-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Kariérní portál
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaOut of Balance: R-loops in Human Disease
R-loops are cellular structures composed of an RNA/DNA hybrid, which is formed when the RNA hybridises to a complementary DNA strand and a displaced single-stranded DNA. R-loops have been detected in various organisms from bacteria to mammals and play crucial roles in regulating gene expression, DNA and histone modifications, immunoglobulin class switch recombination, DNA replication, and genome stability. Recent evidence suggests that R-loops are also involved in molecular mechanisms of neurological diseases and cancer. In addition, mutations in factors implicated in R-loop biology, such as RNase H and SETX (senataxin), lead to devastating human neurodegenerative disorders, highlighting the importance of correctly regulating the level of R-loops in human cells. In this review we summarise current advances in this field, with a particular focus on diseases associated with dysregulation of R-loop structures. We also discuss potential therapeutic approaches for such diseases and highlight future research directions.
Published in the journal: . PLoS Genet 10(9): e32767. doi:10.1371/journal.pgen.1004630
Category: Review
doi: https://doi.org/10.1371/journal.pgen.1004630Summary
R-loops are cellular structures composed of an RNA/DNA hybrid, which is formed when the RNA hybridises to a complementary DNA strand and a displaced single-stranded DNA. R-loops have been detected in various organisms from bacteria to mammals and play crucial roles in regulating gene expression, DNA and histone modifications, immunoglobulin class switch recombination, DNA replication, and genome stability. Recent evidence suggests that R-loops are also involved in molecular mechanisms of neurological diseases and cancer. In addition, mutations in factors implicated in R-loop biology, such as RNase H and SETX (senataxin), lead to devastating human neurodegenerative disorders, highlighting the importance of correctly regulating the level of R-loops in human cells. In this review we summarise current advances in this field, with a particular focus on diseases associated with dysregulation of R-loop structures. We also discuss potential therapeutic approaches for such diseases and highlight future research directions.
Introduction
R-loops are three-stranded structures, which form when RNA hybridises to a complementary DNA strand, forming an RNA/DNA hybrid, resulting in displacement of the other DNA strand in this process (Figure 1). The first R-loops were described in 1976, when their formation in vitro in the presence of 70% formamide was visualised by electron microscopy (Figure 1) [1]. These structures were thermodynamically more stable than duplex DNA, and they remained intact following removal of formamide. This technique of RNA/DNA hybridisation has been used in over 140 studies to map gene organisation, transcription initiation sites, and the direction of transcription, as well as measure the quantities of cellular RNAs [2].
Fig. 1. History of R-loop research. 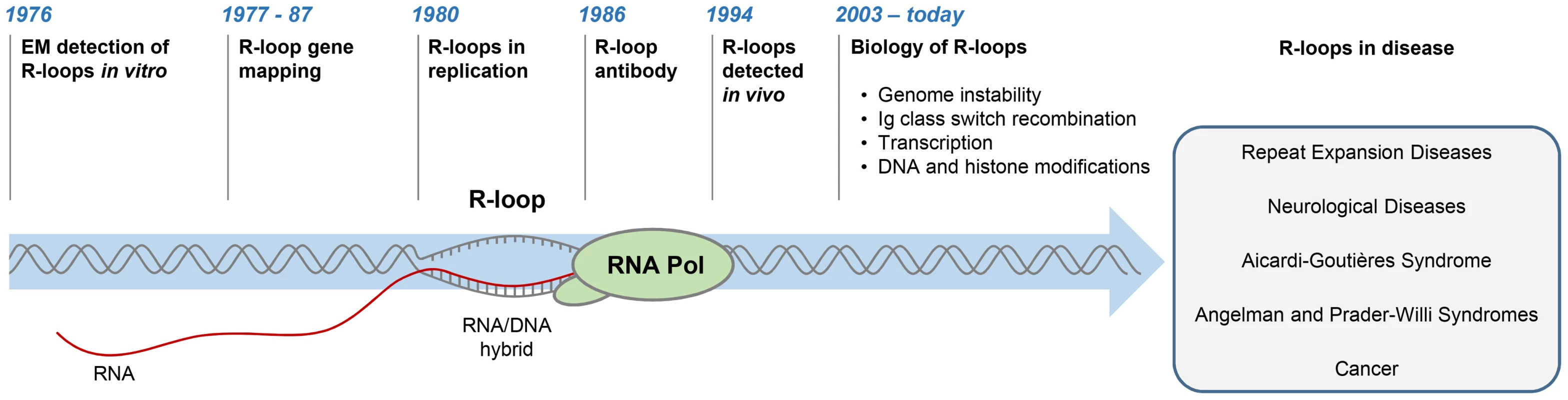
The diagram depicts major developments in the R-loop field and diseases associated with R-loop dysregulation. The first evidence for R-loop formation in live bacteria was obtained in 1994 [3]. This was followed by numerous studies showing that R-loops exist in different organisms (Figure 1) [4]–[6]. In living cells, R-loops are thought to form in cis during transcription, when nascent RNA hybridises to the DNA template behind the elongating RNA polymerase (Pol) [4]. However, in contrast to this popular view of cotranscriptional R-loops, recent studies suggest that RNA transcribed at one locus can hybridise to homologous DNA at another locus, thus leading to R-loop formation in trans [7]. In the last five years, the use of an antibody (S9.6) recognising RNA/DNA hybrids has revolutionised the R-loop field [8]. Initially, the S9.6 antibody, which detects hybrids as small as six bp with an affinity of 0.6 nM, was developed as a tool to enhance the DNA/RNA hybridisation signal in DNA microarray studies [9], [10]. More recently, it has been used to detect R-loops in vivo and uncover their contribution to fundamental biological processes in yeast [11], [12], plants [13], mice [14], [15], and humans [16]–[18].
The picture emerging from these studies suggests that R-loops can be both beneficial and deleterious to cells. Paradoxically, while they are required for important biological processes, they can also promote DNA damage and genome instability. In particular, R-loops have been shown to play an essential positive function in Escherichia coli plasmid and human mitochondrial DNA replication [19], [20] and during immunoglobulin class switch recombination, which contributes to the antibody isotype diversity in activated B cells [21]. R-loops form on many genes in yeast and human cells [18], [22] and have been implicated in regulation of gene expression. R-loops can repress transcription and promote transcriptional termination [16], [23], [24]. Furthermore, R-loops are clearly associated with epigenetic mechanisms governing transcription, including DNA methylation and posttranslational histone modifications [18], . In spite of this growing list of beneficial R-loop functions, it is also evident that R-loops can be a dangerous source of DNA damage. They can sensitize DNA to damaging agents [28], induce transcription-associated recombination [24], double-strand breaks (DSBs) [29], [30], chromosome breaks, and fragile site instability [31]–[33], and cause chromosome loss [34]. Therefore, cells need to tightly regulate the levels of R-loops to exploit their unique features. Altering the physiological R-loop balance can impair R-loop-regulated processes, cause genome instability, and may lead to human diseases. Consequently, defining the roles of R-loops in the multitude of biological processes and human disease is likely to develop into one of the most important and influential areas of R-loop research in the future.
Proteins in R-loop Biology
The number of proteins associated with R-loop biology has increased in the last few years, reflecting the diversity of R-loop processes (Table S1) [4]–[6]. Many proteins can regulate cellular R-loop levels either directly or indirectly, mostly by preventing RNA from hybridising to DNA, thus reducing excessive R-loop accumulation. Among these are proteins required for efficient transcriptional elongation, termination, polyadenylation, RNA splicing, packaging, and export [16], [24], [28], [30], [31], [34], [35]. DNA topology itself can influence hybridisation of RNA to DNA, and topoisomerases consequently play important roles in modulating R-loop levels [27], [33]. Proteins involved in maintenance of genome integrity can also regulate R-loops, suggesting a dynamic interplay between DNA repair and R-loop formation [7]. Importantly, cells possess dedicated enzymes, including the members of the RNase H family that specifically degrade the RNA in R-loops [36], and helicases that can unwind RNA/DNA hybrids [12], [16].
Recent evidence shows that R-loops can directly affect many gene expression–associated processes, including DNA methylation, posttranslational histone modifications, and transcription, by influencing the function of regulatory proteins [16],[18],[25],[26]. Despite the growing number of proteins involved in R-loop homeostasis and human disease, many questions still remain unanswered. For many proteins with documented in vitro RNA/DNA helicase activity (e.g., Pif1, the MCM complex), in vivo evidence is generally still lacking (Table S1) [37], [38]. Moreover, the molecular mechanisms underlying interactions between proteins and R-loops are poorly understood, and in many cases the connections to disease remain obscure.
R-loops and Neurological Diseases
The biological importance of R-loops in humans is supported by the fact that mutations in proteins implicated in R-loop resolution cause devastating human diseases, often related to neurodegeneration. Mutations in the putative RNA/DNA helicase SETX cause neurodegenerative diseases, the dominant juvenile form of amyotrophic lateral sclerosis type 4 (ALS4), and a recessive form of ataxia oculomotor apraxia type 2 (AOA2) (Figure 2A). These diseases are characterised by progressive degeneration of motor neurons in the brain and spinal cord, muscle weakness and atrophy [39]–[41].
