-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praktické
Top novinky
ReklamaAllelic Exchange of Pheromones and Their Receptors Reprograms Sexual Identity in
Cell type specification is a fundamental process that all cells must carry out to ensure appropriate behaviors in response to environmental stimuli. In fungi, cell identity is critical for defining “sexes” known as mating types and is controlled by components of mating type (MAT) loci. MAT–encoded genes function to define sexes via two distinct paradigms: 1) by controlling transcription of components common to both sexes, or 2) by expressing specially encoded factors (pheromones and their receptors) that differ between mating types. The human fungal pathogen Cryptococcus neoformans has two mating types (a and α) that are specified by an extremely unusual MAT locus. The complex architecture of this locus makes it impossible to predict which paradigm governs mating type. To identify the mechanism by which the C. neoformans sexes are determined, we created strains in which the pheromone and pheromone receptor from one mating type (a) replaced the pheromone and pheromone receptor of the other (α). We discovered that these “αa” cells effectively adopt a new mating type (that of a cells); they sense and respond to α factor, they elicit a mating response from α cells, and they fuse with α cells. In addition, αa cells lose the α cell type-specific response to pheromone and do not form germ tubes, instead remaining spherical like a cells. Finally, we discovered that exogenous expression of the diploid/dikaryon-specific transcription factor Sxi2a could then promote complete sexual development in crosses between α and αa strains. These data reveal that cell identity in C. neoformans is controlled fully by three kinds of MAT–encoded proteins: pheromones, pheromone receptors, and homeodomain proteins. Our findings establish the mechanisms for maintenance of distinct cell types and subsequent developmental behaviors in this unusual human fungal pathogen.
Published in the journal: . PLoS Genet 6(2): e32767. doi:10.1371/journal.pgen.1000860
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1000860Summary
Cell type specification is a fundamental process that all cells must carry out to ensure appropriate behaviors in response to environmental stimuli. In fungi, cell identity is critical for defining “sexes” known as mating types and is controlled by components of mating type (MAT) loci. MAT–encoded genes function to define sexes via two distinct paradigms: 1) by controlling transcription of components common to both sexes, or 2) by expressing specially encoded factors (pheromones and their receptors) that differ between mating types. The human fungal pathogen Cryptococcus neoformans has two mating types (a and α) that are specified by an extremely unusual MAT locus. The complex architecture of this locus makes it impossible to predict which paradigm governs mating type. To identify the mechanism by which the C. neoformans sexes are determined, we created strains in which the pheromone and pheromone receptor from one mating type (a) replaced the pheromone and pheromone receptor of the other (α). We discovered that these “αa” cells effectively adopt a new mating type (that of a cells); they sense and respond to α factor, they elicit a mating response from α cells, and they fuse with α cells. In addition, αa cells lose the α cell type-specific response to pheromone and do not form germ tubes, instead remaining spherical like a cells. Finally, we discovered that exogenous expression of the diploid/dikaryon-specific transcription factor Sxi2a could then promote complete sexual development in crosses between α and αa strains. These data reveal that cell identity in C. neoformans is controlled fully by three kinds of MAT–encoded proteins: pheromones, pheromone receptors, and homeodomain proteins. Our findings establish the mechanisms for maintenance of distinct cell types and subsequent developmental behaviors in this unusual human fungal pathogen.
Introduction
One of the most important processes that occurs in cells is the specification of cell type, and it is by this process that cells adopt the genetic state that governs their subsequent behaviors. One mechanism of cell type-specification is the expression of genes encoded only in a given cell type. For example, X and Y chromosomes determine male and female sexes because they encode different genes, such as SRY, a Y-specific protein whose expression establishes male identity [1]. In a similar fashion, the sexes of fungi, known as “mating types,” are also determined through the expression of cell type-specific genes. The most well-characterized cell identity determination system is that of the budding yeast Saccharomyces cerevisiae.
