-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaA Discrete Transition Zone Organizes the Topological and Regulatory Autonomy of the Adjacent and Genes
The specificity of enhancer-gene interactions is fundamental to the execution of gene regulatory programs underpinning embryonic development and cell differentiation. However, our understanding of the mechanisms conferring specificity to enhancers and target gene interactions is limited. In this study, we characterize the cis-regulatory organization of a large genomic locus consisting of two developmental genes, Tfap2c and Bmp7. We show that this locus is structurally partitioned into two distinct domains by the constitutive action of a discrete transition zone located between the two genes. This separation restricts selectively the functional action of enhancers to the genes present within the same domain. Interestingly, the effects of this region as a boundary are relative, as it allows some competing interactions to take place across domains. We show that these interactions modulate the functional output of a brain enhancer on its primary target gene resulting in the spatial restriction of its expression domain. These results support a functional link between topological chromatin domains and allocation of enhancers to genes. They further show that a precise adjustment of chromatin interaction levels fine-tunes gene regulation by long-range enhancers.
Published in the journal: . PLoS Genet 11(1): e32767. doi:10.1371/journal.pgen.1004897
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1004897Summary
The specificity of enhancer-gene interactions is fundamental to the execution of gene regulatory programs underpinning embryonic development and cell differentiation. However, our understanding of the mechanisms conferring specificity to enhancers and target gene interactions is limited. In this study, we characterize the cis-regulatory organization of a large genomic locus consisting of two developmental genes, Tfap2c and Bmp7. We show that this locus is structurally partitioned into two distinct domains by the constitutive action of a discrete transition zone located between the two genes. This separation restricts selectively the functional action of enhancers to the genes present within the same domain. Interestingly, the effects of this region as a boundary are relative, as it allows some competing interactions to take place across domains. We show that these interactions modulate the functional output of a brain enhancer on its primary target gene resulting in the spatial restriction of its expression domain. These results support a functional link between topological chromatin domains and allocation of enhancers to genes. They further show that a precise adjustment of chromatin interaction levels fine-tunes gene regulation by long-range enhancers.
Introduction
Differential regulation of gene expression transforms shared genomic information into the cell type-specific programs underlying organismal development and homeostasis. In vertebrates, it is not uncommon to find gene regulatory elements, in particular enhancers, hundreds of kilobases away from their target gene (reviewed in [1], [2]). The mere scale of this genomic distance raises the question of how enhancers and promoters can find each other, and how enhancers distinguish between their specific target and other neighboring genes, which may even lie much closer. Understanding the molecular basis of such specific interactions is essential as their impairment can lead to mis-expression of the normal target gene [3], [4] or to inappropriate activation of neighboring genes [5]–[8], with often severe phenotypic consequences [7], [9]–[12].
Enhancers can typically activate transcription from different promoters, a property that is part of their initial definition [13] and which has been amply used to assess enhancer activity. Many enhancers act pervasively across their endogenous genomic surroundings [14], [15], and enhancer sharing is not unusual between neighboring genes, particularly within multigenic clusters [16]–[22]. Noteworthy, this can also occur between genes with no functional relationship except genomic proximity [9], [23]–[25]. Nonetheless, in many loci, adjacent genes exhibit distinct expression patterns, implying the existence of mechanisms that limit the promiscuous potential of enhancers.
Different mechanisms and genomic elements have been invoked to explain enhancer-target gene specificity. They can be divided in two main categories, depending on whether they may promote interactions (eg. nature of the promoter, tethering elements [26], [27]), or block them. Amongst the latter, insulators prevent contact of an enhancer with an adjacent promoter, when placed in between [28]–[30]. This capacity of insulators to organize the genome in separate regulatory compartments designate them as critical components in ensuring specificity of cis-regulatory interactions [31]. However, only a handful of insulator elements have been functionally assessed in their native genomic context, and therefore their mode(s) of action is still poorly understood. Contrary to earlier models, a growing body of evidence suggests that insulators do not function autonomously, but rather through higher-order 3D conformations [32].
The necessity to consider the genome's three-dimensional organization is further highlighted by genome-wide high-resolution interaction maps obtained by chromosomal conformation capture techniques [33]. These studies revealed that the genome is compartmentalized in topologically-associating domains (TADs) [34], [35]. TADs have been proposed to contribute to gene expression by limiting enhancer action [36], [37]. In support of this view, genes located within the same TAD tend to be expressed coordinately [35], [38], and TADs have been found to encompass the regulatory domains defined by long-range enhancer activities [15], [39]. Recent works have addressed the finer-scale structural organization of TADs, revealing a complex hierarchy of interactions, which may contribute to mediate long-distance interactions between enhancers and promoters [40], [41] and to subdivide them into distinct regulatory domains [15]. In most instances, the functionality of structural contacts is difficult to evaluate precisely and the causal relationship between structural conformation and gene regulation remains unclear.
To better understand the relationships between 3D structural properties of the genome and enhance-promoter allocations, we focused on a large interval of approximately 0.5 Mb containing two different developmental genes, Bmp7 and Tfap2c. These two genes, which encode a secreted signaling molecule and a nuclear transcription factor, respectively, are active in multiple tissues and organs during embryogenesis [42]–[48]. Both genes have promoter architectures compatible with tissue-specific and long-distance regulatory inputs [49]. Their expression overlaps in the limbs, forebrain and branchial arches of mid-gestation mouse embryos, while in other contexts, their expression is specific of one or the other and exclusive. Therefore this locus constitutes an ideal system to study the control of long-distance enhancer specificities.
To investigate the regulatory organization of this locus, we used a transposon/recombination-based chromosomal engineering approach [14]. We show here that the genomic interval consists of two largely independent regulatory domains, corresponding to each of the two genes. Analysis of the chromatin conformation of re-engineered genomic configurations identified a central transition zone (TZ) that defines different topological sub-domains. Importantly, the allocation of enhancers to one or the other gene is determined by this partition. Altogether, our data support the view that the topological organization of the genome restricts enhancers to specific domains, determining therefore their “specific” target gene choice. Interestingly, we found that the presence of Bmp7 in cis has a mild influence on the expression level of Tfap2c in the developing forebrain, indicating that the position of the two genes to different topological domains does not lead to an absolute insulation.
Results
Mapping the regulatory landscapes of the Tfap2c-Bmp7 locus
To determine the regulatory organization of the Tfap2c-Bmp7 locus, we adapted the GROMIT (Genome Regulatory Organization Mapping with Integrated Transposons) strategy [14]. Firstly, at the 3′ end of the endogenous Bmp7 gene, we inserted a transgene consisting of a Sleeping Beauty transposon comprising 1) a regulatory sensor gene (a LacZ reporter under the control of a short naïve synthetic promoter region derived from the human β-globin gene [14], [50]) and 2) a loxP site. After establishment of a mouse line carrying the correct insertion, we removed the selection marker used to identify candidate targeted ES clones, a step which left behind an additional loxP site, next to the Sleeping Beauty transposon. We designated this allele as SB-B(3end) (Fig. 1). By serial remobilisation of the transposon in vivo [14], we obtained several insertions located in this region of mouse chromosome 2 (S1 Table). Of these, seven insertions were distributed along the Tfap2c-Bmp7 locus (Fig. 1A): three very close to SB-B(3end) (within 23 kb), one (SB-B(up)) 20 kb upstream of Bmp7, and another one in the first intron of Bmp7 (SB-B(in)). The remaining two (SB-A1 and SB-A2) lie within the large intergenic region separating Tfap2c and Bmp7. In parallel, we established a mouse line (BA0758) from an ES clone carrying a βgeo gene-trap insertion in Tfap2c [51].
Fig. 1. The Tfap2c-Bmp7 locus consists of two regulatory domains. 
(A) A schematic representation of the Tfap2c-Bmp7 locus, including the position of the SBlac insertions. Gene bodies are depicted as grey boxes with darker bars representing their exons. Tfap2c and Bmp7 are blue and green, respectively. The centromeric (CEN) and telomeric (TEL) ends of the chromosome are to the left and right of the diagram, respectively. The Sleeping Beauty transposon carrying a loxP site and LacZ reporter was first targeted into the immediate downstream region of Bmp7 along with an additional loxP sequence. Integration sites obtained upon remobilisation of the transposon are indicated by black arrowheads. (B) LacZ staining patterns of the transposon lines in the heart, limb, forebrain and the jaw as well as the staining of the BA0758 gene trap line in the forebrain and jaw are shown. Limbs: E12.5 embryos; other tissues: E11.5 embryos. Note that the intensity of the LacZ staining in the lateral and medial parts of the forebrain varies among BA0758, SB-A1 and SB-A2, as indicated by the blue arrows. Additional stages and views available in S1 Fig. The Tfap2c-Bmp7 locus consists of two distinct regulatory domains
We analyzed the expression pattern of the regulatory sensor at different insertion sites in E10.5 to E12.5 mouse embryos, at stages when Tfap2c and Bmp7 show both shared and specific expression patterns (Fig. 1, S1 Fig.). The two insertions located between Tfap2c and Bmp7 (SB-A1 and -A2) showed very similar LacZ staining in the oro-facial region, the branchial arches, and in the forebrain (Fig. 1B, left). These three expression domains are strikingly consistent with reported expression patterns of Tfap2c [42] and particularly with the Tfap2c LacZ gene-trap allele (Fig. 1B - S1 Fig.). This overlap and agreement in expression suggested that SB-A1 and -A2 were included in the Tfap2c “regulatory domain” [15]. The expression of the reporters showed however different relative intensity between the lateral and medial part of the forebrain: while BA0758 and SB-A1 were preferably expressed in the lateral forebrain, with weaker expression in the medial region, SB-A2 showed the inverse pattern, with a stronger medial than lateral LacZ staining. Such position-effects (the promoter is the same for SB-A2 and SB-A2) are not uncommon within regulatory domains [15], [52]. They may reflect the presence in the locus of several forebrain enhancers with distinct medial/lateral activity and different range of action.