Fig. 2. R-loops and human diseases. 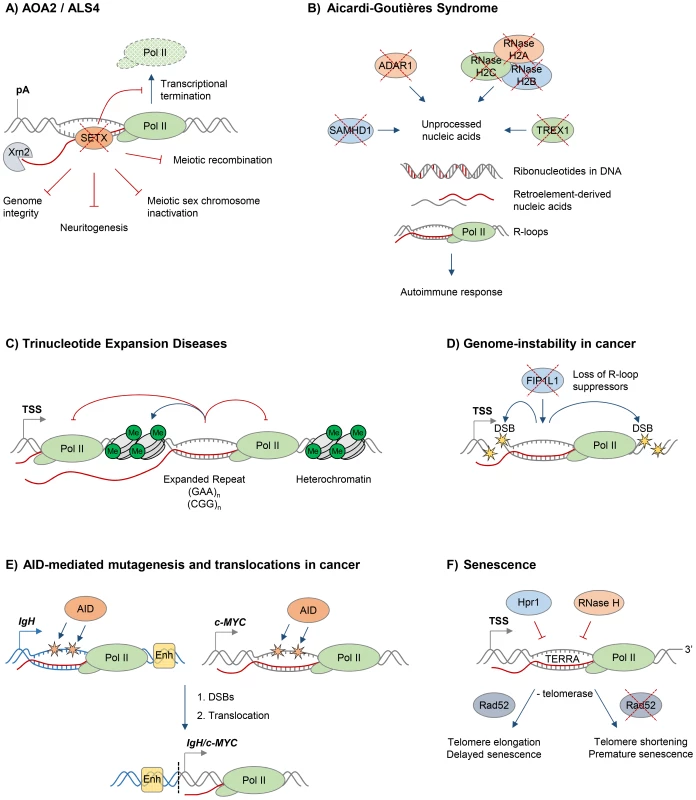
The diagram depicts the role of R-loops in human diseases. Loss of wild type protein function is depicted by red crosses. A. Ataxia and motor neuron diseases. Mutations in human RNA/DNA helicase senataxin are associated with AOA2/ALS4 disorders and lead to R-loop accumulation and defects in transcriptional termination by Pol II [16], the maintenance of genome integrity [46], meiotic recombination during spermatogenesis, gene silencing during meiotic sex chromosome inactivation [14], and neuronal differentiation [49]. B. Aicardi-Goutières syndrome (AGS). AGS is associated with mutations in all three subunits of RNase H2, ssDNA 3′–5′ exonuclease TREX1 (DNASEIII), dsRNA-editing enzyme ADAR1, and dNTP triphosphatase SAMHD1; these trigger accumulation of unprocessed nucleic acids, including genomic DNA with incorporated ribonucleotides, R-loops, and retroelement-derived nucleic acids, and result in the immune response characteristic of AGS [65]. C. Trinucleotide expansion diseases. R-loops form over expanded repeats and result in decreased initiation and elongation of RNA Pol II and formation of repressive chromatin marks, which silence the host gene containing expanded repeats [75]. D. Genome instability in cancer. Loss of proteins protecting against abnormal R-loop accumulation, such as FIP1L1, leads to genome instability, one hallmark of cancer [31]. Yellow stars denote double-stranded DNA breaks. E. AID-mediated mutagenesis and translocations in cancer. Single-stranded DNA in R-loops is a substrate for cytidine deamination by activation-induced cytidine deaminase, leading to mutagenesis as indicated by orange stars [21], [88]. These mutations can cause DSB formation, leading to chromosomal translocations. The IgH/c-MYC translocation brings the strong IgH enhancers, shown as yellow box, close to c-MYC, leading to its overexpression in Burkitt's lymphoma [87]. Transcription of IgH/c-MYC starts from a previously inactive promoter downstream of the translocation break point. The IgH locus is depicted in blue, c-MYC gene is in grey. The translocation breakpoint is indicated by a dashed black line. F. Senescence. R-loops formed by the noncoding RNA TERRA accumulate at telomeres in cells deficient of Hpr1 and RNase H. In the absence of telomerase, these R-loops promote Rad52-dependent telomere elongation and delayed senescence. In the absence of telomerase and Rad52, R-loops promote telomere shortening and premature senescence [94]. In addition to its predicted function as an RNA/DNA helicase, SETX interacts with proteins involved in diverse aspects of RNA metabolism [42]. Moreover, a single amino acid mutation, which compromises the function of the yeast homologue Sen1, dramatically changes the Pol II distribution genome-wide, further supporting the view that SETX/Sen1 functions in the regulation of transcription [43]. Recently, we demonstrated that SETX is implicated in transcriptional termination by Pol II in humans [16]. It is required to resolve R-loops at termination elements, releasing RNA for degradation by the 5′–3′ “torpedo” exonuclease Xrn2 prior to termination (Figure 2A) [16], [44]. Mutations in the yeast homologue, Sen1, also lead to a transcriptional termination defect, associated with accumulation of R-loops and genome instability [12]. In line with its function in R-loop resolution, SETX/Sen1 is also involved in maintaining genome integrity by coordinating transcription, DNA replication, and the DNA damage response [45]–[47]. SETX can target the 3′–5′ RNA degradation complex, the exosome, to sites of transcription-induced DNA damage [48]. Furthermore, SETX protects genome integrity by coordinating meiotic recombination with transcription during spermatogenesis and gene silencing during meiotic sex chromosome inactivation [14]. In particular, Setx knock-out mice accumulated DNA double strand breaks and R-loops and failed to disassemble Rad51 filaments. This resulted in a failure to cross over, likely due to collision between R-loops with Holliday junctions [14]. These defects in Setx knock-out mice lead to male infertility, raising the question as to how this relates to fertility of male AOA2/ALS4 patients.
Studies in neuronal cells have demonstrated a role for SETX in neuronal differentiation through fibroblast growth factor 8 (FGF8) signalling, providing one explanation for the effects of loss-of-function AOA2 mutations [49]. Surprisingly, overexpression of dominant mutant forms of SETX did not affect neuritogenesis, suggesting that a different function of SETX may be affected in ALS4 patients. However, the interplay between the function of SETX in R-loop resolution, genome maintenance, and neuronal differentiation is still unclear. In a recent study, Lavin and colleagues examined cells from mice with disrupted Atm, Tdp1, Setx, or Aptx genes, which cause ataxia telangiectasia (AT), spinocerebellar ataxia with axonal neuropathy 1 (SCAN1), AOA2, and ataxia oculomotor apraxia type 1 (AOA1) disorders, respectively [15]. These diseases are characterised by a defective response to DNA damage, suggesting that R-loops may be implicated in triggering genome instability. Indeed, R-loops were found to be enriched in proliferating cells (testes), but not in the brain tissues from Setx, Atm, Tdp1 or Aptx knock-out mice [15]. The enrichment of R-loops in testes correlated with high levels of DNA damage and apoptosis. The lack of R-loops in brain tissue questions the association between R-loops and neurodegeneration. This result is surprising, because inducible R-loops have been previously detected in neuronal cells at the Snord116 locus, which is associated with the neurodevelopmental disorder Angelman syndrome, as discussed below [50]. Furthermore, R-loops were implicated in inducing DNA damage in nonproliferating cells and post-mitotic neurons and proposed to contribute to the neurodegeneration seen in AT patients [29]. It is possible that R-loops are regulated by different mechanisms in proliferating cells and post-mitotic neurons, thereby leading to different R-loop kinetics and so preventing their detection in some model systems. In particular, R-loop accumulation may reflect collisions between transcription and replication machineries [32], [51], events which do not occur in postmitotic neurons. It should be noted that the mouse models currently used may not fully recapitulate all aspects of human neurodegeneration.
RNase H and Aicardi-Goutières Syndrome (AGS)
In addition to their generation during transcription, RNA/DNA hybrids can arise due to incorporation of ribonucleotides into DNA by DNA polymerases during replication. RNase H enzymes are endonucleases that cleave the RNA of RNA/DNA hybrids in a sequence-independent manner, thus maintaining genome stability by resolving R-loops that form during transcription and by removing misincorporated ribonucleotides from the DNA [36]. Eukaryotic cells have two types of these enzymes, RNase H1 and RNase H2, which have different enzymatic and site-specific activities [52]. In particular, RNase H1 requires a tract of at least four ribonucleotides to cleave the RNA/DNA hybrid, whereas RNase H2 can incise 5′ to a single ribonucleotide incorporated within a DNA molecule [36], [52]. Therefore, only RNase H2 can process single ribonucleotides in the DNA, but both enzymes are capable of eliminating RNA/DNA hybrids. Unlike in bacteria and unicellular eukaryotic organisms, where RNase H enzymes are dispensable for viability, both RNase H enzymes are essential in higher eukaryotes. RNase H1 has been implicated in mitochondrial DNA (mtDNA) replication during mouse development, a process likely to be associated with processing of RNA primers during mtDNA replication [53].