In S. cerevisiae the two haploid cell types, a and α, are distinguished from one another by the actions of specific transcription factors encoded at the mating-type (MAT) locus [2]. MATa encodes the homeodomain transcription factor a1, and MATα encodes α1 and α2, an α-domain protein and a homeodomain protein, respectively. The actions of a1, α1, and α2 govern control of haploid cell behavior through the differential expression of a-specific, and α-specific genes, including pheromone and pheromone receptor genes [3]. It is through a pheromone-pheromone receptor system that cell-cell communication occurs, and cells of opposite mating types can sense one another. Specifically, a cells secrete mating factor a pheromone (MFa), which binds to a receptor (Ste3) on the surface of α cells, and α cells secrete MFα pheromone, which is sensed by a receptor on the surface of a cells (Ste2). In response to the presence of the pheromone of a mating partner, cells undergo a cell cycle arrest and subsequent morphological changes to prepare for mating. After cell fusion, a1 and α2 act in concert to regulate haploid-specific genes, specifying the diploid a/α cell, thus completing a MAT-controlled regulatory circuit [2],[3].
A similar cell-cell recognition process occurs in many other fungi, including the corn smut, Ustilago maydis. In U. maydis specific pheromones and receptors are expressed in different cell types; however, in contrast to S. cerevisiae, these genes are not under the transcriptional control of the homeodomain proteins of the traditional MAT locus [4]. Instead, distinct alleles of the pheromones and their receptors are encoded within a separate MAT locus, and these alleles are sufficiently distinct from one another to confer cell type-specificity. In this case, haploid cells expressing distinct pheromones and receptors from the pheromone MAT locus sense and respond to partners of other mating types and fuse. Once compatible mating types have fused, two transcriptional regulators, bE and bW, which are encoded at the second MAT locus, regulate a transcriptional cascade that promotes further sexual development [4]–[6].
In a related, clinically important human pathogen, Cryptococcus neoformans, the determinants of haploid cell identity are unknown. C. neoformans contains a single MAT locus that is over 100 kb in size and contains 23 genes, some of which have been found to be involved in sexual development and others that appear to be essential “housekeeping” genes [7]. This locus represents an evolutionary transition from the two separate MAT loci found in basidiomycete fungi like U. maydis, to the single MAT locus found in ascomycetes [8]; it is unclear how components in this “fused” MAT locus function to specify haploid cell type. In the C. neoformans MAT locus, there are five genes in each mating type that represent the classic MAT components found in basidiomycete MAT loci. They include the homeodomain transcription factors SXI2a and SXI1α, the six pheromone genes MFa1-3 and MFα1-3, and the pheromone receptors STE3a and STE3α. The transcription factors, Sxi1α and Sxi2a, do not appear to play any role in haploid cells, including establishment of haploid cell identity. In contrast, Sxi1α and Sxi2a are both necessary and sufficient to specify the dikaryotic state following cell fusion and ensure that sexual development continues [9],[10]. Conversely, the pheromones and pheromone receptors of C. neoformans have been shown previously to be necessary for haploid cell behaviors, such as sensing and responding to a mating partner [11]–[16]. What is not clear is whether pheromones and pheromone receptors are sufficient to confer haploid cell identity (as in U. maydis), or whether the actions of other regulators are required to establish the a and α mating types (as in S. cerevisiae).
To test the hypothesis that pheromones and pheromone receptors alone are sufficient to confer haploid cell identity in C. neoformans, we carried out a “swapping” experiment in which components from the a mating-type (STE3a and MFa1) were relocated into a strain of the opposite mating type, α (in which STE3α and all three copies of MFα had been deleted), and the effects of these modifications on mating and development were examined. The results presented here reveal that pheromones and their receptors are both necessary and sufficient to specify haploid cell identity. That is, strains that harbor a receptor and pheromone from the opposite mating type are capable of sensing and fusing with wild type cells of the same mating type. Furthermore, the fused cells are incapable of progressing through sexual development in the absence of both Sxi transcription factors; however, exogenous addition of SXI2a facilitates complete sexual development. These findings reveal that control of haploid cell identity in C. neoformans is mediated by MAT-encoded cell-cell communication components that are not mating type-specific in their downstream effects, and establishes a model for the maintenance of distinct cell types and developmental behaviors in this human fungal pathogen.