These forebrain expression domains were not observed with any of the four insertions located within the 23-kb region at the 3′end of Bmp7 (Fig. 1B, S1 Fig.), suggesting that the telomeric limit of Tfap2c regulatory domain is upstream of this region. More distant insertions in Bmp7 (SB-B(in); SB-B(up)) showed weak medial-only forebrain expression at E11.5, with no lateral expression detected, as also observed for Bmp7 [44]. None of the six telomeric insertions showed the characteristic oro-facial expression observed with the Tfap2c-associated insertions. In contrast, they shared several common expression domains not reported by the SB-A1 and –A2 insertions (Fig. 1B). The four insertions at the 3′end of Bmp7 and SB-B(in) showed all prominent staining for LacZ expression in the developing heart (from E10.5 to E12.5), and in the interdigital mesenchyme (at E12.5). SB-B(up) displayed only faint LacZ staining in the interdigital mesenchyme, and no staining in the heart. However, LacZ expression from this position overlapped characteristically with other SB-B insertions in the whiskers, nasal pits, and forebrain (S1 Fig.), defining collectively a regulatory domain distinct from the one associated with Tfap2c. This domain includes Bmp7, and accordingly, several of the reported activities overlap with known Bmp7 expression domains [47], [53]. Some regions of the Bmp7 expression domain were not reflected accurately in the activity of the SB reporters, being either missing or spatially expanded. These differences may arise from the limited range of action of some promoter-proximal enhancers [53], and/or from the different post-transcriptional stability and dynamics of LacZ compared to the endogenous Bmp7 transcripts.
Overall, the regulatory activities detected by the sensor differed significantly between the centromeric and telomeric part of the locus, and highlighted two distinct and non-overlapping regulatory domains, each defined by multiple distinct tissue-specific activities, one domain corresponding to Tfap2c and the other to Bmp7. We focused for subsequent analyses on the forebrain (medial and lateral) and heart, as representative markers of these two domains. In these two tissues, the expression pattern of the different genes is stable from E10.5 and E12, contrasting with the dynamic expression of these genes in the developing limbs and face. Also, for these two expression domains, it is technically possible to dissect from embryos the part where the gene or the enhancer is active, without the contribution of too many non-expressing cells.
Enhancers in the intergenic region control either Bmp7 or Tfap2c
To further characterize the functional relevance of these two domains and associated enhancers, we used in vivo Cre-mediated recombination to engineer chromosomal deletions removing either the telomeric half or the whole of the intergenic region (Fig. 2). Each deletion was produced using a combination of loxP sites in cis and trans [54] in order to keep the LacZ sensor at the deletion breakpoint (see Materials and Methods). With the TAMERE strategy, we also obtained a large duplication, reciprocal to del3 (S2 Fig.). All three deletions led to a complete loss of LacZ expression in the embryonic heart and forebrain (Fig. 2B) suggesting that the enhancers detected by SB-A1 and SB-B(3end) lie in the region encompassed by del1. Dup3-lacZ embryos showed LacZ expression in the heart similar to SB-B(3end), corroborating the presence of the heart enhancer(s) at the 3′ side of Bmp7 (S2 Fig.). These deletions also provided information on the locations of additional enhancers associated with other expression domains (S2 Fig.).
Fig. 2. Deletion alleles localise enhancers. 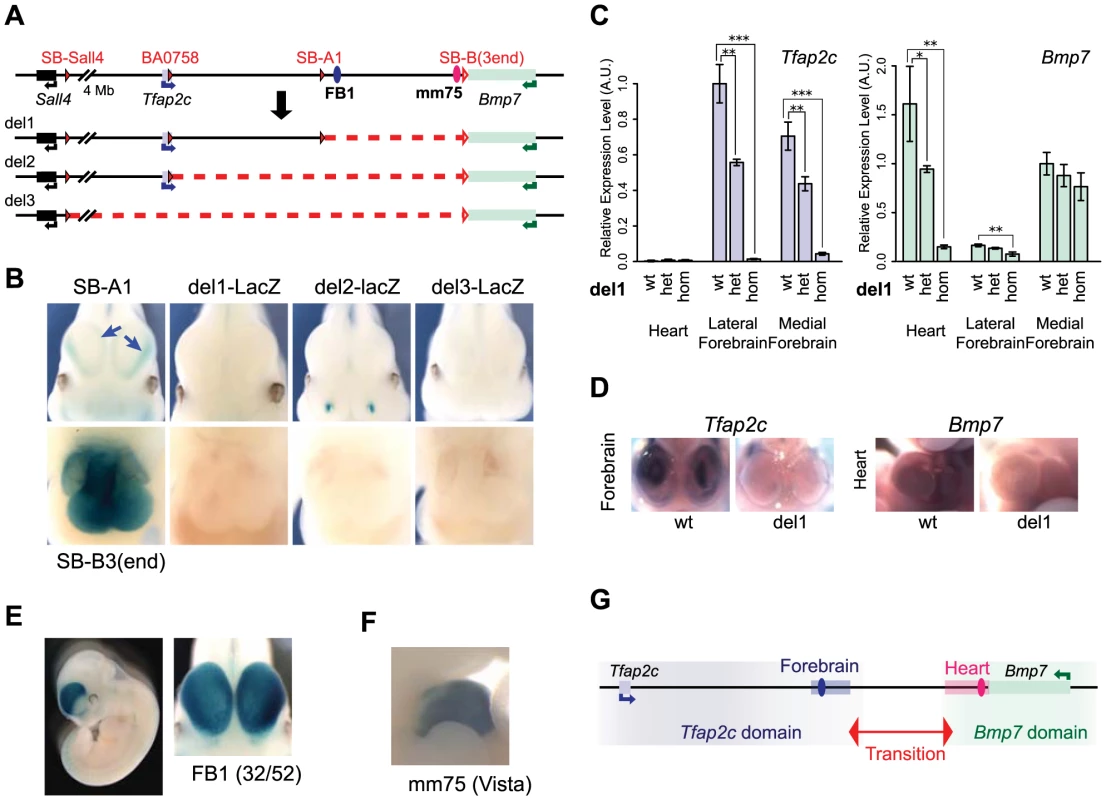
(A) A schematic representation of the deletions generated. The loxP sites used for CRE-mediated recombination are depicted as filled red triangles for the one carried with the transposon. Open triangles indicate both the position of the static loxP at the 3′end of Bmp7 and the position of the LacZ reporter gene in deletions produced by TAMERE. (B) LacZ staining patterns of the three deletion lines in forebrain (top) and heart (bottom) in E11.5 embryos, in comparison to SB-A1 and SB-B3(3end), respectively. The deletions led to complete loss of the both lateral and medial expression in the forebrain (blue arrows in the SB-A1 embryo). (C) Relative expression levels of the Tfap2c and Bmp7 mRNAs in the heart, lateral and medial forebrains from E11.5 embryos measured by RT-qPCR in del1 homozygous, heterozygous, and control (wt) genotypes. For each gene, expression levels were normalized with Gapdh. Expression of the wild type allele in the lateral forebrain for Tfap2c and in the medial forebrain for Bmp7, respectively, was set as 1. The error bars represent s.d. from three biological replicates. Statistical significance was assessed by a two-sided t-test. *p<0.05; **p<0.01; ***p<0.001. (D) In situ hybridization of the wild type and del1 embryos at E10.5 with the anti-sense RNA probes for Tfap2c and Bmp7. (E) Enhancer activity of FB1 on the LacZ reporter gene. 32 out of 52 transgenic embryos showed broad forebrain expression. (F) The mm75 element drives specific expression in the mouse embryos at E11.5 (from VISTA enhancer browser: http://enhancer.lbl.gov). (G). Regulatory domains along the Tfap2c-Bmp7 interval. The forebrain enhancer (FB1) and the heart enhancer (mm75) are depicted with blue and pink ovals. A light blue (resp. pink) rectangle represents the region encompassing the H3K27ac peaks present in the segment deleted in del1, in the forebrain (resp. heart) (S3 Fig.). We next determined if the enhancers present in the del1 interval contributed to Tfap2c and Bmp7 expression by whole-mount in situ hybridization and RT-qPCR (Fig. 2C–D). In del1 homozygous embryos, Bmp7 expression was drastically reduced in the heart compared to wild-type littermates, while the very weak expression of Tfap2c in the heart was unaffected (Fig. 2C). In the forebrain, where both genes are expressed, we found an almost complete loss of Tfap2c expression in both the medial and lateral parts of del1 embryos. In contrast, Bmp7 expression was barely affected and showed only a slight reduction in the lateral forebrain (Fig. 2C).
These analyses demonstrated a critical role of elements located within the del1 segment for the specific expression of Tfap2c in the forebrain and of Bmp7 in the heart, respectively. Several peaks enriched for chromatin marks associated with active enhancers (H3K27ac, EP300) have been detected within this region in the forebrain and the heart of E11.5 embryos [55]–[57] (S3 Fig.). Interestingly, the distribution of these regions is coincident with the location of the two regulatory domains. Many forebrain H3K27ac peaks are located between Tfap2c and SB-A1/A2, while the only ones present around Bmp7 lie in the first intron of the gene. Conversely, heart H3K27ac-enriched elements cluster around the 3′ end of Bmp7. H3K27ac peaks were also identified outside of the del1 region around the locus. The forebrain H3K27ac peak adjacent to Bmp7 could account for its unaffected expression in del1; however, the role of the predicted forebrain and heart enhancers located respectively centromeric and telomeric to del1, respectively, remained unclear, as they were seemingly unable to confer significant activity to the reporter gene or to the endogenous genes in these tissues, in the absence of del1 sequences.
To confirm that del1 contained enhancers with the expected activities, we cloned FB1, an evolutionarily conserved element enriched for both H3K27ac and EP300 in the forebrain, upstream of the regulatory sensor construct. In this transgenic assay, FB1 drove specific and reproducible LacZ expression in the forebrain in E11–12 embryos (Fig. 2E), including the Tfap2c expression domain. However, FB1 appeared broadly and equally active in both medial and lateral forebrain, contrasting with the restricted expression detected by the same reporter gene than the one used in the transgenic assay when inserted in the endogenous locus on either side of FB1. In this context, it showed alternatively preferential expression in the lateral (SB-A1, like Tfap2c) or medial (SB-A2). These differences suggested that additional factors – possibly the other H3K27ac-region present in the vicinity (see below) – may modulate FB1 intrinsic activity in a position-dependent manner. Amongst the predicted heart enhancers, one of them (mm75) had been tested previously [58] and reported to have broad enhancer activity in the heart of E11.5 mouse embryos (Fig. 2F).