RNase H2 is composed of three different subunits, the catalytic subunit 2A, and two other subunits, 2B and 2C, all of which are required for enzyme activity. RNase H2 has been implicated in recognition and removal of ribonucleotides incorporated into DNA and hydrolysis of Okazaki fragment RNA primers during DNA replication [36], [54]–[57]. In addition, recent studies point towards a role of RNase H2 in R-loop resolution during transcription in vivo [11], [58]. In particular, deletion of Saccharomyces cerevisiae RNase H2 imposes transcriptional blocks and R-loop accumulation over rDNA regions in cells depleted of Topoisomerase I [11] and transcriptional down-regulation of genes with higher guanine-cytosine (GC) content at the promoter regions, which are likely to form stable R-loops [58].
In humans, mutations in any of the three subunits of RNase H2 cause Aicardi-Goutières syndrome (AGS), a neurological inflammatory disorder, which resembles a congenital viral infection and is associated with accumulation of ribonucleotides in the DNA (Figure 2B) [59], [60]. Interestingly, AGS can also be triggered by mutations in single-stranded DNA (ssDNA) 3′–5′ exonuclease TREX1(DNASEIII) [61], double-stranded RNA (dsRNA)-editing enzyme ADAR1 [62], and dNTP triphosphatase SAMHD1[63]. These proteins are involved in diverse pathways of nucleic acid metabolism, although their functions are not yet fully understood. They have been implicated in degrading ssDNA arising from endogenous retroelements or replication stress (TREX1), regulating the intracellular dNTPs pool available for replication and reverse transcription of these retroelements (SAMHD1), or altering the immune response to RNA species through RNA editing of retroelements and microRNAs (ADAR1) [64]. Mutations in these proteins are associated with an accumulation of unprocessed nucleic acids, which triggers the immune response characteristic of AGS [64], [65].
So far, pathologies linked to AGS mutations in RNase H2 have been mainly attributed to genome instability caused by accumulation of ribonucleotides in DNA [56], [66]. However, a specific contribution of R-loops and RNA/DNA hybrids to AGS pathology has not been yet investigated. This research has been hampered by the difficulty to uncouple the two activities of RNase H2; its ability to remove ribonucleotides from the DNA and to resolve R-loops, both of which are affected when RNase H2 is deleted [52], [56]. Nevertheless, several lines of evidence suggest that R-loops may be involved in AGS pathology. Thus, an AGS-related mutation in the yeast RNase H2 enzyme resulted in its reduced RNA/DNA cleavage activity [52]. Since RNase H2 constitutes ∼90% of the total cellular RNA/DNA hybrid cleavage activity, its loss due to AGS mutations may lead to significant accumulation of R-loops [56]. The importance of RNase H2 is further highlighted by the fact that mutations in RNase H1 do not cause AGS, suggesting that RNase H2 may have unique properties to degrade RNA/DNA hybrids [52]. Indeed, R-loops arising during DNA replication may be exclusively degraded by RNase H2, as they may be inaccessible to RNase H1 [52], [67]. A recently generated S. cerevisiae RNase H2 mutant, which possesses R-loop degrading activity but fails to remove single ribonucleotides from the DNA [52], will be a useful tool in addressing the contribution of unresolved transcription-associated R-loops to AGS pathology.
TREX1, ADAR1 and SAMHD1 process retroelement-derived nucleic acids and help to suppress retroelements expansion in the host genome and their recognition by the immune system [64]. Interestingly, recent genome-wide studies have demonstrated that RNA/DNA hybrids are particularly enriched at retrotransposon elements in yeast cells [22], suggesting that expansion of retroelements due to mutations in TREX1, ADAR1 or SAMHD1 may lead to increased RNA/DNA hybrid levels, contributing to autoimmunitity in AGS. Indeed, it has recently been demonstrated that RNA/DNA hybrids can be sensed by toll-like receptor 9 (TLR9) to induce pro-inflammatory cytokine and antiviral interferon production in dendritic cells [68].
R-loops in Nucleotide Expansion Diseases
Expansions of repetitive sequences have been linked to over forty human diseases [69], and R-loops have been proposed to play a role in their pathology [70]–[73]. Remarkably, R-loops are formed following transcription of trinucleotide repeats in vitro, in bacteria and human cells [70], [71], [73]. Interestingly, the nontemplate DNA strand in many repetitive sequences can adopt unusual DNA structures, including G-quadruplexes and DNA triplexes, which may further stabilise R-loops [74]. Moreover, R-loops formed at CTG repeats promote repeat instability characteristic of these diseases [71].
Recently, we demonstrated that R-loops form over expanded GAA and CGG repeats in cells from Friedreich's Ataxia (FRDA) and Fragile X syndrome (FXS) patients, respectively (Figure 2C) [75]. The abundance of these stable R-loops correlates with expansion size, and they colocalise with the repressive chromatin marks characteristic of these diseases (Figure 2C). R-loops can also trigger the formation of repressive chromatin and cause transcriptional silencing of the FXN gene, providing a molecular link between R-loops and the pathology of expansion diseases [75]. In line with R-loops formed on expanded “premutation” and “full mutation” CGG-repeat-containing alleles of the FMR1 gene [75], [76], promoter-bound FMR1 mRNA containing trinucleotide repeats was shown to promote epigenetic silencing in FXS [77]. Importantly, the involvement of R-loops in expansion diseases is not limited to trinucleotide repeats, since R-loops associated with expanded hexanucleotide GGGGCC repeats in C9orf72 contribute to the molecular event leading to amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD) [78].
R-loops could contribute to the pathology of expansion diseases in various ways. Similar to R-loops at the 3′ends of human genes, expansion-associated R-loops may form a structural block, directly interfering with Pol II transcriptional elongation [16], [24]. Alternatively, R-loops may nucleate repressive chromatin over the expansion region, by analogy with heterochromatin formation at centromeres in Schizosaccharomyces pombe [25], or promote chromatin compaction associated with histone H3S10 phosphorylation, as observed in S. cerevisiae, Caenorhabditis elegans, and human cells [26]. Furthermore, R-loops could cause the characteristic intergenerational and somatic instability of repeat sequences [72].
R-loops in Cancer
Genome instability is a hallmark of cancer, and it may actively drive hereditary tumour development [79], [80]. Research in the last decade has clearly demonstrated that dysregulation of R-loops can corrupt genome integrity, resulting in increased DNA sensitivity to damaging agents, formation of DSBs, chromosome breaks, fragile site instability, chromosome loss, and recombination events [5]. Several mechanisms have therefore evolved to maintain R-loop levels in balance, and alterations in genome caretaker processes can affect R-loop levels and genome stability [4]. Moreover, mutations in proteins controlling R-loop levels have been identified in tumours (Figure 2D). For example, in eosinophilic leukemia, an oncogenic translocation renders cleavage and polyadenylation factor FIP1L1 inactive, which has been previously shown to cause increased R-loop levels, DNA damage and chromosome instability (Figure 2D) [31]. A similar mechanism was suggested for RNA kinase CLP1, which is associated with a translocation in mixed lineage leukemia (MLL) [31]. The histone ubiquitin ligase BRE1 also limits R-loop levels, and its decreased expression may contribute to the high levels of genomic instability observed in testicular seminoma [81].
The link between R-loops and cancer has been further substantiated by the finding that the tumour suppressor BRCA2, which is mutated in breast and ovarian cancer, is required to prevent R-loop accumulation and genome instability [82]. These observations raise the interesting possibility that R-loops may provide proliferative advantages to tumour cells by promoting genome instability. This will in turn increase the probability of accumulating mutations favourable to tumour growth and metastasis. Intriguingly, recent evidence demonstrates that human oncogenic viruses may also promote genomic instability through accumulation of R-loops after infection. Kaposi's sarcoma-associated herpesvirus (KSHV), which causes multiple AIDS-related cancers, encodes the ORF57 protein, which can sequester the host hTREX complex, important for mRNA processing and export [83]. Sequestration of hTREX leads to KSHV-induced accumulation of R-loops and causes damage to the host DNA, contributing to tumourigenesis [83].
Whilst some proteins suppress R-loop formation, others may promote R-loops and so increase genome instability leading to tumour development. This unexpected function has been shown in yeast for transcription elongation factor Spt2 and DNA repair protein Rad51 [7], [84]. Overexpression of Spt2 leads to transcription-dependent chromosomal rearrangements, which are prevented by RNase H overexpression [84]. Spt2 is structurally related to human HMG1, which is overexpressed in gastric cancers and malignant melanomas [84]. However, it is not clear if increased HMG1 levels promote R-loops and DNA damage in cancer cells. In contrast to its well-established role in DNA strand exchange during homologous recombination and DNA repair [85], recent studies have shown that Rad51 can also mediate R-loop formation and genome instability in trans, extending the prevailing view that R-loops form cotranscriptionally [7]. Similar to HMG1, RAD51 is overexpressed in human cancers [7]. However, it remains to be elucidated if RAD51 overexpression in cancers is a consequence of activated DNA repair pathways, or a cause of genome instability [7].