Results
Construction of modified α strains
To test the possibility that haploid cell identity in C. neoformans is controlled by pheromones and pheromone receptors, a strain where the pheromone and pheromone receptor genes from one mating type were replaced with those from the opposite mating type was constructed. Specifically, we created an altered α strain by functionally replacing the STE3α receptor gene and the three MFα pheromone genes of an α cell with the STE3a receptor gene and three copies of the MFa1 gene from an a cell (Figure 1).
Fig. 1. αa cells are α cells that express a pheromone and the pheromone receptor from a cells. 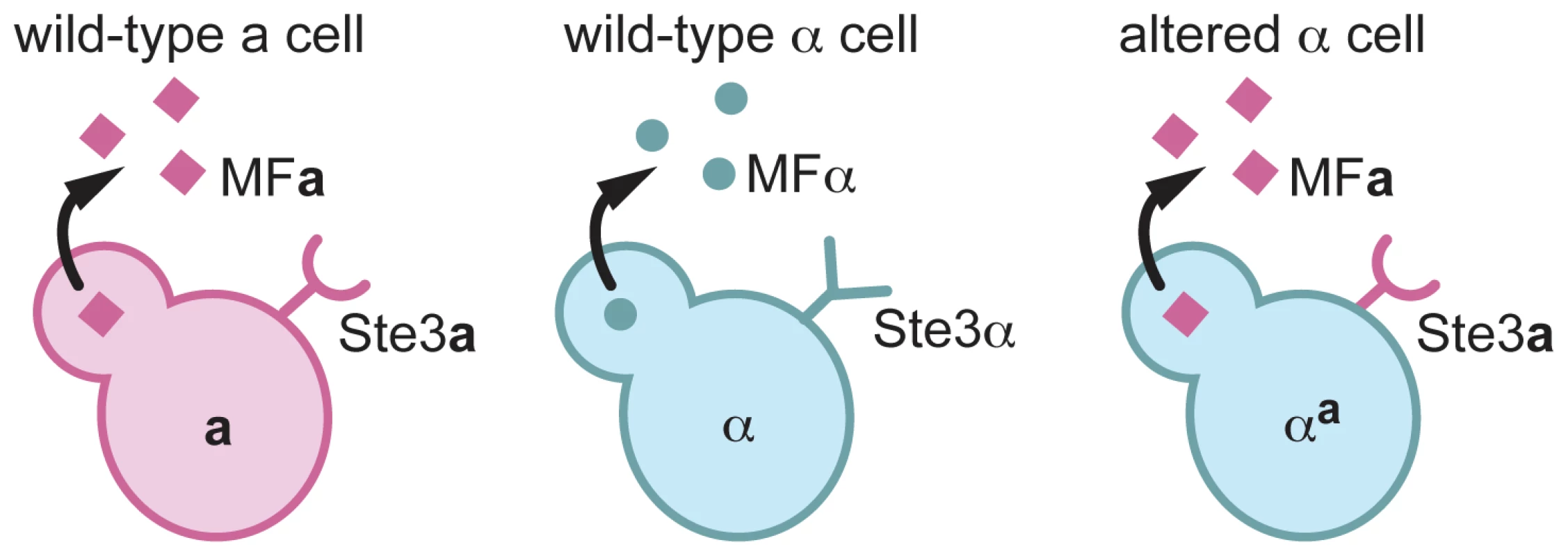
A schematic of pheromones and pheromone receptors produced by a cells (left), α cells (middle), and αa cells (right) is represented. a cells (pink) produce the Ste3a pheromone receptor (pink half circle/stick structure) and MFa pheromones (pink squares). α cells (blue) produce the Ste3α receptor (blue Y-shaped structure) and MFα pheromones (blue circles). αa cells produce only the Ste3a pheromone receptor and the MFa1 pheromone. To construct the altered α strain, α cells in which all three copies of the MFα gene had been deleted (α mfαΔ), were transformed with a construct to replace the endogenous copy of STE3α with STE3a (Figure 2A) [13]. This replacement resulted in a complete deletion of the STE3α open reading frame and the precise insertion of STE3a under the control of the endogenous STE3α promoter. This strain, designated αmfαΔ, STE3a was confirmed by PCR (data not shown) and Southern analysis (Figure 2A) to contain a single, targeted integration of the STE3a gene at the STE3α locus. This αmfαΔ, STE3a strain was then transformed with a construct expressing MFa1 (under the control of the MFα1 promoter) designed to integrate randomly into the genome (Figure 2B). Multiple transformants of the resulting strain, αa, were assessed by Southern blot analysis, and several strains containing integrated copies of the MFa construct were carried forward in further analyses. All of the transformants confirmed to contain the MFa construct exhibited identical behaviors in subsequent phenotype analyses. A representative αa transformant containing three copies of the MFa1 gene is shown in Figure 2B.
