr, ParthunMR, DelrowJ, TsukiyamaT (2006) Isw1 functions in parallel with the NuA4 and Swr1 complexes in stress-induced gene repression. Mol Cell Biol 26 : 6117–6129.
63. MitchellL, LambertJP, GerdesM, Al-MadhounAS, SkerjancIS, et al. (2008) Functional dissection of the NuA4 histone acetyltransferase reveals its role as a genetic hub and that Eaf1 is essential for complex integrity. Mol Cell Biol 28 : 2244–2256.
64. CoronaDF, ClapierCR, BeckerPB, TamkunJW (2002) Modulation of ISWI function by site-specific histone acetylation. EMBO Rep 3 : 242–247.
65. Shogren-KnaakM, IshiiH, SunJM, PazinMJ, DavieJR, et al. (2006) Histone H4-K16 acetylation controls chromatin structure and protein interactions. Science 311 : 844–847.
66. FerreiraR, EberharterA, BonaldiT, ChiodaM, ImhofA, et al. (2007) Site-specific acetylation of ISWI by GCN5. BMC Mol Biol 8 : 73.
67. LugerK, DechassaML, TremethickDJ (2012) New insights into nucleosome and chromatin structure: an ordered state or a disordered affair? Nat Rev Mol Cell Biol 13 : 436–447.
68. LusserA, UrwinDL, KadonagaJT (2005) Distinct activities of CHD1 and ACF in ATP-dependent chromatin assembly. Nat Struct Mol Biol 12 : 160–166.
69. MaierVK, ChiodaM, RhodesD, BeckerPB (2008) ACF catalyses chromatosome movements in chromatin fibres. EMBO J 27 : 817–826.
70. StockdaleC, FlausA, FerreiraH, Owen-HughesT (2006) Analysis of nucleosome repositioning by yeast ISWI and Chd1 chromatin remodeling complexes. J Biol Chem 281 : 16279–16288.
71. GangarajuVK, BartholomewB (2007) Dependency of ISW1a chromatin remodeling on extranucleosomal DNA. Mol Cell Biol 27 : 3217–3225.
72. Amberg D, Burke D, Strathern J (2005) Methods in yeast genetics: a Cold Spring Harbor Laboratory course manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
73. LongtineMS, McKenzieA3rd, DemariniDJ, ShahNG, WachA, et al. (1998) Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 14 : 953–961.
74. EngebrechtJ, HirschJ, RoederGS (1990) Meiotic gene conversion and crossing over: their relationship to each other and to chromosome synapsis and segregation. Cell 62 : 927–937.
75. DarstRP, GarciaSN, KochMR, PillusL (2008) Slx5 promotes transcriptional silencing and is required for robust growth in the absence of Sir2. Mol Cell Biol 28 : 1361–1372.
76. KentNA, MellorJ (1995) Chromatin structure snap-shots: rapid nuclease digestion of chromatin in yeast. Nucleic Acids Res 23 : 3786–3787.
77. WuL, WinstonF (1997) Evidence that Snf-Swi controls chromatin structure over both the TATA and UAS regions of the SUC2 promoter in Saccharomyces cerevisiae. Nucleic Acids Res 25 : 4230–4234.
78. WangSS, ZhouBO, ZhouJQ (2011) Histone H3 lysine 4 hypermethylation prevents aberrant nucleosome remodeling at the PHO5 promoter. Mol Cell Biol 31 : 3171–3181.