R-loops have been detected in immunoglobulin (Ig) genes, where they initiate class switch recombination by exposing single-stranded DNA, thus providing the substrate for activation-induced cytidine deaminase (AID), which promotes DSBs and subsequent translocation between Ig heavy chains [21], [86]. Although this process is essential for generation of antibody isotype diversity, AID-mediated mutagenesis has also been implicated in pathological translocations between the Ig loci and other active genes, leading to production of fusion proteins or oncogenic gene expression, observed in B cell malignancies (Figure 2E) [87]. Interestingly, R-loops are also found in common translocation partners of Ig genes, including the oncogene c-MYC [18], [27]. Therefore, the simultaneous formation of R-loops in Ig and transcribed non-Ig genes may induce AID-mediated DSB formation, leading to pathological translocations (Figure 2E) [27], [88], [89]. Interestingly, overexpression of the APOBEC family of AID-related enzymes in breast cancer have been linked to genomic mutations, pointing to a potentially broader role of R-loops and AID/APOBEC-mediated genome instability in cancer [90].
Changes in gene expression are another central aspect of cancer [79]. In healthy cells, the expression of tumour suppressor genes prevents abnormal proliferation and other aspects of tumourigenesis [79]. Tumour suppressors are frequently silenced in cancer by excessive promoter DNA methylation [91]. It has been proposed that R-loop formation at promoters protects against DNA methylation by de novo DNA methyltransferase DNMT3B, thereby keeping genes active [18]. Since R-loops have been computationally predicted to form at promoters of tumour suppressor genes BRCA1, RASSF1A, and CDKN2A [92], it is important to investigate if R-loop levels at these genes are reduced in cancer and how this relates to the observed DNA hypermethylation.
In contrast to this, efficient transcription of the oncogene c-MYC requires that R-loop levels are kept low by the activity of DNA topoisomerase IIIB, which is recruited to arginine-methylated histones by the tudor domain containing 3 (TDRD3) protein [27]. This R-loop-mediated mechanism of c-MYC gene regulation may be relevant to tumour progression in breast cancer, which frequently shows overexpression of both c-MYC and TDRD3 [27], [93]. Therefore, it is tempting to speculate that increased TDRD3 levels suppress R-loops in c-MYC, thereby allowing its enhanced expression, which correlates with poor cancer prognosis [93]. However, it still remains to be determined if R-loops play a specific role in transcription dysregulation in cancer and if this process differs from R-loop-mediated transcriptional programmes associated with housekeeping genes.
More recently, R-loops have been implicated in cell senescence, a mechanism protecting against tumour cell proliferation [79]. In particular, the telomeric noncoding (nc) RNA TERRA forms R-loops which are induced when R-loop suppressors such as RNase H or Thp2 are lost [94], [95]. In the absence of telomerase, telomeric R-loops promote recombination-mediated telomere elongation via Rad52, and this delays the onset of cellular senescence [94]. In contrast, in Rad52-deficient cells, R-loop accumulation leads to telomere shortening and premature senescence [94]. Interestingly, cells from AOA2 patients with senataxin mutations contain shorter telomeres, suggesting a possible involvement of SETX in telomere stability [96]. Telomeric R-loops therefore play a complex and dynamic role in telomere length maintenance and cellular proliferative potential (Figure 2F).
In conclusion, multiple lines of evidence point to an involvement of R-loops in cancer biology. Yet it still remains to be investigated if R-loop levels are indeed regulated differentially in normal and tumour tissues and if they can directly influence tumourigenesis.
R-loop Therapies
R-loops represent a potential therapeutic target. Despite their importance in gene regulation, they have yet to be fully exploited in drug design [97]. Various ligands can target RNA/DNA hybrids, including ethidium bromide, the aminoglycosides neomycin and paramomycin, and the polyamides distamycin and netropsin [98]. These compounds recognise RNA/DNA hybrids through intercalation and binding to the nucleic acid groove. Although exhibiting high binding affinities to RNA/DNA hybrids, many of these molecules also bind dsDNA and RNA and are mutagenic, limiting their potential biological applications [98]. However, recent studies suggest that combining the properties of these ligands can achieve subnanomolar affinity for RNA/DNA hybrids. In particular, this has been demonstrated for ligands linking aminoglycosides to derivatives of ethidium bromide [99], providing a possible approach for the development of potent and specific RNA/DNA hybrid ligands in future drug design efforts.
Various compounds that modulate DNA supercoiling and inhibit DNA topoisomerases, including topotecan and camptothecin, can also affect R-loop formation in vivo [29], [50]. In particular, topoisomerase inhibitors have recently been used to reactivate the silenced paternal Ube3a gene, which encodes a ubiquitin E3 ligase, to compensate for the deleted maternal Ube3a in Angelman syndrome (AS). AS and Prader-Willi syndrome (PWS) are imprinted neurodevelopmental disorders that are often caused by large deletions of human chromosome 15q11–q13 over the Snord116 gene locus, but the deletion differs in its parent-of-origin [100]. In neurons, only the maternal Ube3a allele is expressed, because the paternal Ube3a allele is silenced by expression of the ncRNA Ube3a-ATS (Figure 3A) [101]. AS therapies therefore seek to reactivate the silenced, but genetically intact, paternal Ube3a allele. Interestingly, R-loops were recently shown to regulate the neuronal expression of the paternal Ube3a-ATS transcript, which is essential for transcriptional silencing of the paternal Ube3a gene [50]. In particular, treatment with the topoisomerase inhibitor topotecan increased R-loop levels over the Snord116 locus, resulting in chromatin decondensation, inhibition of Pol II transcription of Ube3a-ATS, and concomitant increase in Ube3a expression from the paternal allele (Figure 3B). This R-loop-mediated reactivation of paternal Ube3a could therefore compensate for the loss of maternal Ube3a in AS and so potentially holds promise for targeted therapies for both AS and PWS (Figure 3B).
Fig. 3. Potential R-loop-based therapeutic approach in Angelman Syndrome (AS). 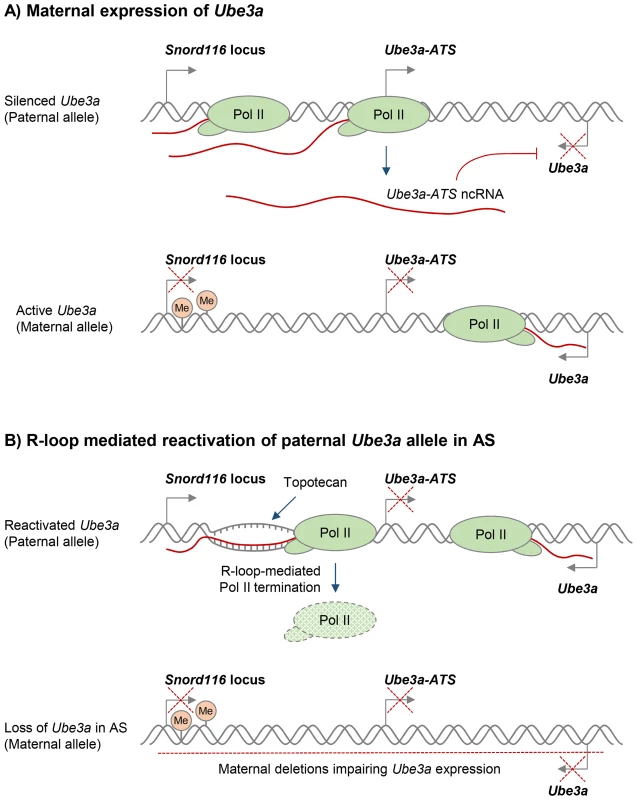
A. Neuronal expression of the paternal ncRNA Ube3a-ATS represses paternal Ube3a gene in cis [101]. DNA methylation of the Snord116 locus on the maternal allele prevents Ube3a-ATS transcription, resulting in Ube3a expression from the maternal allele. Transcriptional repression is indicated by red crosses. B. R-loop-mediated re-activation of silent paternal Ube3a gene provides a targeted therapy for AS. Deletion leading to the loss of maternal Ube3a expression detected in AS is indicated by the red dashed line. Topotecan treatment increases R-loop levels over the Snord116 locus, resulting in chromatin decondensation, inhibition of Pol II transcription through Ube3a-ATS, and increased expression of Ube3a from the paternal allele [50]. It has previously been proposed that R-loops in trinucleotide expansion diseases could be targeted to suppress repeat expansions or reactivate silenced genes [72]. A recent study provided direct evidence that a small molecule is able to suppress R-loop formation at expanded CGG repeats in the FMR1 gene, thereby preventing FMR1 epigenetic silencing in FXS [77]. As an alternative approach, R-loop levels may be indirectly modulated by treatments that target proteins involved in R-loop biology (Table S1). For instance, genomic instability caused by a widespread increase of R-loops due to loss of an R-loop suppressing protein could potentially be reverted by introduction of an alternative R-loop suppressor.