Fig. 2. Southern and northern analyses of wild-type and modified α strains. 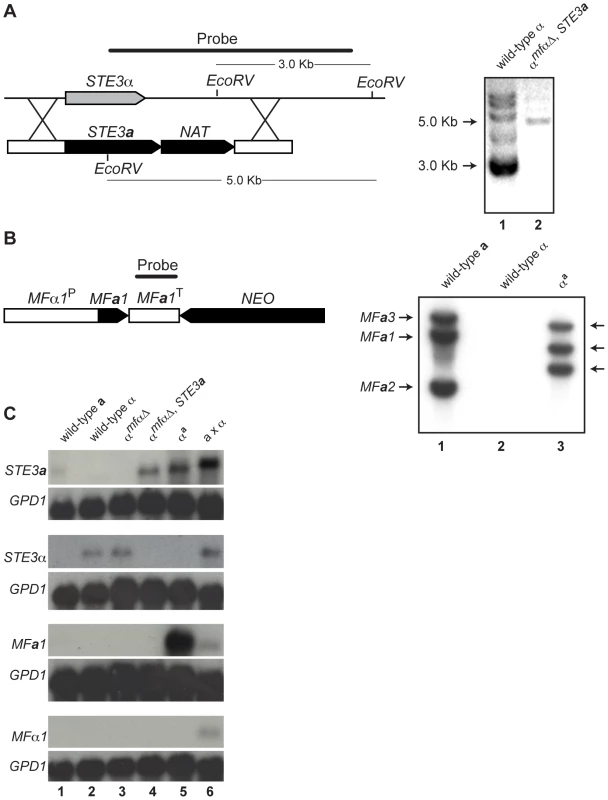
(A) Schematic of the STE3a/NAT integration event to generate the αmfαΔ, STE3a strain (left). Southern blot analysis confirms a single integration of the STE3a-NAT construct into the STE3α locus (Lane 1: ∼3 kb wild type band. Lane 2: ∼5 kb insertion band, right). (B) Schematic of the MFa1/NEO construct to generate the αa strain (left). Southern blot analysis reveals that 3 copies of the MFa1 gene integrated into the genome. Lane 1: three wild type bands present in a cells. Lane 2: no MFa signal in α cells. Lane 3: three randomly integrated copies of MFa in αa cells (right). (C) Northern blot analysis of pheromone receptors (STE3a and STE3α) and pheromones (MFa and MFα). Genotypes of the test strains are indicated over the panels and probes are indicated to the left. A probe to GPD1 was used as a loading and hybridization control. Each lane contains RNA from the following strains or crosses: Lane 1 wild type a, Lane 2 wild type α, Lane 3 αmfαΔ, Lane 4 α mfαΔ, STE3a, Lane 5 αa, and Lane 6 a x α. To assess the expression patterns of pheromones and their receptors in the constructed strains, northern blot analysis was carried out. Haploid strains were grown on V8 medium for 24 hours, and their relative transcript levels were examined (Figure 2C). The STE3a and STE3α transcripts were detected in wild type a and α cells, respectively, and the STE3α transcript was also present in the αmfαΔ strain. However, as expected, in the αmfαΔ, STE3a strain where the STE3a ORF had replaced the STE3α ORF, only STE3a transcript was observed. In summary, the transcripts detected from the constructed strains were in accordance with the predicted expression patterns for each of the strains tested based on their genotypes.