Taken together, these data demonstrated that the del1 region contained heart-specific and forebrain-specific regulatory element(s) critical for the expression of Bmp7 in the heart, and of Tfap2c in the forebrain, respectively. Importantly, these elements appeared to be dispensable for the regulation of one another's genes. These selective influences and the separate location of the different enhancers further confirmed the partition of this genomic interval into two distinct regulatory domains containing enhancers which act exclusively on one or the other gene (Fig. 2G).
Topological organisation of the locus
We next investigated how the regulatory subdivision of the locus corresponded to its topological organization. Hi-C data available for mouse ES cells and cortex [34] suggests that the locus has a relatively loose topological structure, confined between two prominent topologically associating domains (Fig. 3A, S4 Fig.). To determine the pattern of physical contacts involving Tfap2c and Bmp7, we carried out circular chromatin conformation capture experiments followed by high-throughput sequencing (4C-Seq) using the promoters of these two genes as viewpoints (Fig. 3). We performed these 4C-Seq analyses on dissected samples where one and/or the other gene were expressed (E11.5 heart, medial and lateral forebrain) and whole body of E11.5 embryos (where most cells are non-expressing either of the two genes). We also included samples from E12.5 limbs, which comprised a majority of non-expressing cells.
Fig. 3. 4C profiles describing the conformational structure of the Tfap2c-Bmp7 locus. 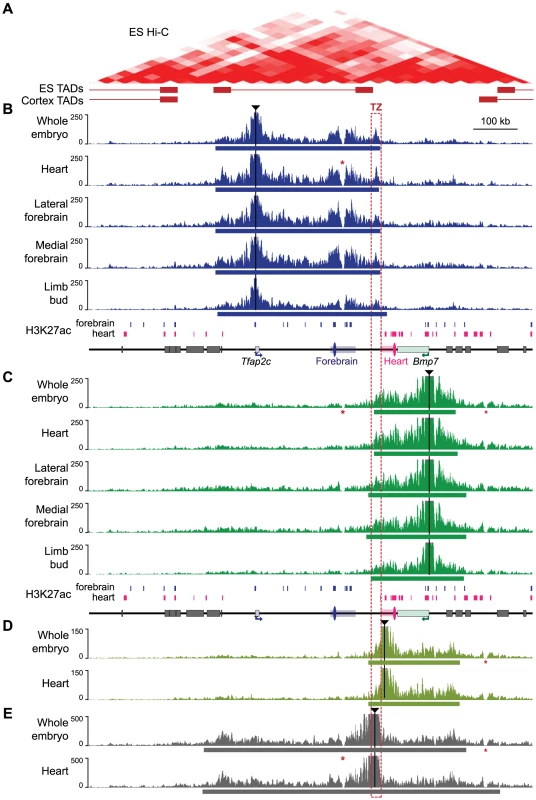
(A) Hi-C heat-map of the Tfap2c-Bmp7 locus in mouse ES cells (top) and corresponding TADs identified in ES cells and adult cortex (bottom; shown by whiskered red bars) (data from Dixon et al. 2012; aligned with the other panels). (B–E) 4C-contact profiles for the different viewpoints (indicated by black triangles): promoter of Tfap2c (B) and Bmp7 (C), adjacent to the TZ (D) and within the TZ (E). Five different tissues (whole embryos, heart, lateral forebrain, medial forebrain at stage E11.5, limb buds at E12.5) were examined with the two promoter viewpoints (B, C). Only whole embryos and heart samples were used for the additional viewpoints (D, E). The estimated primary interaction domains are indicated by a bar below the corresponding 4C plots. The region of overlap of the different primary interaction domains (TZ) is outlined with a dashed red box. Stars (*) indicate two regions with low mappability [84] accounting for the absence of signal over these positions. For comparison, a schematic representation of the region including H3K27ac peaks detected in forebrain and heart chromatin (S3 Fig.) is shown below panels B and C. Surrounding gene bodies are represented with grey boxes. The FB1 and mm75 enhancers are shown as blue and pink ovals, respectively. Associated rectangles indicate the extended enhancer regions encompassing the additional H3K27ac regions detected within the del1 segment. For both viewpoints, the 4C profiles highlighted a large primary interaction domain characterized by high 4C read counts (Fig. 3B, C). We applied a segmentation algorithm [59] to delineate this primary domain in the different conditions (S2 Table). The calculated primary interaction domains for a given viewpoint were nearly identical across the different tissue samples. The 4C profiles were predominantly similar between samples, with the exception of a moderate increase of the 4C signals over the enhancers associated with each gene in the tissues in which they are active (for Tfap2c: FB1 and flanking H3K27ac-enriched regions in the brain samples; for Bmp7 mm75 and surrounding H3K27ac-enriched regions in the heart sample). We confirmed the increased interactions of Tfap2c with FB1 and of Bmp7 with mm75 in an independent 3C experiment (S5 Fig.). Importantly, the reciprocal 3C experiment with FB1 as a viewpoint showed that it contacted strongly Tfap2c in the forebrain, but not in the heart, and had much weaker/rarer contacts with Bmp7.
Noteworthy, the Tfap2c domain and the Bmp7 domain end shortly before the edges of the flanking TADs detected in mouse ES cells [34], consistent with the notion that these 4C primary domains corresponded to the structural conformation adopted by the locus. In all samples, the primary contact domains of one gene included the enhancer regions we found associated with it, but excluded the ones associated with the other gene. Nevertheless, we observed a consistent overlap between the two domains, demarcating a region of about 10 - to 30-kb region, which we termed the transition zone (TZ). To further characterize this region, we used two additional viewpoints for 4C analysis (Fig. 3D–E). Contacts observed from a viewpoint located just before the centromeric end of the Bmp7 primary domain showed extensive overlap with the latter, extending broadly over Bmp7 but not stopping almost abruptly at the TZ (Fig. 3D). Similarly, FB1 showed only weak contact with positions located on the other side of the TZ (S5C Fig.). This asymmetry in the distribution of contacts suggested the TZ indeed corresponds to a conformational transition between two different conformations. Importantly, a viewpoint located in the TZ itself showed prominent contacts extending towards both genes (Fig. 3E), consistent with the strong 4C signals observed over the TZ in the reciprocal 4C experiments.
Next, we performed 4C analyses on del1 homozygous embryos, where the TZ region was deleted together with a larger part of the locus, including the different enhancers (S6 Fig.). In this context, we observed a wide extension of the contacts made by Tfap2c (resp. Bmp7) in the telomeric (resp. centromeric) region, over distances larger than the size of the deleted region. At the same time, the centromeric (resp. telomeric) profiles remained highly similar between WT and del1. Interestingly, the intervals with frequent contacts by Tfap2c and Bmp7 now largely overlapped, as if they “merged” into one domain only limited by the adjacent TADs (S6 Fig., S3 Table). These new extended contacts supported the notion that the TZ may contribute to delineate two distinct structural domains. However, as del1 also significantly reduced the linear distance between Tfap2c and Bmp7, we decided to use other types of alleles to challenge the structural and regulatory organization of the locus and to test the influence of the TZ on these.
Chromosomal rearrangements led to enhancer re-allocation
We used insertions carrying loxP sites in the opposite orientation to the one left at the SB-B(3end) position in cis to engineer three balanced inversions by CRE-mediated recombination (Fig. 4A, S1 Table). In INV-L1 and -L2, the distance between Bmp7 and the heart enhancer increased to 5.7 and 1.1 Mb, respectively, whereas the relative order and distances between Tfap2c, the enhancers and the TZ region were unchanged (S7 Fig.). In INV-M, the heart enhancer was now equidistant from Bmp7 and Tfap2c (187 and 207 kb, compared to distances of 80 kb and 312 kb in the wild-type allele, with mm75 taken as reference). However, in this allele, the TZ was now located between Bmp7 and the heart enhancer(s). With each inversion, the LacZ reporter remained adjacent to the heart enhancer region and displayed its normal heart expression (Fig. 4B, S7 Fig.), demonstrating that these rearrangements did not disrupt heart enhancer activity. In the three inversions, Bmp7 expression was strongly reduced in the heart, comparable to levels observed with del1 (Fig. 4C). In contrast, Tfap2c expression was enhanced by a thousand-fold in the heart of INV-M animals (Fig. 4D), implying that in this genomic configuration, the heart enhancers now activated Tfap2c instead of Bmp7. This complete switch of the heart enhancer(s) from Bmp7 to Tfap2c coincided with the new relative position of the TZ. The importance of the position of the TZ was further supported by a lack of up-regulation of Tfap2c in INV-L1 and INV-L2 (Fig. 4D), where its location with regards to the TZ/heart enhancers remained unchanged. In INV-L1, we instead found an up-regulation of Ptgis (Fig. 4E), which was now located on the other side of the TZ, next to mm75. As Ptgis was closer to the heart enhancer (S7A Fig.) we were unable in this case to fully rule out a possible influence of distance on promoter choice. However, in INV-L2, Dok5, the new gene juxtaposed “next to” the heart enhancer(s) opposite to TZ was much further away than Tfap2c (1.1 Mb versus 0.3 Mb). In this context, neither Dok5 (Fig. 4F) nor Tfap2c were up-regulated in the heart, ruling out the possibility that the heart enhancer(s) act simply by default the nearest gene.