Recent identification of small-molecule inhibitors for RNase H2 may also provide a powerful new tool for the study of R-loop biology in health and disease [102]. Furthermore, the S9.6 antibody offers new opportunities for research and development. In particular, it has already been used in the development of biosensor systems [103], detection of miRNA targets [104], and as a key component of human papillomavirus (HPV) diagnostic kit (Qiagen).
The explosion of studies uncovering the role of R-loops in health and disease in recent years provides the exciting prospect of developing new targeted therapeutics for many human disorders. However, due to the ubiquitous nature of R-loops it will be important to ensure that efficient treatments are specific.
Conclusions and Future Challenges
R-loops have been implicated in many biological processes in different organisms. R-loops can play positive and negative roles in gene expression; they can mediate Ig class switch recombination and transcriptional termination, affect genome stability, transcription, cell cycle progression, and cell viability. Despite the diversity of these biological processes, the molecular mechanisms associated with R-loop formation in mammalian cells remain largely unknown. It is unclear how R-loops can regulate gene expression, how they are maintained and eliminated in the cells, and which proteins are involved in the regulation of these processes.
The connections between R-loops and human diseases suggest that cells have evolved mechanisms to distinguish between deleterious and beneficial R-loops. However, the evidence discussed above raises an important question: how can R-loop dysregulation be mechanistically linked to a variety of human diseases with such diverse pathologies? One explanation may be that R-loops form in many genomic locations in healthy cells [16], [18], [22], [27]. Therefore, unsurprisingly, their dysregulation can affect a large number of disease-associated genes. This is in contrast to gene-specific R-loop pathologies, associated with mutations, which result in altered R-loop levels locally, as observed in the repeat expansion diseases FRDA and FXS [75],[77]. Furthermore, R-loops can have different intrinsic properties. R-loops at expanded GAA repeats in the FXN gene are highly stable and trigger transcriptional repression, while R-loops in the highly-expressed γ-Actin gene are easily turned over [75]. This could, in part, be due to differential activity of R-loop processing proteins on different classes of genes, as proposed in yeast [22]. Adding another layer of complexity, the formation of R-loops can be influenced by cell type [77], cell cycle stage [15], gene length, and/or GC content and transcriptional level [22], [105]. Epigenetic marks including DNA methylation and post-translational histone modifications can contribute to further modulation of R-loop levels [18], [27]. Thus, R-loops represent cellular structures that share the same elementary composition, but may possess different dynamic properties, which can be affected by any of the aforementioned processes, thus explaining the wide range of diseases associated with R-loops.
Despite the lack of mechanistic insights into R-loop-associated diseases, some common themes, underlying their pathology, are already becoming obvious. First, there is a strong connection between R-loop dysregulation and induction of DNA damage and loss of genome integrity, which contributes to cancer development [31], [81], [88], repeat expansion diseases [71], and neurodegeneration [29], [45]. Secondly, R-loops can mediate changes in transcription locally or globally, contributing to pathologies associated with repeat expansion diseases [75], [77], [78], Angelman syndrome [50], and cancer [27]. However, it is a strong possibility that both of these pathological themes may overlap in many disorders, as observed in repeat expansion diseases [71], [75], [77], and novel disease themes may be revealed in the future.
One of the major challenges in R-loop field is to investigate the causes and consequences of R-loop formation in additional models of human disease. Uncovering further aspects of R-loop biology in human cells will certainly shed light on many basic biological questions and have major implications for our understanding of human disease. Future studies will undoubtedly reveal more diseases associated with R-loop dysregulation and will provide the basis for novel therapeutic approaches targeting these so far overlooked structures in gene expression.
Supporting Information
Zdroje
1. ThomasM, WhiteRL, DavisRW (1976) Hybridization of RNA to double-stranded DNA: formation of R-loops. Proc Natl Acad Sci U S A 73 : 2294–2298.
2. WhiteRL, HognessDS (1977) R loop mapping of the 18S and 28S sequences in the long and short repeating units of Drosophila melanogaster rDNA. Cell 10 : 177–192.
3. DroletM, BiX, LiuLF (1994) Hypernegative supercoiling of the DNA template during transcription elongation in vitro. J Biol Chem 269 : 2068–2074.
4. AguileraA, Garcia-MuseT (2012) R loops: from transcription byproducts to threats to genome stability. Mol Cell 46 : 115–124.
5. HamperlS, CimprichKA (2014) The contribution of co-transcriptional RNA∶DNA hybrid structures to DNA damage and genome instability. DNA Repair (Amst) 19 : 84–94.
6. Skourti-StathakiK, ProudfootNJ (2014) A double-edged sword: R loops as threats to genome integrity and powerful regulators of gene expression. Genes Dev 28 : 1384–1396.
7. WahbaL, GoreSK, KoshlandD (2013) The homologous recombination machinery modulates the formation of RNA-DNA hybrids and associated chromosome instability. Elife 2: e00505.
8. BoguslawskiSJ, SmithDE, MichalakMA, MickelsonKE, YehleCO, et al. (1986) Characterization of monoclonal antibody to DNA.RNA and its application to immunodetection of hybrids. J Immunol Methods 89 : 123–130.
9. PhillipsDD, GarbocziDN, SinghK, HuZ, LepplaSH, et al. (2013) The sub-nanomolar binding of DNA-RNA hybrids by the single-chain Fv fragment of antibody S9.6. J Mol Recognit 26 : 376–381.
10. HuZ, ZhangA, StorzG, GottesmanS, LepplaSH (2006) An antibody-based microarray assay for small RNA detection. Nucleic Acids Res 34: e52.
11. El HageA, FrenchSL, BeyerAL, TollerveyD (2010) Loss of Topoisomerase I leads to R-loop-mediated transcriptional blocks during ribosomal RNA synthesis. Genes Dev 24 : 1546–1558.
12. MischoHE, Gomez-GonzalezB, GrzechnikP, RondonAG, WeiW, et al. (2011) Yeast Sen1 helicase protects the genome from transcription-associated instability. Mol Cell 41 : 21–32.
13. SunQ, CsorbaT, Skourti-StathakiK, ProudfootNJ, DeanC (2013) R-loop stabilization represses antisense transcription at the Arabidopsis FLC locus. Science 340 : 619–621.
14. BecherelOJ, YeoAJ, StellatiA, HengEY, LuffJ, et al. (2013) Senataxin plays an essential role with DNA damage response proteins in meiotic recombination and gene silencing. PLoS Genet 9: e1003435.
15. YeoAJ, BecherelOJ, LuffJE, CullenJK, WongsurawatT, et al. (2014) R-Loops in Proliferating Cells but Not in the Brain: Implications for AOA2 and Other Autosomal Recessive Ataxias. PLoS ONE 9: e90219.
16. Skourti-StathakiK, ProudfootNJ, GromakN (2011) Human senataxin resolves RNA/DNA hybrids formed at transcriptional pause sites to promote Xrn2-dependent termination. Mol Cell 42 : 794–805.
17. GinnoPA, LimYW, LottPL, KorfI, ChedinF (2013) GC skew at the 5′ and 3′ ends of human genes links R-loop formation to epigenetic regulation and transcription termination. Genome Res 23 : 1590–1600.
18. GinnoPA, LottPL, ChristensenHC, KorfI, ChedinF (2012) R-loop formation is a distinctive characteristic of unmethylated human CpG island promoters. Mol Cell 45 : 814–825.
19. XuB, ClaytonDA (1996) RNA-DNA hybrid formation at the human mitochondrial heavy-strand origin ceases at replication start sites: an implication for RNA-DNA hybrids serving as primers. EMBO J 15 : 3135–3143.
20. ItohT, TomizawaJ (1980) Formation of an RNA primer for initiation of replication of ColE1 DNA by ribonuclease H. Proc Natl Acad Sci U S A 77 : 2450–2454.
21. YuK, ChedinF, HsiehCL, WilsonTE, LieberMR (2003) R-loops at immunoglobulin class switch regions in the chromosomes of stimulated B cells. Nat Immunol 4 : 442–451.
22. ChanYA, AristizabalMJ, LuPY, LuoZ, HamzaA, et al. (2014) Genome-Wide Profiling of Yeast DNA∶RNA Hybrid Prone Sites with DRIP-Chip. PLoS Genet 10: e1004288.
23. TousC, AguileraA (2007) Impairment of transcription elongation by R-loops in vitro. Biochem Biophys Res Commun 360 : 428–432.
24. HuertasP, AguileraA (2003) Cotranscriptionally formed DNA∶RNA hybrids mediate transcription elongation impairment and transcription-associated recombination. Mol Cell 12 : 711–721.