While pheromone transcript was not detected in either haploid cell type (2C, bottom two panels, lanes 1 and 2), this result was not surprising because pheromone genes are transcribed at low levels in haploid cells grown on minimal medium [13]. Furthermore, pheromone transcript was not observed in strains where these genes had been deleted (αmfαΔ and αmfαΔ, STE3a) (lanes 3 and 4). However, when the MFa1 gene was introduced into the αmfαΔ, STE3a strain (resulting in the αa strain), high levels of MFa transcript were detected, and this high level of expression occurred in all of the MFa transformants (lane 5 and data not shown). Because MFa1, in this case, is under the control of the MFα1 promoter (defined here as 1 kb of sequence upstream of the translational start site), a likely explanation for our findings is that normal repression of MFa1 in haploid cells is disrupted. That is, the elements required for wild type levels of expression were likely not included in our construct, resulting in higher levels of MFa1 gene expression. Because pheromones expressed from a non-native promoter do not interfere with development and the mating response, the αa strain was used in our subsequent analyses [13],[14].
Altered α strains mate with wild-type α cells
To evaluate how the series of modified α strains interacted with mating partners under sexual development conditions, crosses using various test strains were carried out and assessed microscopically for the presence of fusants after 18 hours on V8 medium. The cells used in each cross were stained with either rhodamine (Alexa Fluor 594) or fluorescein (Alexa Fluor 488) prior to mixing with partner cells so that the original mating types could be discerned after cell fusion.
In crosses between wild type a and α strains, fusants were identified as “dumbbell” shaped cells, consisting of one a cell (red) connected to one α cell (green) via a conjugation tube (Figure 3A, panel 1). In contrast, no fusants could be identified in crosses between wild type a cells (red) and any modified α cells (green) (panels 2–4). Specifically, in crosses between a cells and αmfαΔ cells, the αmfαΔ cells responded to the presence of a cells by forming conjugation tubes; however, in the absence of α pheromone production, the frequency of fusion with a cells is predicted to be very low, and consistent with this expectation, we did not observe any fusants. Accordingly, in strains no longer harboring the STE3α receptor (αmfαΔ,STE3a and αa), the cells appear to neither form conjugation tubes (in the presence of a cells) nor fuse with a cells (panels 3 & 4). These results indicate that modified α strains, in which the STE3α receptor gene has been deleted, do not respond to or mate with a cells.