Fig. 4. Inversion alleles reallocate the target of the heart enhancer. 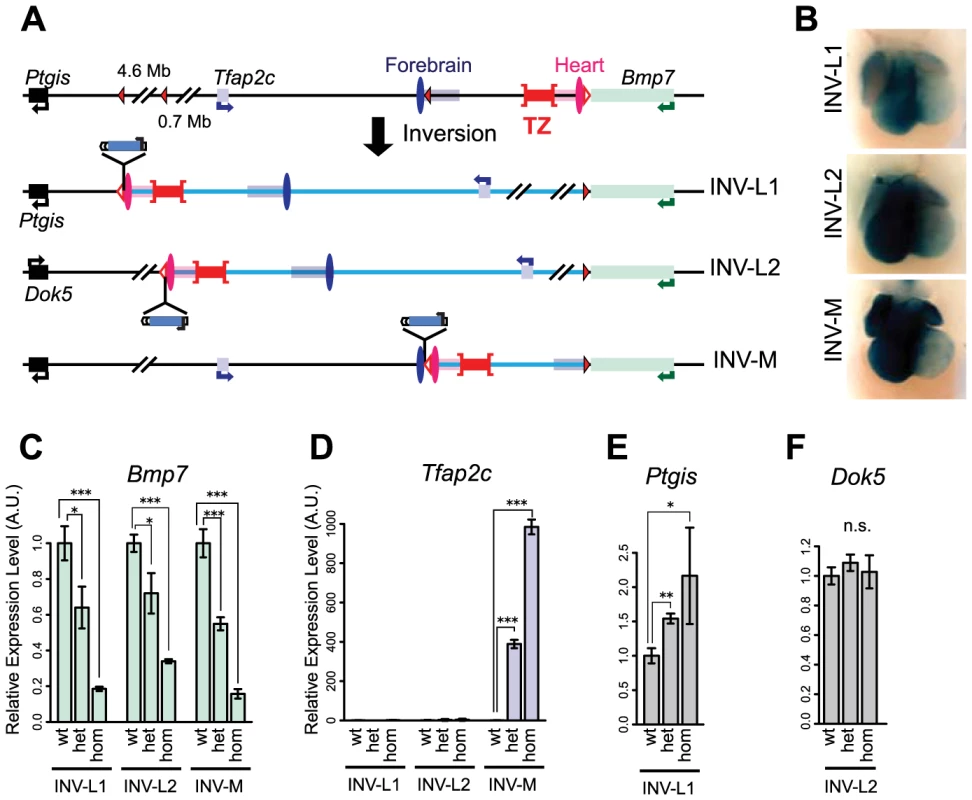
(A) A schematic representation of the three different inversions obtained in this study, with loxP sites as triangles, genes as plain boxes, and enhancers as ovals (FB1 or mm75) or grouped in rectangles for the ones predicted by chromatin marks (FB1/forebrain H3K27ac: blue, mm75/heart H3K27ac: pink). The TZ is represented by a whiskered red bar. In the generated inversion lines, the heart enhancer(s) were brought next to the LacZ reporter. (B) LacZ staining of the three inversion lines in E11.5 heart. Quantification by mRNA RT-qPCR of expression levels of Tfap2c (C), Bmp7 (D), Ptgis (E) and Dok5 (F) in the inversion alleles. Expression levels in wild type (wt) were normalized as 1. The error bars represent the s.d. of three biological replicates. The statistical significance was assessed by a two-sided Student's t-test. *p<0.05; **p<0.01; ***p<0.001; n.s.: non-significant. Intrinsic asymmetric distribution of interactions around the TZ
To examine at the consequences of these rearrangements on the structural conformation of the region, we performed 4C experiments on INV-M and INV-L2 embryos (Fig. 5, S8–S10 Figs.). In INV-M, as in WT controls, Tfap2c showed robust interactions over a domain extending up to the TZ. Due to the inversion, this domain now included the heart enhancer, which displayed much stronger interaction with Tfap2c than those observed in WT (S8A Fig., pink versus grey arrow), a result consistent with mm75 now activating Tfap2c. Conversely, the new primary interaction domain of Bmp7 stopped at the TZ, with a very reduced 4C signal over the heart enhancer in INV-M when compared to WT (S8D Fig., grey versus pink arrow). The viewpoint located between mm75 and TZ, which was part of the Bmp7 interaction domain in WT, showed in INV-M broad and extended contacts overlapping with the Tfap2c interaction domain, ending at the TZ region (Fig. 5B). Interestingly, the inversion had no effect on the 4C profile of the TZ-associated viewpoint, which extended on both sides in all configurations. Thus, in INV-M as in WT, the locus appeared structurally partitioned at the TZ: instead of maintaining their normal contacts and regulatory preferences, genes and regulatory elements established new interactions, depending on their respective position in relation to the TZ.
Fig. 5. Redistribution of the interaction domains upon chromosomal inversions. 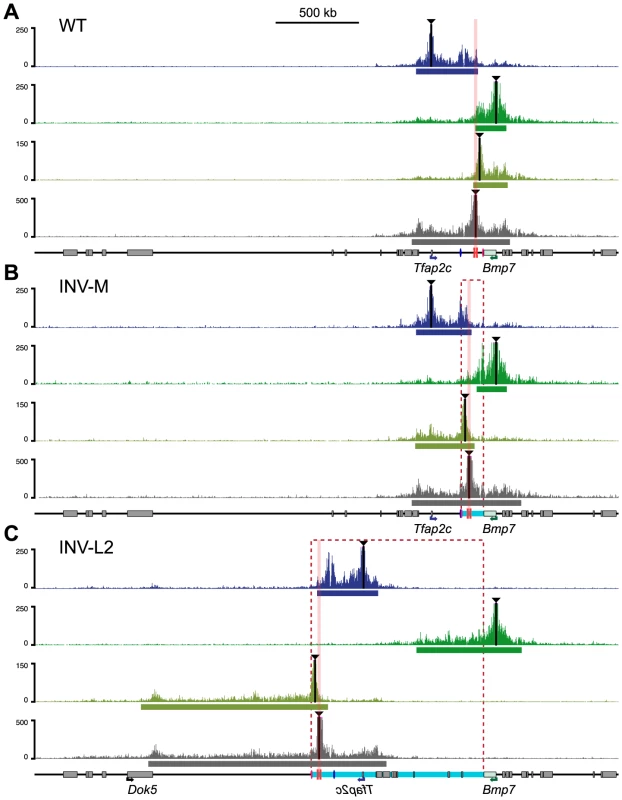
4C profiles were compared amongst WT control (A), INV-M (B) and INV-L2 (C) alleles for the four viewpoints indicated with black triangles. For inversion plots, the genomic coordinates were reordered to take the genomic rearrangements into account: hence, representated profiles correspond to the actual genomic structure of each allele. Representations of the data aligned on the reference (WT) genome are available in S8 and S9 Figs. Dashed rectangles and light-blue bars represent the regions inverted in the INV-M and INV-L2 alleles. The position of the TZ is marked by pink columns. The heart (mm75) and forebrain (FB1) enhancers are depicted as pink and blue ovals, respectively. The bars below each plot represent the corresponding primary interaction domain. In INV-L2 embryos, the 4C profile of Tfap2c appeared generally unchanged and did not expand across the TZ into its new flanking region. The TZ-flanking viewpoint remained still limited by the TZ, but highlighted on the other side a broad domain of nearly 1 Mb in the Dok5-Cbln4 gene desert, which is now adjacent to it. The 4C signal was strongly diminished before reaching the promoter of Dok5, which may explain the lack of up-regulation of this gene in the heart of INV-L2 embryos (Fig. 4F). Again, the TZ itself contacted both flanking regions, the relocated Tfap2c domain, and the new Dok5-Cbln4. Importantly, in INV-L2, Bmp7 showed broad contacts over the region now present at its 3′end, extending for up to 0.5 Mb further in the Cbln4 locus, supporting the notion that the presence of the TZ limited the extent of the Bmp7 contact range (Fig. 5C, S3 Table).
Remarkably, the new distribution of 4C contacts in the different rearrangements appeared to follow quite strictly the relative position of the TZ. It did not appear to depend on the nature of the flanking sequences themselves. The directional bias of contacts made by the viewpoint flanking the TZ is the same in the different configurations (WT, INV-M and INV-L2) (S10 Fig., on the right), irrespectively of the flanking sequences.
Fine-tuning of Tfap2c forebrain expression across the TZ
The expression and structural changes observed in the heart suggested that the TZ behaved as a simple insulator region. In INV-L1 and INV-L2, the Tfap2c domain was fully maintained and unaffected by the genomic rearrangements. Therefore one would expect little impact on Tfap2c. However, we observed an up-regulation of Tfap2c in the medial telencephalon in both alleles (Fig. 6A–B). This up-regulation is unlikely to be caused by the juxtaposition of new forebrain enhancers, as the regulatory sensor did not detect any forebrain activity in L1 and L2 position, in either the inverted or non-inverted configurations (Fig. 6C). We noted that in INV-L1 and –L2, Bmp7, which is strongly expressed in the medial forebrain, was relocated away from Tfap2c and its forebrain enhancer. This rearrangement had no effect on Bmp7 expression in the forebrain, suggesting that it was the presence of Bmp7 in cis that negatively influenced Tfap2c. Supporting this hypothesis, we did not observe any up-regulation of Tfap2c in the medial forebrain of INV-M embryos (Fig. 6D), where Bmp7 remained adjacent to the Tfap2c. These observations prompted us to re-examine the 4C profiles. As stated before, the intensity of the 4C signals diminished strongly beyond the TZ region. However, we observed that the 4C contacts made by the Bmp7 promoter, albeit weak, were stronger over the Tfap2c domain than over the region located symmetrically from the viewpoint (S9 Fig., green boxes). Reciprocally, Tfap2c showed weak but consistent interactions with the Bmp7 region in WT and INV-M (S9 Fig., blue boxes), interactions which are not observed with a symmetrically located region, or with the region at the equivalent place in INV-L2. To further test if the INV-L1 and –L2 up-regulation of Tfap2c depended on the removal of Bmp7, we produced INV-Bmp7 which consists in a simple inversion of the gene itself. Consequently, Bmp7 remained adjacent to the Tfap2c domain, and separated from it by the TZ (S11A Fig.). In this configuration, we did not observe significant changes of Bmp7 or Tfap2c expression, with the exception of a small reduction of Bmp7 expression in the lateral forebrain. Altogether, these results supported that the simple presence of an active Bmp7 in cis, despite the presence of the TZ region, can affect Tfap2c expression in the medial forebrain.