Štítky
Genetika Reprodukční medicína
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2012 Číslo 10
-
Všechny články tohoto čísla
- The Germline Genome Provides a Niche for Intragenic Parasitic DNA: Evolutionary Dynamics of Internal Eliminated Sequences
- Classical Genetics Meets Next-Generation Sequencing: Uncovering a Genome-Wide Recombination Map in
- Calpain-5 Mutations Cause Autoimmune Uveitis, Retinal Neovascularization, and Photoreceptor Degeneration
- Cofilin-1: A Modulator of Anxiety in Mice
- The Date of Interbreeding between Neandertals and Modern Humans
- Embryos of Robertsonian Translocation Carriers Exhibit a Mitotic Interchromosomal Effect That Enhances Genetic Instability during Early Development
- Viral Evasion of a Bacterial Suicide System by RNA–Based Molecular Mimicry Enables Infectious Altruism
- Phosphatase-Dead Myotubularin Ameliorates X-Linked Centronuclear Myopathy Phenotypes in Mice
- Full-Length Synaptonemal Complex Grows Continuously during Meiotic Prophase in Budding Yeast
- MOV10 RNA Helicase Is a Potent Inhibitor of Retrotransposition in Cells
- A Likelihood-Based Framework for Variant Calling and Mutation Detection in Families
- The Contribution of RNA Decay Quantitative Trait Loci to Inter-Individual Variation in Steady-State Gene Expression Levels
- New Partners in Regulation of Gene Expression: The Enhancer of Trithorax and Polycomb Corto Interacts with Methylated Ribosomal Protein L12 Its Chromodomain
- Mining the Unknown: A Systems Approach to Metabolite Identification Combining Genetic and Metabolic Information
- Mutations in (Hhat) Perturb Hedgehog Signaling, Resulting in Severe Acrania-Holoprosencephaly-Agnathia Craniofacial Defects
- The Many Landscapes of Recombination in
- Faster-X Evolution of Gene Expression in
- Loss of Slc4a1b Chloride/Bicarbonate Exchanger Function Protects Mechanosensory Hair Cells from Aminoglycoside Damage in the Zebrafish Mutant
- Regulation of ATG4B Stability by RNF5 Limits Basal Levels of Autophagy and Influences Susceptibility to Bacterial Infection
- and the BTB Adaptor Are Key Regulators of Sleep Homeostasis and a Dopamine Arousal Pathway in Drosophila
- Mutation and Fetal Ethanol Exposure Synergize to Produce Midline Signaling Defects and Holoprosencephaly Spectrum Disorders in Mice
- Specific Missense Alleles of the Arabidopsis Jasmonic Acid Co-Receptor COI1 Regulate Innate Immune Receptor Accumulation and Function
- Deep Genome-Wide Measurement of Meiotic Gene Conversion Using Tetrad Analysis in
- Mismatch Repair Balances Leading and Lagging Strand DNA Replication Fidelity
- Distinguishing between Selective Sweeps from Standing Variation and from a Mutation
- Cytokinesis-Based Constraints on Polarized Cell Growth in Fission Yeast
- Deposition of Histone Variant H2A.Z within Gene Bodies Regulates Responsive Genes
- Functional Antagonism between Sas3 and Gcn5 Acetyltransferases and ISWI Chromatin Remodelers
- The SET-Domain Protein SUVR5 Mediates H3K9me2 Deposition and Silencing at Stimulus Response Genes in a DNA Methylation–Independent Manner
- Morphogenesis and Cell Fate Determination within the Adaxial Cell Equivalence Group of the Zebrafish Myotome
- Muscle-Specific Splicing Factors ASD-2 and SUP-12 Cooperatively Switch Alternative Pre-mRNA Processing Patterns of the ADF/Cofilin Gene in
- Maize Is Required for Maintaining Silencing Associated with Paramutation at the and Loci
- Increasing Signal Specificity of the TOL Network of mt-2 by Rewiring the Connectivity of the Master Regulator XylR
- Use of Pleiotropy to Model Genetic Interactions in a Population
- RAB-Like 2 Has an Essential Role in Male Fertility, Sperm Intra-Flagellar Transport, and Tail Assembly
- Variants Affecting Exon Skipping Contribute to Complex Traits
- Topoisomerase II– and Condensin-Dependent Breakage of -Sensitive Fragile Sites Occurs Independently of Spindle Tension, Anaphase, or Cytokinesis
- Comparison of Family History and SNPs for Predicting Risk of Complex Disease
- Recovery of Arrested Replication Forks by Homologous Recombination Is Error-Prone
- A Mutation in the Gene Causes Alternative Splicing Defects and Deafness in the Bronx Waltzer Mouse
- Comparative Genomics Suggests an Independent Origin of Cytoplasmic Incompatibility in
- It Was Heaven: An Interview with Evelyn Witkin
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- A Mutation in the Gene Causes Alternative Splicing Defects and Deafness in the Bronx Waltzer Mouse
- Mutations in (Hhat) Perturb Hedgehog Signaling, Resulting in Severe Acrania-Holoprosencephaly-Agnathia Craniofacial Defects
- Classical Genetics Meets Next-Generation Sequencing: Uncovering a Genome-Wide Recombination Map in
- Regulation of ATG4B Stability by RNF5 Limits Basal Levels of Autophagy and Influences Susceptibility to Bacterial Infection
Přihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