Fig. 3. αa strains fuse with wild-type α cells. 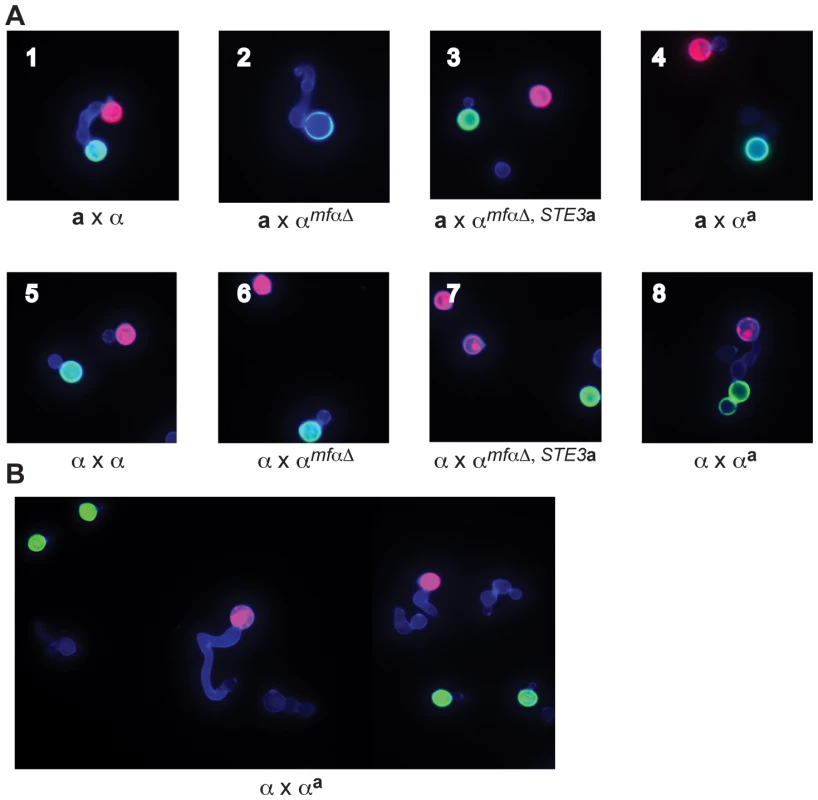
(A) Sexual development assays were carried out on V8 media using differentially labeled a and α strains (red or green) to detect the formation of germ tubes (by α cells) and fusants (dumbbell shaped structures). A representative view from the indicated cross is displayed in each panel (1-8). In panels 1–4, a cells are labeled red, and α cells are labeled green. Panel 1: wild type a x α fusant. Panel 2: germ tube from an αmfαΔ cell; no fusants. Panels 3 and 4: no germ tubes; no fusants. In panel 5, wild type α cells are labeled red or green. In panels 6–8, wild type α cells are labeled red and modified α cells are labeled green. Panels 5–7: no germ tubes; no fusants. Panel 8: wild type α x αa fusant. (B) αa cells do not form germ tubes. Wild type α cells were labeled red, and αa cells were labeled green. Wild type α cells (red) produce germ tubes in response to a cells (A, panel 1) and to αa cells (A, panel 8). Similar to wild-type a cells, αa cells do not form germ tubes in response to α cells. In parallel experiments, wild type α strains were crossed with the modified α strains. In a control cross, differentially labeled α cells (either red or green) were crossed and evaluated for the presence of fusants. As expected for wild type cells of the same mating type, α cells did not respond to one another or fuse (panel 5). Fusants were also not detected in crosses between wild type α cells (red) and the modified αmfαΔ and αmfαΔ,STE3a (green) strains (panels 6 & 7, respectively). It was not until a wild type α cell (red) was crossed with the altered αa strain (green) that fusants could be recovered at roughly wild type a x α levels (panel 8). Subsequent quantitative fusion assays confirmed that α x αa fusion levels are comparable to those of wild type a x α crosses (data not shown). This finding indicates that, as expected, pheromones and pheromone receptors are required for wild type fusion between a and α cells. It also reveals that the altered αa strain fuses only with wild type α cells, indicating that by simply replacing the α pheromones and receptors with those from a cells, cell identify was switched from α to a. Moreover, that change was sufficient to allow the altered αa cells to fuse with wild type α cells as though they were of opposite mating types.
In addition, the introduction of pheromone and pheromone receptor from a cells into α cells changes not only their ability to fuse but also the initial mating response. That is, αa strains do not produce germ tubes in response to wild type α cells, instead appearing to behave like wild type a cells (Figure 3B). These findings indicate that pheromones and pheromone receptors specifically control both the ability to sense a mating partner and the subsequent morphological response.