Fig. 6. Changes of gene expression in the forebrain following genomic inversion. 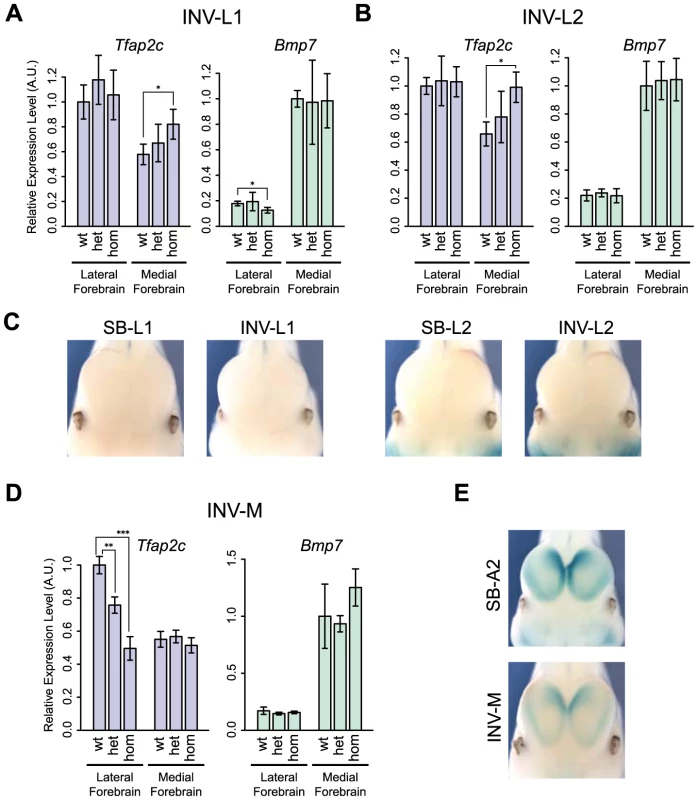
Quantification by RT-qPCR of the relative expression levels of the Tfap2c and Bmp7 mRNAs in the lateral and medial forebrain for INV-L1 (A), INV-L2 (B) and INV-M (D), normalized as in Fig. 2. Error bars represent the s.d. of three biological replicates. The statistical significance was assessed by a two-sided Student's t-test. *p<0.05; **p<0.01; ***p<0.001. (C) Absence of LacZ staining in the forebrain of SB/INV-L1 and SB/INV-L2 E11.5 embryos. (E) LacZ staining of SB-A2 (up) and INV-M (bottom) E11.5 embryos. We also noted that INV-M led to a significant reduction of Tfap2c expression in the lateral forebrain (Fig. 6D), even if the genomic region between Tfap2c and FB1 was unaffected. This reduction could result from the relocation to the other side of the TZ of two forebrain-specific H3K27ac-enriched regions included in INV-M. As we observed neither a concomitant up-regulation of Bmp7 (Fig. 6D) nor changes in the activity reported by the sensor (Fig. 6E), it is possible that these elements may not act autonomously but rather modulate the long-range action of FB1.
Discussion
We show here that the neighboring Tfap2c and Bmp7 genes are controlled by distinct set of enhancers acting specifically on one or the other gene. Since we observed in a balanced genomic rearrangement a switch of enhancer-gene preferences, the specificity of these enhancers for one or the other gene cannot result exclusively from differences in their promoter structures, as proposed for other situations [49], [60]. In contrast, our results indicate that, for this locus, the regulatory interactions are in a large part determined by the relative position of the different elements, as reported for other complex regions [7], [61], [62].
Our 4C experiments showed that Bmp7 and Tfap2c lie in genomic domains that share limited physical contacts. These domains were only weakly demarcated in the available Hi-C data in ES cells [34]. Therefore, it is unclear if the Tfap2c and Bmp7 domains correspond to adjacent sub-TADs [41], or weak TADs in a rather unstructured region. However, the distinction between these different levels of spatial segregation of the genome may in part be semantic, based on arbitrary thresholds, which may not be pertinent for gene regulation. We showed here that the distinct enhancers that regulate each gene (this work, [53], [63]) reside and act within the corresponding conformational domain, further supporting the functional relevance of the structural partition we described in establishing distinct domains of regulation [15]. Furthermore, we showed that a balanced rearrangement exchanging the relative position of genes, enhancers and the TZ region led to a concomitant redistribution of physical and regulatory interactions. The switch of the heart enhancer from Bmp7 to Tfap2c and the patterns of contacts observed in this configuration demonstrate together that the topological separation in two distinct domains is key to allocate distant enhancers to one or the other gene.
We observed extensive similarities in the 4C profiles between the different cell tissues assayed, irrespective of the expression state of the corresponding genes. This indicates that the Tfap2c-Bmp7 locus adopts a rather generic conformation which undergoes limited changes in response to transcriptional activity. Such a constitutive folding has also been described for other loci [34], [40], [64]–[66]. It suggests that the structural partitioning of the locus into two domains pre-exists and guides regulatory interactions, instead of deriving from directed interactions between active genes and enhancers.
Our functional dissection of the locus highlights that the transition zone separating the two domains has an important role in organizing this topological subdivision. The fusion of the interaction profiles of the two promoters and the centromeric extension of the Bmp7 interaction domain upon removal of the TZ strongly argue in favor of the TZ preventing interactions between Bmp7 and Tfap2c. The different balanced inversions further demonstrate that the TZ organizes this topological separation irrespectively of the nature of its flanking sequences. Interestingly, the TZ region interacts robustly with both flanking regions, suggesting that the topological segregation between Tfap2c and Bmp7 may arise from its action as an interaction sink or decoy, not as a blocker or repulsive element. TAD “boundaries” often displayed strong interactions with regions flanking them on both side [34], suggesting that this behavior could be a rather general feature of topological transitions. The TZ does not appear to coincide with a region of constitutive transcription, contrarily to a large subset of typical TAD boundaries [34]. It is flanked by and includes several constitutive CTCF sites [38]. CTCF sites have been proposed to anchor long-range interactions and to act, together with cohesin and Mediator complexes, as master regulators of the chromosomal 3D conformation [67], [68]. However, as only a subset of CTCF sites act as insulators [15], [69], and as depletion of CTCF only mildly impacts chromosomal topologies [70] and long-range gene regulation [71], the precise role of these sequences – and of other regions of the TZ – would need to be directly assessed.
With regard to the allocation of the heart enhancer, the TZ behave similarly to a classical insulator (Fig. 7). However, the analysis of INV-L1 and –L2 indicates that the TZ does not provide complete shielding from external influences, as the presence, beyond the TZ, of an active Bmp7 promoter can interfere with the expression of Tfap2c in the medial forebrain. Although contacts between Bmp7 and Tfap2c and its associated forebrain enhancer(s) are limited and even insufficient to lead to productive interactions (i.e. activation of Bmp7), they are nonetheless present at higher than background level. Our data suggests that they may be frequent and/or strong enough to perturb the regulation of Tfap2c by its forebrain enhancer(s), most probably through promoter competition. Several studies have reported that promoters have a tendency to come into close proximity [40], [72], [73], particularly when they are co-active and linked. Our analysis indicates that the TZ appears to counteract this generic promoter clustering by limiting admixing of the two domains, but it does not however totally prevent the diffusion of regulatory influences between them. The functional impact of these influences underscores the difficulties of defining functional thresholds for the interaction data obtained with 4C or Hi-C. It also emphasizes that topological domains should not be considered as strict autarchic units: topological separation does not exclude neighborly relationships and semipermeable borders. Transformation of the intrinsically broad forebrain activity of FB1 into the graded expression pattern shown by Tfap2c may involve additional neighboring enhancer elements, as hinted to by the INV-M data. However, our observations suggest that the permeability of the TZ to active Bmp7 may also contribute to this fine-tuning (Fig. 7C). In operational terms, the TZ should be considered as a rheostatic controller rather than as a strict insulator.
Fig. 7. Structural partitioning controls enhancer-target gene allocation and modulates enhancers' effective activity on target genes. 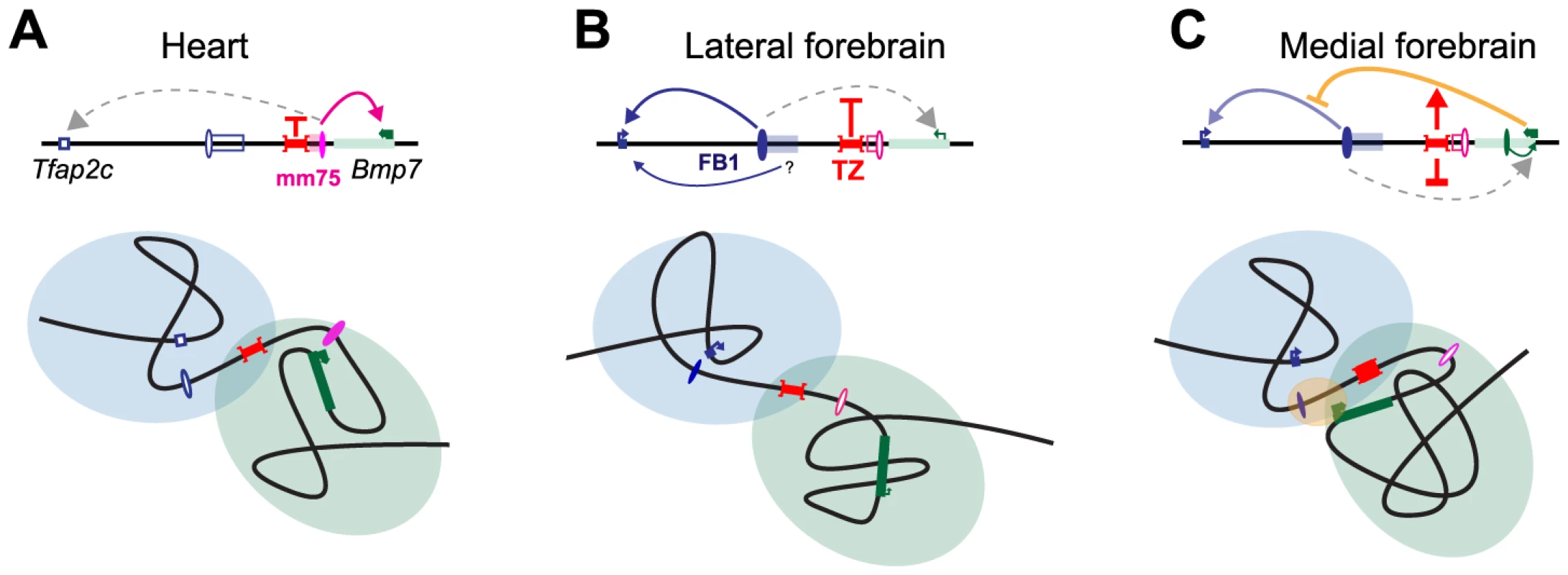
Genes and enhancers are shown as rectangles and ovals, respectively. Active promoters and enhancers are marked with arrows and plain colors. The TZ organizes the locus into two distinct, partially overlapping spatial conformations (represented by light blue and green circles), where genes and enhancers can interact. In the heart (A) and forebrain (B), this situation prevents action of one enhancer on a gene in the other domain. In the lateral forebrain, enhancers adjacent to FB1 may contribute to Tfap2c expression. In the medial forebrain (C), the active Bmp7 promoter may compete, non productively, for the forebrain enhancer, and interferes (marked by a yellow oval) with its action on Tfap2c. The TZ may control the strength and therefore the consequences of this interference on Tfap2c expression. Interestingly, a sequence orthologous to FB1 is present between Tfap2c and Bmp7 in the coelacanth, but not in teleosts or sharks (S12 Fig.). This indicates that the origin of FB1 can be traced back to the ancestor of the lobe-finned fishes. In contrast, the sequence of the TZ region is far less conserved, suggesting a more recent origin. Expression of Bmp7 in the forebrain is likely an ancestral feature, as it is shared amongst Bmp7 orthologues and paralogues [44]. Conversely, Tfap2c is the only member of its family expressed in the forebrain [42], [45], and the only one directly adjacent to a Bmp gene. The evolution of FB1 as a forebrain enhancer may have been favoured by the pre-existing expression of Bmp7 in this tissue, as suggested for other loci [74]–[76]. In this scenario, we suggest that Bmp7 may have initially been the primary target of this emerging enhancer. The evolution of a region with insulating-like activity would have make FB1 available to Tfap2c. Interestingly, the forebrain expression of Tfap2c regulates the formation of basal progenitors in the developing cortex in mammals [77] and variations of this expression levels, in space and time, have been proposed to account for the increased number of cortical neurons present in higher primates [77]. Changes in gene expression changes are usually attributed to evolution of enhancers or promoters [78]. Our results indicate that a simple change of the filtering capacity of the TZ may also provide evolution with means of modulating gene expression.