25. NakamaM, KawakamiK, KajitaniT, UranoT, MurakamiY (2012) DNA-RNA hybrid formation mediates RNAi-directed heterochromatin formation. Genes Cells 17 : 218–233.
26. Castellano-PozoM, Santos-PereiraJM, RondonAG, BarrosoS, AndujarE, et al. (2013) R loops are linked to histone H3 S10 phosphorylation and chromatin condensation. Mol Cell 52 : 583–590.
27. YangY, McBrideKM, HensleyS, LuY, ChedinF, et al. (2014) Arginine methylation facilitates the recruitment of TOP3B to chromatin to prevent R loop accumulation. Mol Cell 53 : 484–497.
28. Santos-PereiraJM, HerreroAB, Garcia-RubioML, MarinA, MorenoS, et al. (2013) The Npl3 hnRNP prevents R-loop-mediated transcription-replication conflicts and genome instability. Genes Dev 27 : 2445–2458.
29. SordetO, RedonCE, Guirouilh-BarbatJ, SmithS, SolierS, et al. (2009) Ataxia telangiectasia mutated activation by transcription - and topoisomerase I-induced DNA double-strand breaks. EMBO Rep 10 : 887–893.
30. LiX, ManleyJL (2005) Inactivation of the SR protein splicing factor ASF/SF2 results in genomic instability. Cell 122 : 365–378.
31. StirlingPC, ChanYA, MinakerSW, AristizabalMJ, BarrettI, et al. (2012) R-loop-mediated genome instability in mRNA cleavage and polyadenylation mutants. Genes Dev 26 : 163–175.
32. HelmrichA, BallarinoM, ToraL (2011) Collisions between replication and transcription complexes cause common fragile site instability at the longest human genes. Mol Cell 44 : 966–977.
33. TuduriS, CrabbeL, ContiC, TourriereH, Holtgreve-GrezH, et al. (2009) Topoisomerase I suppresses genomic instability by preventing interference between replication and transcription. Nat Cell Biol 11 : 1315–1324.
34. WahbaL, AmonJD, KoshlandD, Vuica-RossM (2011) RNase H and multiple RNA biogenesis factors cooperate to prevent RNA∶DNA hybrids from generating genome instability. Mol Cell 44 : 978–988.
35. Herrera-MoyanoE, MerguiX, Garcia-RubioML, BarrosoS, AguileraA (2014) The yeast and human FACT chromatin-reorganizing complexes solve R-loop-mediated transcription-replication conflicts. Genes Dev 28 : 735–748.
36. CerritelliSM, CrouchRJ (2009) Ribonuclease H: the enzymes in eukaryotes. FEBS J 276 : 1494–1505.
37. BouleJB, ZakianVA (2007) The yeast Pif1p DNA helicase preferentially unwinds RNA DNA substrates. Nucleic Acids Res 35 : 5809–5818.
38. ShinJH, KelmanZ (2006) The replicative helicases of bacteria, archaea, and eukarya can unwind RNA-DNA hybrid substrates. J Biol Chem 281 : 26914–26921.
39. MoreiraMC, KlurS, WatanabeM, NemethAH, Le BerI, et al. (2004) Senataxin, the ortholog of a yeast RNA helicase, is mutant in ataxia-ocular apraxia 2. Nat Genet 36 : 225–227.
40. AnheimM, MongaB, FleuryM, CharlesP, BarbotC, et al. (2009) Ataxia with oculomotor apraxia type 2: clinical, biological and genotype/phenotype correlation study of a cohort of 90 patients. Brain 132 : 2688–2698.
41. ChenYZ, BennettCL, HuynhHM, BlairIP, PulsI, et al. (2004) DNA/RNA helicase gene mutations in a form of juvenile amyotrophic lateral sclerosis (ALS4). Am J Hum Genet 74 : 1128–1135.
42. SuraweeraA, LimY, WoodsR, BirrellGW, NasimT, et al. (2009) Functional role for senataxin, defective in ataxia oculomotor apraxia type 2, in transcriptional regulation. Hum Mol Genet 18 : 3384–3396.
43. SteinmetzEJ, WarrenCL, KuehnerJN, PanbehiB, AnsariAZ, et al. (2006) Genome-wide distribution of yeast RNA polymerase II and its control by Sen1 helicase. Mol Cell 24 : 735–746.
44. WestS, GromakN, ProudfootNJ (2004) Human 5′→3′ exonuclease Xrn2 promotes transcription termination at co-transcriptional cleavage sites. Nature 432 : 522–525.
45. SuraweeraA, BecherelOJ, ChenP, RundleN, WoodsR, et al. (2007) Senataxin, defective in ataxia oculomotor apraxia type 2, is involved in the defense against oxidative DNA damage. J Cell Biol 177 : 969–979.
46. YuceO, WestSC (2013) Senataxin, defective in the neurodegenerative disorder ataxia with oculomotor apraxia 2, lies at the interface of transcription and the DNA damage response. Mol Cell Biol 33 : 406–417.
47. AlzuA, BermejoR, BegnisM, LuccaC, PicciniD, et al. (2012) Senataxin associates with replication forks to protect fork integrity across RNA-polymerase-II-transcribed genes. Cell 151 : 835–846.
48. RichardP, FengS, ManleyJL (2013) A SUMO-dependent interaction between Senataxin and the exosome, disrupted in the neurodegenerative disease AOA2, targets the exosome to sites of transcription-induced DNA damage. Genes Dev 27 : 2227–2232.
49. VantaggiatoC, BondioniS, AiroldiG, BozzatoA, BorsaniG, et al. (2011) Senataxin modulates neurite growth through fibroblast growth factor 8 signalling. Brain 134 : 1808–1828.
50. PowellWT, CoulsonRL, GonzalesML, CraryFK, WongSS, et al. (2013) R-loop formation at Snord116 mediates topotecan inhibition of Ube3a-antisense and allele-specific chromatin decondensation. Proc Natl Acad Sci U S A 110 : 13938–13943.
51. HelmrichA, BallarinoM, NudlerE, ToraL (2013) Transcription-replication encounters, consequences and genomic instability. Nat Struct Mol Biol 20 : 412–418.
52. ChonH, SparksJL, RychlikM, NowotnyM, BurgersPM, et al. (2013) RNase H2 roles in genome integrity revealed by unlinking its activities. Nucleic Acids Res 41 : 3130–3143.
53. CerritelliSM, FrolovaEG, FengC, GrinbergA, LovePE, et al. (2003) Failure to produce mitochondrial DNA results in embryonic lethality in Rnaseh1 null mice. Mol Cell 11 : 807–815.
54. RydbergB, GameJ (2002) Excision of misincorporated ribonucleotides in DNA by RNase H (type 2) and FEN-1 in cell-free extracts. Proc Natl Acad Sci U S A 99 : 16654–16659.
55. Nick McElhinnySA, KumarD, ClarkAB, WattDL, WattsBE, et al. (2010) Genome instability due to ribonucleotide incorporation into DNA. Nat Chem Biol 6 : 774–781.
56. ReijnsMA, RabeB, RigbyRE, MillP, AstellKR, et al. (2012) Enzymatic removal of ribonucleotides from DNA is essential for mammalian genome integrity and development. Cell 149 : 1008–1022.
57. LazzaroF, NovarinaD, AmaraF, WattDL, StoneJE, et al. (2012) RNase H and postreplication repair protect cells from ribonucleotides incorporated in DNA. Mol Cell 45 : 99–110.
58. AranaME, KernsRT, WhareyL, GerrishKE, BushelPR, et al. (2012) Transcriptional responses to loss of RNase H2 in Saccharomyces cerevisiae. DNA Repair (Amst) 11 : 933–941.
59. CrowYJ, LeitchA, HaywardBE, GarnerA, ParmarR, et al. (2006) Mutations in genes encoding ribonuclease H2 subunits cause Aicardi-Goutieres syndrome and mimic congenital viral brain infection. Nat Genet 38 : 910–916.
60. CrowYJ, RehwinkelJ (2009) Aicardi-Goutieres syndrome and related phenotypes: linking nucleic acid metabolism with autoimmunity. Hum Mol Genet 18: R130–136.
61. CrowYJ, HaywardBE, ParmarR, RobinsP, LeitchA, et al. (2006) Mutations in the gene encoding the 3′-5′ DNA exonuclease TREX1 cause Aicardi-Goutieres syndrome at the AGS1 locus. Nat Genet 38 : 917–920.
62. RiceGI, KasherPR, ForteGM, MannionNM, GreenwoodSM, et al. (2012) Mutations in ADAR1 cause Aicardi-Goutieres syndrome associated with a type I interferon signature. Nat Genet 44 : 1243–1248.
63. RiceGI, BondJ, AsipuA, BrunetteRL, ManfieldIW, et al. (2009) Mutations involved in Aicardi-Goutieres syndrome implicate SAMHD1 as regulator of the innate immune response. Nat Genet 41 : 829–832.
64. Lee-KirschMA, WolfC, GuntherC (2014) Aicardi-Goutieres syndrome: a model disease for systemic autoimmunity. Clin Exp Immunol 175 : 17–24.