Complete sexual development between αa and wild-type α cells occurs only in the presence of both Sxi1α and Sxi2a
To assess the ability of the series of modified α strains to undergo sexual development, crosses were carried out and assessed for the production of filaments, basidia, and spores. In crosses between wild type a and α strains, florid filamentation, basidia formation, and sporulation were all visible at the periphery of the cross (Figure 4A, panel 1). However, when the modified α strains (αmfαΔ, αmfαΔ,STE3a, and αa) were crossed with either wild type a or α cells, none of the combinations resulted in wild type sexual development (panels 2–8). The a x αmfαΔ cross (panel 2) revealed a significant reduction in sexual development, and this result is consistent with previously published data [13]. In addition, aberrant filaments were observed in the α x αa cross (panel 8). This limited, mutant filamentation was very close in appearance to crosses between strains containing deletions of SXI2a (panel 9). In crosses between wild type α strains and sxi2aΔ mutants, the strains fuse at wild type levels; however, only aberrant filaments are formed, and they do not progress through sexual development [9]. This phenotype occurs because both Sxi1α and Sxi2a must be present to specify the fused state and initiate either diploid-specific or dikaryon-specific developmental programs. Because the products of an α by αa cross contain only the Sxi1α developmental regulator (and not the Sxi2a regulator found only in a cells), such crosses would not be expected to progress through sexual development and would be expected to exhibit a sxi2aΔ cross phenotype. This is, in fact, the case. Conversely, if the Sxi proteins, pheromone, and pheromone receptors were necessary and sufficient to specify cellular identify (dikaryon vs. haploid), then simply supplying the SXI2a developmental regulator to an α by αa cross would result in complete sexual development.
Fig. 4. α x αa crosses proceed through sexual development only in the presence of Sxi2a. 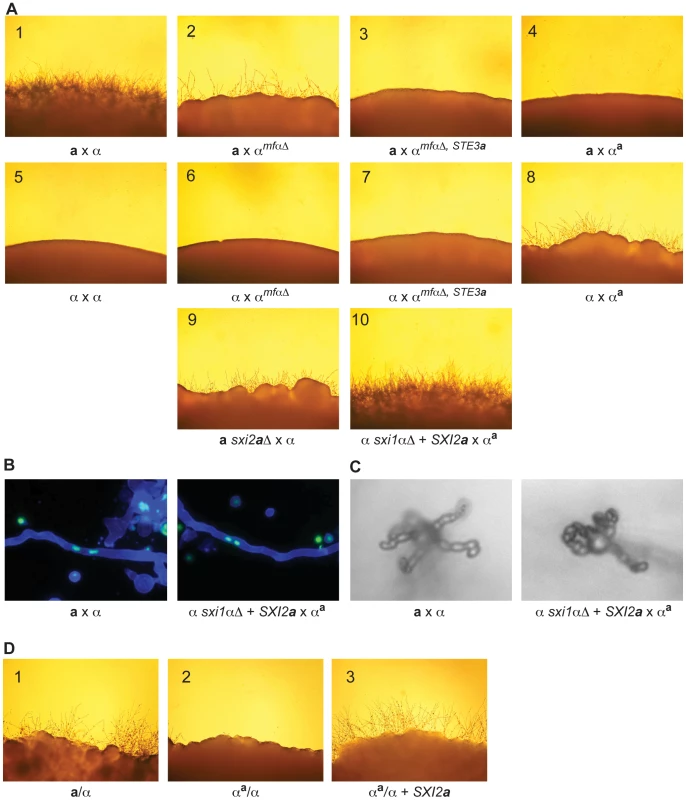
(A) Sexual development assays were carried out on V8 for 72 hours, and the peripheries of crosses are shown under 200× magnification. Panels: 1) a x α, 2) a x αmfαΔ, 3) a x αmfαΔ, STE3a, 4) a x αa, 5) α x α, 6) α x αmfαΔ, 7) α x αmfαΔ, STE3a, 8) α x αa, 9) a sxi2aΔ x α, 10) α sxi1αΔ + SXI2a x αa. Complete sexual development is observed only in panels 1 and 10. (B) Dikaryotic filaments are produced in the α sxi1αΔ + SXI2a x αa cross. Calcofluor stained filaments appear blue, and Sytox Green strained nuclei appear green. Both an a x α cross (left) and the α sxi1αΔ + SXI2a x αa cross (right) produce dikaryotic filaments (400× magnification). (C) Basidia and spores are produced in the α sxi1αΔ + SXI2a x αa cross. High resolution microscopy reveals the formation of basidia and spores from an a x α cross (left), and from the α sxi1αΔ + SXI2a x αa cross (right) (1000× magnification). (D) Addition of the SXI2a gene to αa/α diploids results in sexual development. Diploids were incubated on V8 for 72 hours, and test spot peripheries are shown under 200× magnification as follows: wild type a/α diploid (panel 1), αa/α diploid (panel 2), and αa/α + SXI2a (panel 3). To test the hypothesis that α x αa crosses were simply in need of SXI2a to continue through sexual