Materials and Methods
(See S1 Text for details)
Generation of the different transgenic lines and chromosomal rearrangements
The initial allele used to produce SB-B(3end) was obtained by homologous recombination in ES cells (E14). The targeting construct comprised: the SB8 transposon [79]; an additional loxP site outside of the transposon; a neomycin resistance gene under the control of the PGK promoter that are flanked by two FRT sequences. The homology arms (chr2 : 172686051–172689701 and chr2 : 172689702–172694528 (NCBI37/mm9)) were amplified by PCR and then attached to the targeting construct above. After transformation and selection in ES cells, correctly targeted clones were injected into donor C57BL/6J blastocyst. Germline transmission was obtained from one chimera. The FRT-flanked selection cassette was then removed by breeding with hACTB-FLPe mice, leaving only the transposon and the loxP sequence outside of it at the site (allele SB-B(3end)). The ES clone BA0758 was obtained from BayGenomics, verified by PCR genotyping, and injected to establish a Tfap2c-gene trap line.
The SB transposon was remobilised and new insertions were mapped as described before [14]. Alleles carrying the different deletions, duplications and inversions were produced by in vivo genomic engineering [18], [54], using the 129S1/Sv-Hprttm1(cre)Mnn/J CRE line [80]. Deletions del1 and del3 were obtained by recombination in cis between the static loxP site at the end of Bmp7 and the one moved along with the transposed insertion SB-A1 and SB-Sall4, respectively. To keep the regulatory sensor at the deletion breakpoint, we also produced another version of these deletions, del1-LacZ and del3-LacZ, by CRE-mediated recombination in trans [54], between the loxP site from SB-B(3end) and the one at SB-A1 and SB-Sall4, respectively. For the del2-lacZ allele, we used a recombination in trans, between SB-B(3end) and BA0758. Mice were genotyped by PCR (see Supplemental Experimental Procedures).
Mouse experiments were conducted in accordance with the principles and guidelines in place at European Molecular Biology Laboratory, as defined and overseen by its Institutional Animal Care and Use Committee, in accordance with the European Convention 18/3/1986 and Directives 86/609/EEC and 2010/63/EU.
Gene and reporter gene expression analysis
LacZ staining and whole-mount in situ hybridization was carried out following standard protocols. For RT-qPCR, total RNA was extracted from the frozen tissues using RNeasy kit (QIAGEN), and then cDNA was synthesized using the ProtoScript II First Strand cDNA Synthesis Kit (New England Biolabs). The quantitative PCR was performed using StepOne Real-Time PCR System with SYBR green reagent (Applied Biosystems). Gapdh was used to normalize expression level for each sample. The extra-embryonic membranes were used for PCR-genotyping of the embryos.
In vivo enhancer assay
We cloned the FB1 enhancer (chr2 : 172551998–172555000, NCBI37/mm9) upstream of the reporter gene used in SB8, in a lentiviral vector [81]. The transgenic provirus was produced in HEK293 cells as described elsewhere [81]. Briefly, the virus was micro-injected under the zona pellucida of one-cell embryos which were maintained in culture up to the blastocyst stage. Embryos were then reimplanted into foster mothers and, at stage E11.5 or E12.5, stained for LacZ activity and genotyped.
3C assay, 4C library preparation, sequencing and data analysis
To prepare the 3C library we dissected out the heart and the lateral and medial forebrains from E11.5 C57BL/6 embryos. The cells were dissociated, fixed and then processed following the protocol in Splinter et al. [82]. The fixed genomic DNA was digested with NlaIII enzyme and subsequently self-ligated. To quantify the ligation products of interest, we conducted qPCR with TaqMan probes. qPCR was performed with four technical replicates, and for each value, mean and standard deviation were plotted.
For the 4C analyses, the 3C libraries were first prepared as described above from the respective tissues with NlaIII enzyme. They were then subjected to digestion by DpnII and ligation. After purification of the circularized DNA, inverse PCR was performed to obtain 4C libraries. Reading primers had 3–6 nucleotides of tag sequence, to allow for demultiplexing of the pooled libraries after sequencing. PCR products were purified, mixed altogether and sequenced on a HiSeq 2000 (Illumina). For data analysis, we first demultiplexed the FASTQ files of the 4C sequencing libraries and then aligned them to the mm9 reference genome using Bowtie version 1.0.0 [83]. To normalize with regard to library size, we divided the counts by the total number of counts on the viewpoint chromosome (chr2) for each library and multiplied these values by 1,000,000 (“RPM normalization”). We then smoothed the counts over adjacent fragments, using a window size of 11 fragments. Details are available in Supplementary Information. Sequencing data of the 4C libraries is deposited at ENA (Study Accession ERP005557)
Supporting Information
Zdroje
1. ViselA, RubinEM, RubinEM, PennacchioLA, PennacchioLA (2009) Genomic views of distant-acting enhancers. Nature 461 : 199–205.
2. SakabeNJ, SavicD, NobregaMA (2012) Transcriptional enhancers in development and disease. Genome Biol 13 : 238.
3. De GobbiM (2006) A Regulatory SNP Causes a Human Genetic Disease by Creating a New Transcriptional Promoter. Science 312 : 1215–1217.
4. MontavonT, ThevenetL, DubouleD (2012) Impact of copy number variations (CNVs) on long-range gene regulation at the HoxD locus. Proc Natl Acad Sci U S A 109 : 20204–20211.
5. PeichelCL, PrabhakaranB, VogtTF (1997) The mouse Ulnaless mutation deregulates posterior HoxD gene expression and alters appendicular patterning. Development 124 : 3481–3492.
6. KokubuC, WilmB, KokubuT, WahlM, RodrigoI, et al. (2003) Undulated short-tail deletion mutation in the mouse ablates Pax1 and leads to ectopic activation of neighboring Nkx2-2 in domains that normally express Pax1. Genetics 165 : 299–307.
7. MarinićM, AktasT, RufS, SpitzF (2013) An integrated holo-enhancer unit defines tissue and gene specificity of the Fgf8 regulatory landscape. Dev Cell 24 : 530–542.
8. LetticeLA, DanielsS, SweeneyE, VenkataramanS, DevenneyPS, et al. (2011) Enhancer-adoption as a mechanism of human developmental disease. Hum Mutat 32 : 1492–1499.
9. SpitzF, GonzalezF, DubouleD (2003) A global control region defines a chromosomal regulatory landscape containing the HoxD cluster. Cell 113 : 405–417.
10. SpielmannM, BrancatiF, KrawitzPM, RobinsonPN, IbrahimDM, et al. (2012) Homeotic Arm-to-Leg Transformation Associated with Genomic Rearrangements at the PITX1 Locus. Am J of Hum Genet 91 : 629–635.
11. KlopockiE, LohanS, BrancatiF, KollR, BrehmA, et al. (2011) Copy-number variations involving the IHH locus are associated with syndactyly and craniosynostosis. Am J Hum Genet 88 : 70–75.
12. GostissaM, YanCT, BiancoJM, CognéM, PinaudE, et al. (2009) Long-range oncogenic activation of Igh-c-myc translocations by the Igh 3' regulatory region. Nature 462 : 803–807.
13. BanerjiJ, RusconiS, SchaffnerW (1981) Expression of a beta-globin gene is enhanced by remote SV40 DNA sequences. Cell 27 : 299–308.
14. RufS, SymmonsO, UsluVV, DolleD, HotC, et al. (2011) Large-scale analysis of the regulatory architecture of the mouse genome with a transposon-associated sensor. Nat Genet 43 : 379–386.
15. SymmonsO, UsluVV, TsujimuraT, RufS, NassariS, et al. (2014) Functional and topological characteristics of mammalian regulatory domains. Genome Res 24 : 390–400.
16. JinL, LongL, GreenMA, SpearBT (2009) The alpha-fetoprotein enhancer region activates the albumin and alpha-fetoprotein promoters during liver development. Dev Biol 336 : 294–300.
17. NickolJM, FelsenfeldG (1988) Bidirectional control of the chicken beta - and epsilon-globin genes by a shared enhancer. Proc Natl Acad Sci USA 85 : 2548–2552.
18. SpitzF, HerkenneC, MorrisMA, DubouleD (2005) Inversion-induced disruption of the Hoxd cluster leads to the partition of regulatory landscapes. Nat Genet 37 : 889–893.
19. TsujimuraT, HosoyaT, KawamuraS (2010) A Single Enhancer Regulating the Differential Expression of Duplicated Red-Sensitive Opsin Genes in Zebrafish. PLoS Genet 6: e1001245.
20. SumiyamaK, IrvineSQ, StockDW, WeissKM, KawasakiK, et al. (2002) Genomic structure and functional control of the Dlx3–7 bigene cluster. Proc Natl Acad Sci USA 99 : 780–785.
21. XuX, ScottMM, DenerisES (2006) Shared long-range regulatory elements coordinate expression of a gene cluster encoding nicotinic receptor heteromeric subtypes. Mol Cell Biol 26 : 5636–5649.