65. RabeB (2013) Aicardi-Goutieres syndrome: clues from the RNase H2 knock-out mouse. J Mol Med (Berl) 91 : 1235–1240.
66. HillerB, AchleitnerM, GlageS, NaumannR, BehrendtR, et al. (2012) Mammalian RNase H2 removes ribonucleotides from DNA to maintain genome integrity. J Exp Med 209 : 1419–1426.
67. BubeckD, ReijnsMA, GrahamSC, AstellKR, JonesEY, et al. (2011) PCNA directs type 2 RNase H activity on DNA replication and repair substrates. Nucleic Acids Res 39 : 3652–3666.
68. RigbyRE, WebbLM, MackenzieKJ, LiY, LeitchA, et al. (2014) RNA∶DNA hybrids are a novel molecular pattern sensed by TLR9. EMBO J 33 : 542–558.
69. Lopez CastelA, ClearyJD, PearsonCE (2010) Repeat instability as the basis for human diseases and as a potential target for therapy. Nat Rev Mol Cell Biol 11 : 165–170.
70. GrabczykE, MancusoM, SammarcoMC (2007) A persistent RNA.DNA hybrid formed by transcription of the Friedreich ataxia triplet repeat in live bacteria, and by T7 RNAP in vitro. Nucleic Acids Res 35 : 5351–5359.
71. LinY, DentSY, WilsonJH, WellsRD, NapieralaM (2010) R loops stimulate genetic instability of CTG.CAG repeats. Proc Natl Acad Sci U S A 107 : 692–697.
72. McIvorEI, PolakU, NapieralaM (2010) New insights into repeat instability: role of RNA*DNA hybrids. RNA Biol 7 : 551–558.
73. ReddyK, TamM, BowaterRP, BarberM, TomlinsonM, et al. (2011) Determinants of R-loop formation at convergent bidirectionally transcribed trinucleotide repeats. Nucleic Acids Res 39 : 1749–1762.
74. BelotserkovskiiBP, MirkinSM, HanawaltPC (2013) DNA sequences that interfere with transcription: implications for genome function and stability. Chem Rev 113 : 8620–8637.
75. GrohM, LufinoMM, Wade-MartinsR, GromakN (2014) R-loops associated with triplet repeat expansions promote gene silencing in Friedreich ataxia and fragile X syndrome. PLoS Genet 10: e1004318.
76. LoomisEW, SanzLA, ChedinF, HagermanPJ (2014) Transcription-Associated R-Loop Formation across the Human FMR1 CGG-Repeat Region. PLoS Genet 10: e1004294.
77. ColakD, ZaninovicN, CohenMS, RosenwaksZ, YangWY, et al. (2014) Promoter-bound trinucleotide repeat mRNA drives epigenetic silencing in fragile X syndrome. Science 343 : 1002–1005.
78. HaeuslerAR, DonnellyCJ, PerizG, SimkoEA, ShawPG, et al. (2014) C9orf72 nucleotide repeat structures initiate molecular cascades of disease. Nature 507 : 195–200.
79. HanahanD, WeinbergRA (2011) Hallmarks of cancer: the next generation. Cell 144 : 646–674.
80. NegriniS, GorgoulisVG, HalazonetisTD (2010) Genomic instability–an evolving hallmark of cancer. Nat Rev Mol Cell Biol 11 : 220–228.
81. ChernikovaSB, RazorenovaOV, HigginsJP, SishcBJ, NicolauM, et al. (2012) Deficiency in mammalian histone H2B ubiquitin ligase Bre1 (Rnf20/Rnf40) leads to replication stress and chromosomal instability. Cancer Res 72 : 2111–2119.
82. BhatiaV, BarrosoSI, Garcia-RubioML, TuminiE, Herrera-MoyanoE, et al. (2014) BRCA2 prevents R-loop accumulation and associates with TREX-2 mRNA export factor PCID2. Nature 511 : 362–365.
83. JacksonBR, NoerenbergM, WhitehouseA (2014) A novel mechanism inducing genome instability in Kaposi's sarcoma-associated herpesvirus infected cells. PLoS Pathog 10: e1004098.
84. SikdarN, BanerjeeS, ZhangH, SmithS, MyungK (2008) Spt2p defines a new transcription-dependent gross chromosomal rearrangement pathway. PLoS Genet 4: e1000290.
85. San FilippoJ, SungP, KleinH (2008) Mechanism of eukaryotic homologous recombination. Annu Rev Biochem 77 : 229–257.
86. ChaudhuriJ, TianM, KhuongC, ChuaK, PinaudE, et al. (2003) Transcription-targeted DNA deamination by the AID antibody diversification enzyme. Nature 422 : 726–730.
87. RobbianiDF, NussenzweigMC (2013) Chromosome translocation, B cell lymphoma, and activation-induced cytidine deaminase. Annu Rev Pathol 8 : 79–103.
88. RuizJF, Gomez-GonzalezB, AguileraA (2011) AID induces double-strand breaks at immunoglobulin switch regions and c-MYC causing chromosomal translocations in yeast THO mutants. PLoS Genet 7: e1002009.
89. DuquetteML, PhamP, GoodmanMF, MaizelsN (2005) AID binds to transcription-induced structures in c-MYC that map to regions associated with translocation and hypermutation. Oncogene 24 : 5791–5798.
90. BurnsMB, LackeyL, CarpenterMA, RathoreA, LandAM, et al. (2013) APOBEC3B is an enzymatic source of mutation in breast cancer. Nature 494 : 366–370.
91. KulisM, EstellerM (2010) DNA methylation and cancer. Adv Genet 70 : 27–56.
92. WongsurawatT, JenjaroenpunP, KwohCK, KuznetsovV (2011) Quantitative model of R-loop forming structures reveals a novel level of RNA-DNA interactome complexity. Nucleic Acids Res 40: e16.
93. HynesNE, StoelzleT (2009) Key signalling nodes in mammary gland development and cancer: Myc. Breast Cancer Res 11 : 210.
94. BalkB, MaicherA, DeesM, KlermundJ, Luke-GlaserS, et al. (2013) Telomeric RNA-DNA hybrids affect telomere-length dynamics and senescence. Nat Struct Mol Biol 20 : 1199–1205.
95. PfeifferV, CrittinJ, GrolimundL, LingnerJ (2013) The THO complex component Thp2 counteracts telomeric R-loops and telomere shortening. EMBO J 32 : 2861–2871.
96. De AmicisA, PianeM, FerrariF, FanciulliM, DeliaD, et al. (2011) Role of senataxin in DNA damage and telomeric stability. DNA Repair (Amst) 10 : 199–209.
97. WheelhouseRT, ChairesJB (2010) Drug binding to DNA×RNA hybrid structures. Methods Mol Biol 613 : 55–70.
98. ShawNN, AryaDP (2008) Recognition of the unique structure of DNA∶RNA hybrids. Biochimie 90 : 1026–1039.
99. ShawNN, XiH, AryaDP (2008) Molecular recognition of a DNA∶RNA hybrid: sub-nanomolar binding by a neomycin-methidium conjugate. Bioorg Med Chem Lett 18 : 4142–4145.
100. CassidySB, DykensE, WilliamsCA (2000) Prader-Willi and Angelman syndromes: sister imprinted disorders. Am J Med Genet 97 : 136–146.
101. MengL, PersonRE, BeaudetAL (2012) Ube3a-ATS is an atypical RNA polymerase II transcript that represses the paternal expression of Ube3a. Hum Mol Genet 21 : 3001–3012.
102. WhiteR, SaxtyB, LargeJ, KettleboroughCA, JacksonAP (2013) Identification of small-molecule inhibitors of the ribonuclease H2 enzyme. J Biomol Screen 18 : 610–620.
103. SipovaH, ZhangS, DudleyAM, GalasD, WangK, et al. (2010) Surface plasmon resonance biosensor for rapid label-free detection of microribonucleic acid at subfemtomole level. Anal Chem 82 : 10110–10115.
104. QaviAJ, KindtJT, GleesonMA, BaileyRC (2011) Anti-DNA∶RNA antibodies and silicon photonic microring resonators: increased sensitivity for multiplexed microRNA detection. Anal Chem 83 : 5949–5956.
105. Gomez-GonzalezB, Garcia-RubioM, BermejoR, GaillardH, ShirahigeK, et al. (2011) Genome-wide function of THO/TREX in active genes prevents R-loop-dependent replication obstacles. EMBO J 30 : 3106–3119.
Štítky
Genetika Reprodukční medicína
Článek An Evolutionarily Conserved Role for the Aryl Hydrocarbon Receptor in the Regulation of MovementČlánek Requirement for Drosophila SNMP1 for Rapid Activation and Termination of Pheromone-Induced ActivityČlánek Co-regulated Transcripts Associated to Cooperating eSNPs Define Bi-fan Motifs in Human Gene NetworksČlánek Identification of a Regulatory Variant That Binds FOXA1 and FOXA2 at the Type 2 Diabetes GWAS LocusČlánek tRNA Modifying Enzymes, NSUN2 and METTL1, Determine Sensitivity to 5-Fluorouracil in HeLa CellsČlánek Derlin-1 Regulates Mutant VCP-Linked Pathogenesis and Endoplasmic Reticulum Stress-Induced ApoptosisČlánek A Genetic Assay for Transcription Errors Reveals Multilayer Control of RNA Polymerase II FidelityČlánek The Proprotein Convertase KPC-1/Furin Controls Branching and Self-avoidance of Sensory Dendrites inČlánek Regulation of p53 and Rb Links the Alternative NF-κB Pathway to EZH2 Expression and Cell SenescenceČlánek BMPs Regulate Gene Expression in the Dorsal Neuroectoderm of and Vertebrates by Distinct MechanismsČlánek Unkempt Is Negatively Regulated by mTOR and Uncouples Neuronal Differentiation from Growth Control
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2014 Číslo 9
-
Všechny články tohoto čísla
- Translational Regulation of the Post-Translational Circadian Mechanism
- An Evolutionarily Conserved Role for the Aryl Hydrocarbon Receptor in the Regulation of Movement
- Eliminating Both Canonical and Short-Patch Mismatch Repair in Suggests a New Meiotic Recombination Model
- Requirement for Drosophila SNMP1 for Rapid Activation and Termination of Pheromone-Induced Activity
- Co-regulated Transcripts Associated to Cooperating eSNPs Define Bi-fan Motifs in Human Gene Networks
- Targeted H3R26 Deimination Specifically Facilitates Estrogen Receptor Binding by Modifying Nucleosome Structure
- Role for Circadian Clock Genes in Seasonal Timing: Testing the Bünning Hypothesis
- The Tandem Repeats Enabling Reversible Switching between the Two Phases of β-Lactamase Substrate Spectrum
- The Association of the Vanin-1 N131S Variant with Blood Pressure Is Mediated by Endoplasmic Reticulum-Associated Degradation and Loss of Function
- Identification of a Regulatory Variant That Binds FOXA1 and FOXA2 at the Type 2 Diabetes GWAS Locus
- Regulation of Flowering by the Histone Mark Readers MRG1/2 via Interaction with CONSTANS to Modulate Expression
- The Actomyosin Machinery Is Required for Retinal Lumen Formation
- Plays a Conserved Role in Assembly of the Ciliary Motile Apparatus
- Hidden Diversity in Honey Bee Gut Symbionts Detected by Single-Cell Genomics
- Ribosome Rescue and Translation Termination at Non-Standard Stop Codons by ICT1 in Mammalian Mitochondria
- tRNA Modifying Enzymes, NSUN2 and METTL1, Determine Sensitivity to 5-Fluorouracil in HeLa Cells
- Causal Variation in Yeast Sporulation Tends to Reside in a Pathway Bottleneck
- Tissue-Specific RNA Expression Marks Distant-Acting Developmental Enhancers
- WC-1 Recruits SWI/SNF to Remodel and Initiate a Circadian Cycle
- Clonal Expansion of Early to Mid-Life Mitochondrial DNA Point Mutations Drives Mitochondrial Dysfunction during Human Ageing
- Methylation QTLs Are Associated with Coordinated Changes in Transcription Factor Binding, Histone Modifications, and Gene Expression Levels
- Differential Management of the Replication Terminus Regions of the Two Chromosomes during Cell Division
- Obesity-Linked Homologues and Establish Meal Frequency in
- Derlin-1 Regulates Mutant VCP-Linked Pathogenesis and Endoplasmic Reticulum Stress-Induced Apoptosis
- Stress-Induced Nuclear RNA Degradation Pathways Regulate Yeast Bromodomain Factor 2 to Promote Cell Survival
- The MAPK p38c Regulates Oxidative Stress and Lipid Homeostasis in the Intestine
- Widespread Genome Reorganization of an Obligate Virus Mutualist
- Trans-kingdom Cross-Talk: Small RNAs on the Move
- The Vip1 Inositol Polyphosphate Kinase Family Regulates Polarized Growth and Modulates the Microtubule Cytoskeleton in Fungi
- Myosin Vb Mediated Plasma Membrane Homeostasis Regulates Peridermal Cell Size and Maintains Tissue Homeostasis in the Zebrafish Epidermis
- GLD-4-Mediated Translational Activation Regulates the Size of the Proliferative Germ Cell Pool in the Adult Germ Line
- Genome Wide Association Studies Using a New Nonparametric Model Reveal the Genetic Architecture of 17 Agronomic Traits in an Enlarged Maize Association Panel
- Translational Regulation of the DOUBLETIME/CKIδ/ε Kinase by LARK Contributes to Circadian Period Modulation
- Positive Selection and Multiple Losses of the LINE-1-Derived Gene in Mammals Suggest a Dual Role in Genome Defense and Pluripotency
- Out of Balance: R-loops in Human Disease
- A Genetic Assay for Transcription Errors Reveals Multilayer Control of RNA Polymerase II Fidelity
- Altered Behavioral Performance and Live Imaging of Circuit-Specific Neural Deficiencies in a Zebrafish Model for Psychomotor Retardation
- Nipbl and Mediator Cooperatively Regulate Gene Expression to Control Limb Development
- Meta-analysis of Mutations in Autism Spectrum Disorders: A Gradient of Severity in Cognitive Impairments
- The Proprotein Convertase KPC-1/Furin Controls Branching and Self-avoidance of Sensory Dendrites in
- Hydroxymethylated Cytosines Are Associated with Elevated C to G Transversion Rates
- Memory and Fitness Optimization of Bacteria under Fluctuating Environments
- Regulation of p53 and Rb Links the Alternative NF-κB Pathway to EZH2 Expression and Cell Senescence
- Interspecific Tests of Allelism Reveal the Evolutionary Timing and Pattern of Accumulation of Reproductive Isolation Mutations
- PRO40 Is a Scaffold Protein of the Cell Wall Integrity Pathway, Linking the MAP Kinase Module to the Upstream Activator Protein Kinase C
- Low Levels of p53 Protein and Chromatin Silencing of p53 Target Genes Repress Apoptosis in Endocycling Cells
- SPDEF Inhibits Prostate Carcinogenesis by Disrupting a Positive Feedback Loop in Regulation of the Foxm1 Oncogene
- RRP6L1 and RRP6L2 Function in Silencing Regulation of Antisense RNA Synthesis
- BMPs Regulate Gene Expression in the Dorsal Neuroectoderm of and Vertebrates by Distinct Mechanisms
- Unkempt Is Negatively Regulated by mTOR and Uncouples Neuronal Differentiation from Growth Control
- Atkinesin-13A Modulates Cell-Wall Synthesis and Cell Expansion in via the THESEUS1 Pathway
- Dopamine Signaling Leads to Loss of Polycomb Repression and Aberrant Gene Activation in Experimental Parkinsonism
- Histone Methyltransferase MMSET/NSD2 Alters EZH2 Binding and Reprograms the Myeloma Epigenome through Global and Focal Changes in H3K36 and H3K27 Methylation
- Bipartite Recognition of DNA by TCF/Pangolin Is Remarkably Flexible and Contributes to Transcriptional Responsiveness and Tissue Specificity of Wingless Signaling
- The Olfactory Transcriptomes of Mice
- Muscular Dystrophy-Associated and Variants Disrupt Nuclear-Cytoskeletal Connections and Myonuclear Organization
- Interplay of dFOXO and Two ETS-Family Transcription Factors Determines Lifespan in
- Evidence for Widespread Positive and Negative Selection in Coding and Conserved Noncoding Regions of
- Genome-Wide Association Meta-analysis of Neuropathologic Features of Alzheimer's Disease and Related Dementias
- Rejuvenation of Meiotic Cohesion in Oocytes during Prophase I Is Required for Chiasma Maintenance and Accurate Chromosome Segregation
- Admixture in Latin America: Geographic Structure, Phenotypic Diversity and Self-Perception of Ancestry Based on 7,342 Individuals
- Local Effect of Enhancer of Zeste-Like Reveals Cooperation of Epigenetic and -Acting Determinants for Zygotic Genome Rearrangements
- Differential Responses to Wnt and PCP Disruption Predict Expression and Developmental Function of Conserved and Novel Genes in a Cnidarian
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Admixture in Latin America: Geographic Structure, Phenotypic Diversity and Self-Perception of Ancestry Based on 7,342 Individuals
- Nipbl and Mediator Cooperatively Regulate Gene Expression to Control Limb Development
- Genome Wide Association Studies Using a New Nonparametric Model Reveal the Genetic Architecture of 17 Agronomic Traits in an Enlarged Maize Association Panel
- Histone Methyltransferase MMSET/NSD2 Alters EZH2 Binding and Reprograms the Myeloma Epigenome through Global and Focal Changes in H3K36 and H3K27 Methylation
Přihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