22. CarvajalJJ, CoxD, SummerbellD, RigbyPW (2001) A BAC transgenic analysis of the Mrf4/Myf5 locus reveals interdigitated elements that control activation and maintenance of gene expression during muscle development. Development 128 : 1857–1868.
23. ZunigaA, MichosO, SpitzF, HaramisA-PG, PanmanL, et al. (2004) Mouse limb deformity mutations disrupt a global control region within the large regulatory landscape required for Gremlin expression. Genes Dev 18 : 1553–1564.
24. CajiaoI, ZhangA, YooEJ, CookeNE, LiebhaberSA (2004) Bystander gene activation by a locus control region. EMBO J 23 : 3854–3863.
25. LowerKM, HughesJR, De GobbiM, HendersonS, ViprakasitV, et al. (2009) Adventitious changes in long-range gene expression caused by polymorphic structural variation and promoter competition. Proc Natl Acad Sci U S A 106 : 21771–21776.
26. CalhounVC, LevineM (2003) Long-range enhancer-promoter interactions in the Scr-Antp interval of the Drosophila Antennapedia complex. Proc Natl Acad Sci USA 100 : 9878–9883.
27. OhtsukiS, LevineM, CaiHN (1998) Different core promoters possess distinct regulatory activities in the Drosophila embryo. Genes Dev 12 : 547–556.
28. CaiH, LevineM (1995) Modulation of enhancer-promoter interactions by insulators in the Drosophila embryo. Nature 376 : 533–536 doi:10.1038/376533a0
29. GeyerPK, CorcesVG (1992) DNA position-specific repression of transcription by a Drosophila zinc finger protein. Genes Dev 6 : 1865–1873.
30. KellumR, SchedlP (1992) A group of scs elements function as domain boundaries in an enhancer-blocking assay. Mol Cell Biol 12 : 2424–2431.
31. YangJ, CorcesVG (2011) Chromatin insulators: a role in nuclear organization and gene expression. Adv Cancer Res 110 : 43–76.
32. ChetverinaD, AokiT, ErokhinM, GeorgievP, SchedlP (2014) Making connections: Insulators organize eukaryotic chromosomes into independent cis-regulatory networks. Bioessays 36 : 163–172.
33. DekkerJ, Marti-RenomMA, MirnyLA (2013) Exploring the three-dimensional organization of genomes: interpreting chromatin interaction data. Nat Rev Genet 14 : 390–403.
34. DixonJR, SelvarajS, YueF, KimA, LiY, et al. (2012) Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485 : 376–380.
35. NoraEP, LajoieBR, SchulzEG, GiorgettiL, OkamotoI, et al. (2012) Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485 : 381–385.
36. NoraEP, DekkerJ, HeardE (2013) Segmental folding of chromosomes: a basis for structural and regulatory chromosomal neighborhoods? Bioessays 35 : 818–828.
37. GibcusJH, DekkerJ (2013) The Hierarchy of the 3D Genome. Mol Cell 49 : 773–782.
38. ShenY, YueF, McClearyDF, YeZ, EdsallL, et al. (2012) A map of the cis-regulatory sequences in the mouse genome. Nature 488 : 116–120.
39. AndreyG, MontavonT, MascrezB, GonzalezF, NoordermeerD, et al. (2013) A switch between topological domains underlies HoxD genes collinearity in mouse limbs. Science 340 : 1234167.
40. JinF, LiY, DixonJR, SelvarajS, YeZ, et al. (2013) A high-resolution map of the three-dimensional chromatin interactome in human cells. Nature 503 : 290–294.
41. Phillips-CreminsJE, SauriaMEG, SanyalA, GerasimovaTI, LajoieBR, et al. (2013) Architectural Protein Subclasses Shape 3D Organization of Genomes during Lineage Commitment. Cell 153 : 1281–1295.
42. ChazaudC, Oulad-AbdelghaniM, BouilletP, DécimoD, ChambonP, et al. (1996) AP-2.2, a novel gene related to AP-2, is expressed in the forebrain, limbs and face during mouse embryogenesis. Mech Dev 54 : 83–94.
43. HofmannC, LuoG, BallingR, KarsentyG (1996) Analysis of limb patterning in BMP-7-deficient mice. Dev Genet 19 : 43–50.
44. FurutaY, PistonDW, HoganBL (1997) Bone morphogenetic proteins (BMPs) as regulators of dorsal forebrain development. Development 124 : 2203–2212.
45. ZhaoF, LufkinT, GelbBD (2003) Expression of Tfap2d, the gene encoding the transcription factor Ap-2 delta, during mouse embryogenesis. Gene Expr Patterns 3 : 213–217.
46. GrayPA, FuH, LuoP, ZhaoQ, YuJ, et al. (2004) Mouse brain organization revealed through direct genome-scale TF expression analysis. Science 306 : 2255–2257.
47. ZouvelouV, LuderH-U, MitsiadisTA, GrafD (2009) Deletion of BMP7 affects the development of bones, teeth, and other ectodermal appendages of the orofacial complex. Journal of experimental zoology Part B, Molecular and developmental evolution. 312B: 361–374 d.
48. DaneshSM, VillasenorA, ChongD, SoukupC, CleaverO (2009) BMP and BMP receptor expression during murine organogenesis. Gene Expr Patterns 9 : 255–265.
49. AkalinA, FredmanD, ArnerE, DongX, BryneJC, et al. (2009) Transcriptional features of genomic regulatory blocks. Genome Biol 10: R38.
50. YeeSP, RigbyPW (1993) The regulation of myogenin gene expression during the embryonic development of the mouse. Genes Dev 7 : 1277–1289.
51. StrykeD, KawamotoM, HuangCC, JohnsSJ, KingLA, et al. (2003) BayGenomics: a resource of insertional mutations in mouse embryonic stem cells. Nucleic Acids Res 31 : 278–281.
52. KondoT, DubouleD (1999) Breaking colinearity in the mouse HoxD complex. Cell 97 : 407–417.
53. AdamsD, KarolakM, RobertsonE, OxburghL (2007) Control of kidney, eye and limb expression of Bmp7 by an enhancer element highly conserved between species. Developmental biology 311 : 679–690.
54. HéraultY, RassoulzadeganM, CuzinF, DubouleD (1998) Engineering chromosomes in mice through targeted meiotic recombination (TAMERE). Nat Genet 20 : 381–384.
55. NordAS, BlowMJ, AttanasioC, AkiyamaJA, HoltA, et al. (2013) Rapid and pervasive changes in genome-wide enhancer usage during mammalian development. Cell 155 : 1521–1531.
56. ViselA, TaherL, GirgisH, MayD, GolonzhkaO, et al. (2013) A High-Resolution Enhancer Atlas of the Developing Telencephalon. Cell 152 : 895–908.
57. BlowMJ, McCulleyDJ, LiZ, ZhangT, AkiyamaJA, et al. (2010) ChIP-Seq identification of weakly conserved heart enhancers. Nat Genet 42 : 806–810.
58. ViselA, MinovitskyS, DubchakI, PennacchioLA (2007) VISTA Enhancer Browser—a database of tissue-specific human enhancers. Nucleic Acids Res 35: D88–D92.
59. HuberW, ToedlingJ, SteinmetzLM (2006) Transcript mapping with high-density oligonucleotide tiling arrays. Bioinformatics 22 : 1963–1970.
60. NolisIK, McKayDJ, MantouvalouE, LomvardasS, MerikaM, et al. (2009) Transcription factors mediate long-range enhancer-promoter interactions. Proc Natl Acad Sci U S A 106 : 20222–20227.
61. TanimotoK, LiuQ, BungertJ, EngelJD (1999) Effects of altered gene order or orientation of the locus control region on human beta-globin gene expression in mice. Nature 398 : 344–348.
62. KmitaM, FraudeauN, HéraultY, DubouleD (2002) Serial deletions and duplications suggest a mechanism for the collinearity of Hoxd genes in limbs. Nature 420 : 145–150.
63. ParkJ-S, MaW, O'BrienLL, ChungE, GuoJ-J, et al. (2012) Six2 and Wnt regulate self-renewal and commitment of nephron progenitors through shared gene regulatory networks. Dev Cell 23 : 637–651.
64. MontavonT, SoshnikovaN, MascrezB, JoyeE, ThevenetL, et al. (2011) A regulatory archipelago controls Hox genes transcription in digits. Cell 147 : 1132–1145.
65. DostieJ, RichmondTA, ArnaoutRA, SelzerRR, LeeWL, et al. (2006) Chromosome Conformation Capture Carbon Copy (5C): a massively parallel solution for mapping interactions between genomic elements. Genome Res 16 : 1299–1309.
66. Ghavi-HelmY, KleinFA, PakozdiT, CiglarL, NoordermeerD, et al. (2014) Enhancer loops appear stable during development and are associated with paused polymerase. Nature 512 : 96–100.
67. PhillipsJE, CorcesVG (2009) CTCF: master weaver of the genome. Cell 137 : 1194–1211.
68. MerkenschlagerM, OdomDT (2013) CTCF and cohesin: linking gene regulatory elements with their targets. Cell 152 : 1285–1297.
69. SanyalA, LajoieBR, JainG, DekkerJ (2012) The long-range interaction landscape of gene promoters. Nature 489 : 109–113.
70. ZuinJ, DixonJR, van der ReijdenMIJA, YeZ, KolovosP, et al. (2014) Cohesin and CTCF differentially affect chromatin architecture and gene expression in human cells. Proc Natl Acad Sci U S A 111 : 996–1001.
71. SoshnikovaN, MontavonT, LeleuM, GaljartN, DubouleD (2010) Functional Analysis of CTCF During Mammalian Limb Development. Dev Cell 19 : 819–830.
72. SchoenfelderS, SextonT, ChakalovaL, CopeNF, HortonA, et al. (2010) Preferential associations between co-regulated genes reveal a transcriptional interactome in erythroid cells. Nat Genet 42 : 53–61.
73. LiG, RuanX, AuerbachRK, SandhuKS, ZhengM, et al. (2012) Extensive promoter-centered chromatin interactions provide a topological basis for transcription regulation. Cell 148 : 84–98.
74. EichenlaubMP, EttwillerL (2011) De novo genesis of enhancers in vertebrates. PLoS Biol 9: e1001188.
75. GompelN, Prud'hommeB, WittkoppPJ, KassnerVA, CarrollSB (2005) Chance caught on the wing: cis-regulatory evolution and the origin of pigment patterns in Drosophila. Nature 433 : 481–487.
76. FranchiniLF, López-LealR, NasifS, BeatiP, GelmanDM, et al. (2011) Convergent evolution of two mammalian neuronal enhancers by sequential exaptation of unrelated retroposons. Proc Natl Acad Sci U S A 108 : 15270–15275.
77. PintoL, DrechselD, SchmidM-T, NinkovicJ, IrmlerM, et al. (2009) AP2gamma regulates basal progenitor fate in a region - and layer-specific manner in the developing cortex. Nat Neurosci 12 : 1229–1237.
78. WittkoppPJ, KalayG (2012) Cis-regulatory elements: molecular mechanisms and evolutionary processes underlying divergence. Nat Rev Genet 13 : 59–69.
79. ChenC-K, SymmonsO, UsluVV, TsujimuraT, RufS, et al. (2013) TRACER: a resource to study the regulatory architecture of the mouse genome. BMC Genomics 14 : 215.
80. TangS-HE, SilvaFJ, TsarkWMK, MannJR (2002) A Cre/loxP-deleter transgenic line in mouse strain 129S1/SvImJ. Genesis 32 : 199–202.
81. FriedliM, BardeI, ArcangeliM, VerpS, QuazzolaA, et al. (2010) A systematic enhancer screen using lentivector transgenesis identifies conserved and non-conserved functional elements at the Olig1 and Olig2 locus. PLoS ONE 5: e15741.
82. SplinterE, de WitE, van de WerkenHJG, KlousP, de LaatW (2012) Determining long-range chromatin interactions for selected genomic sites using 4C-seq technology: From fixation to computation. Methods 58 : 221–230.
83. LangmeadB, TrapnellC, PopM, SalzbergSL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10: R25.
84. DerrienT, EstelléJ, SolaSM, KnowlesDG, RaineriE, et al. (2012) Fast Computation and Applications of Genome Mappability. PLoS ONE 7: e30377.
85. AttanasioC, NordAS, ZhuY, BlowMJ, BiddieSC, et al. (2014) Tissue-specific SMARCA4 binding at active and repressed regulatory elements during embryogenesis. Genome Res 24 : 920–929.
86. ViselA, BlowMJ, LiZ, ZhangT, AkiyamaJA, et al. (2009) ChIP-seq accurately predicts tissue-specific activity of enhancers. Nature 457 : 854–858.
Štítky
Genetika Reprodukční medicína
Článek Phosphorylation of Elp1 by Hrr25 Is Required for Elongator-Dependent tRNA Modification in YeastČlánek Naturally Occurring Differences in CENH3 Affect Chromosome Segregation in Zygotic Mitosis of HybridsČlánek Insight in Genome-Wide Association of Metabolite Quantitative Traits by Exome Sequence AnalysesČlánek ALIX and ESCRT-III Coordinately Control Cytokinetic Abscission during Germline Stem Cell DivisionČlánek Deciphering the Genetic Programme Triggering Timely and Spatially-Regulated Chitin Deposition
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2015 Číslo 1
-
Všechny články tohoto čísla
- The Combination of Random Mutagenesis and Sequencing Highlight the Role of Unexpected Genes in an Intractable Organism
- Ataxin-3, DNA Damage Repair, and SCA3 Cerebellar Degeneration: On the Path to Parsimony?
- α-Actinin-3: Why Gene Loss Is an Evolutionary Gain
- Origins of Context-Dependent Gene Repression by Capicua
- Transposable Elements Contribute to Activation of Maize Genes in Response to Abiotic Stress
- No Evidence for Association of Autism with Rare Heterozygous Point Mutations in Contactin-Associated Protein-Like 2 (), or in Other Contactin-Associated Proteins or Contactins
- Nur1 Dephosphorylation Confers Positive Feedback to Mitotic Exit Phosphatase Activation in Budding Yeast
- A Regulatory Hierarchy Controls the Dynamic Transcriptional Response to Extreme Oxidative Stress in Archaea
- Genetic Variants Modulating CRIPTO Serum Levels Identified by Genome-Wide Association Study in Cilento Isolates
- Small RNA Sequences Support a Host Genome Origin of Satellite RNA
- Phosphorylation of Elp1 by Hrr25 Is Required for Elongator-Dependent tRNA Modification in Yeast
- Genetic Mapping of MAPK-Mediated Complex Traits Across
- An AP Endonuclease Functions in Active DNA Demethylation and Gene Imprinting in
- Developmental Regulation of the Origin Recognition Complex
- End of the Beginning: Elongation and Termination Features of Alternative Modes of Chromosomal Replication Initiation in Bacteria
- Naturally Occurring Differences in CENH3 Affect Chromosome Segregation in Zygotic Mitosis of Hybrids
- Imputation of the Rare G84E Mutation and Cancer Risk in a Large Population-Based Cohort
- Polycomb Protein SCML2 Associates with USP7 and Counteracts Histone H2A Ubiquitination in the XY Chromatin during Male Meiosis
- A Genetic Strategy for Probing the Functional Diversity of Magnetosome Formation
- Interactions of Chromatin Context, Binding Site Sequence Content, and Sequence Evolution in Stress-Induced p53 Occupancy and Transactivation
- The Yeast La Related Protein Slf1p Is a Key Activator of Translation during the Oxidative Stress Response
- Integrative Analysis of DNA Methylation and Gene Expression Data Identifies as a Key Regulator of COPD
- Proteasomes, Sir2, and Hxk2 Form an Interconnected Aging Network That Impinges on the AMPK/Snf1-Regulated Transcriptional Repressor Mig1
- Functional Interplay between the 53BP1-Ortholog Rad9 and the Mre11 Complex Regulates Resection, End-Tethering and Repair of a Double-Strand Break
- Estrogenic Exposure Alters the Spermatogonial Stem Cells in the Developing Testis, Permanently Reducing Crossover Levels in the Adult
- Protein Poly(ADP-ribosyl)ation Regulates Immune Gene Expression and Defense Responses
- Sumoylation Influences DNA Break Repair Partly by Increasing the Solubility of a Conserved End Resection Protein
- A Discrete Transition Zone Organizes the Topological and Regulatory Autonomy of the Adjacent and Genes
- Elevated Mutation Rate during Meiosis in
- The Intersection of the Extrinsic Hedgehog and WNT/Wingless Signals with the Intrinsic Hox Code Underpins Branching Pattern and Tube Shape Diversity in the Airways
- MiR-24 Is Required for Hematopoietic Differentiation of Mouse Embryonic Stem Cells
- Tissue-Specific Effects of Genetic and Epigenetic Variation on Gene Regulation and Splicing
- Heterologous Aggregates Promote Prion Appearance via More than One Mechanism
- The Tumor Suppressor BCL7B Functions in the Wnt Signaling Pathway
- , A -Acting Locus that Controls Chromosome-Wide Replication Timing and Stability of Human Chromosome 15
- Regulating Maf1 Expression and Its Expanding Biological Functions
- A Polyubiquitin Chain Reaction: Parkin Recruitment to Damaged Mitochondria
- RecFOR Is Not Required for Pneumococcal Transformation but Together with XerS for Resolution of Chromosome Dimers Frequently Formed in the Process
- An Intracellular Transcriptomic Atlas of the Giant Coenocyte
- Insight in Genome-Wide Association of Metabolite Quantitative Traits by Exome Sequence Analyses
- The Role of the Mammalian DNA End-processing Enzyme Polynucleotide Kinase 3’-Phosphatase in Spinocerebellar Ataxia Type 3 Pathogenesis
- The Global Regulatory Architecture of Transcription during the Cell Cycle
- Identification and Functional Characterization of Coding Variants Influencing Glycemic Traits Define an Effector Transcript at the Locus
- Altered Ca Kinetics Associated with α-Actinin-3 Deficiency May Explain Positive Selection for Null Allele in Human Evolution
- Genetic Variation in the Nuclear and Organellar Genomes Modulates Stochastic Variation in the Metabolome, Growth, and Defense
- PRDM9 Drives Evolutionary Erosion of Hotspots in through Haplotype-Specific Initiation of Meiotic Recombination
- Transcriptional Control of an Essential Ribozyme in Reveals an Ancient Evolutionary Divide in Animals
- ALIX and ESCRT-III Coordinately Control Cytokinetic Abscission during Germline Stem Cell Division
- Century-scale Methylome Stability in a Recently Diverged Lineage
- A Re-examination of the Selection of the Sensory Organ Precursor of the Bristle Sensilla of
- Antagonistic Cross-Regulation between Sox9 and Sox10 Controls an Anti-tumorigenic Program in Melanoma
- A Dependent Pool of Phosphatidylinositol 4,5 Bisphosphate (PIP) Is Required for G-Protein Coupled Signal Transduction in Photoreceptors
- Deciphering the Genetic Programme Triggering Timely and Spatially-Regulated Chitin Deposition
- Aberrant Gene Expression in Humans
- Fascin1-Dependent Filopodia are Required for Directional Migration of a Subset of Neural Crest Cells
- The SWI2/SNF2 Chromatin Remodeler BRAHMA Regulates Polycomb Function during Vegetative Development and Directly Activates the Flowering Repressor Gene
- Evolutionary Constraint and Disease Associations of Post-Translational Modification Sites in Human Genomes
- A Truncated NLR Protein, TIR-NBS2, Is Required for Activated Defense Responses in the Mutant
- The Genetic and Mechanistic Basis for Variation in Gene Regulation
- Inactivation of PNKP by Mutant ATXN3 Triggers Apoptosis by Activating the DNA Damage-Response Pathway in SCA3
- DNA Damage Response Factors from Diverse Pathways, Including DNA Crosslink Repair, Mediate Alternative End Joining
- hnRNP K Coordinates Transcriptional Silencing by SETDB1 in Embryonic Stem Cells
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- The Global Regulatory Architecture of Transcription during the Cell Cycle
- A Truncated NLR Protein, TIR-NBS2, Is Required for Activated Defense Responses in the Mutant
- Proteasomes, Sir2, and Hxk2 Form an Interconnected Aging Network That Impinges on the AMPK/Snf1-Regulated Transcriptional Repressor Mig1
- The SWI2/SNF2 Chromatin Remodeler BRAHMA Regulates Polycomb Function during Vegetative Development and Directly Activates the Flowering Repressor Gene
Přihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