development, we carried out crosses in which exogenous SXI2a was provided. Because strains containing both SXI1α and SXI2a are self-filamentous under sexual development conditions, we crossed the αa strain with an α sxi1αΔ strain carrying an integrated copy of SXI2a under the control of a constitutive promoter (GPD1). This cross resulted in the restoration of complete sexual development (panel 10). In addition, fluorescence and high-resolution microscopy show that dikaryotic filaments basidia, and spore chains were formed in αa x α + SXI2a crosses (Figures 4B, 4C). Because complete sexual development occurred between cells with nearly identical genomes (αa x α + SXI2a), no additional information from a cells is required for this process.
This finding was further supported by the results of assays for diploid formation. Previous studies have shown that a/α diploids are self-filamentous on V8 medium, undergoing sexual development [17]. To test the ability of the αa strain to form diploids, crosses of αa by wild type a or α cells were carried out. αa cells fused to form mononucleate cells (presumed diploids) with wild type α cells, but not with wild type a cells, consistent with αa cells exhibiting a cell behavior. In this assay, αa cells (ura5, NEOR) were mixed with either wild type α or a cells (URA5) and incubated on V8. After 24 hours at room temperature, the co-cultures were plated under double selection (Ura− + G418) at 37°C to select for mononucleate diploids [17]. No colonies grew from the a x αa crosses, indicating that no a/αa diploids were formed. In contrast, abundant colonies were recovered from the α x αa crosses, indicating that αa/α diploids were formed. The resulting αa/α strains were determined to be mononucleate (data not shown) and were evaluated for the ability to undergo sexual development by incubation on V8 at room temperature (Figure 4D). As in α x αa crosses, sexual development was not observed (panel 2); however, the addition of the SXI2a gene to the αa/α strain resulted in full sexual development on V8, similar to that of a wild type a/α diploid (panels 3 and 1, respectively). Taken together, these results demonstrate that cell identity and sexual development are controlled entirely by three kinds of genes in C. neoformans: pheromones, pheromone receptors, and homeodomain proteins.
The αa strain undergoes α fruiting
Monokaryotic or α fruiting is another form of sexual development that takes place in C. neoformans; however, this process is specific to α cells, which form filaments, basidia, and spores under severe nutrient limitation and desiccation [10],[18],[19]. Although fruiting is known to be an α specific process, the factors responsible for α fruiting are unknown. To assess whether changing haploid cell identity influences α fruiting, fruiting assays were carried out with wild type a, wild type α, and αa strains. Strains were incubated on filament agar for 14 days at room temperature in the dark and evaluated for the production of filaments and spores. We observed that the αa strain undergoes α fruiting that is indistinguishable from wild type α cells (Figure 5), indicating that this α-specific process does not require α-specific pheromones or pheromone receptor, and is not influenced by the presence of a-specific pheromones and pheromone receptor.
Fig. 5. αa cells undergo α fruiting. 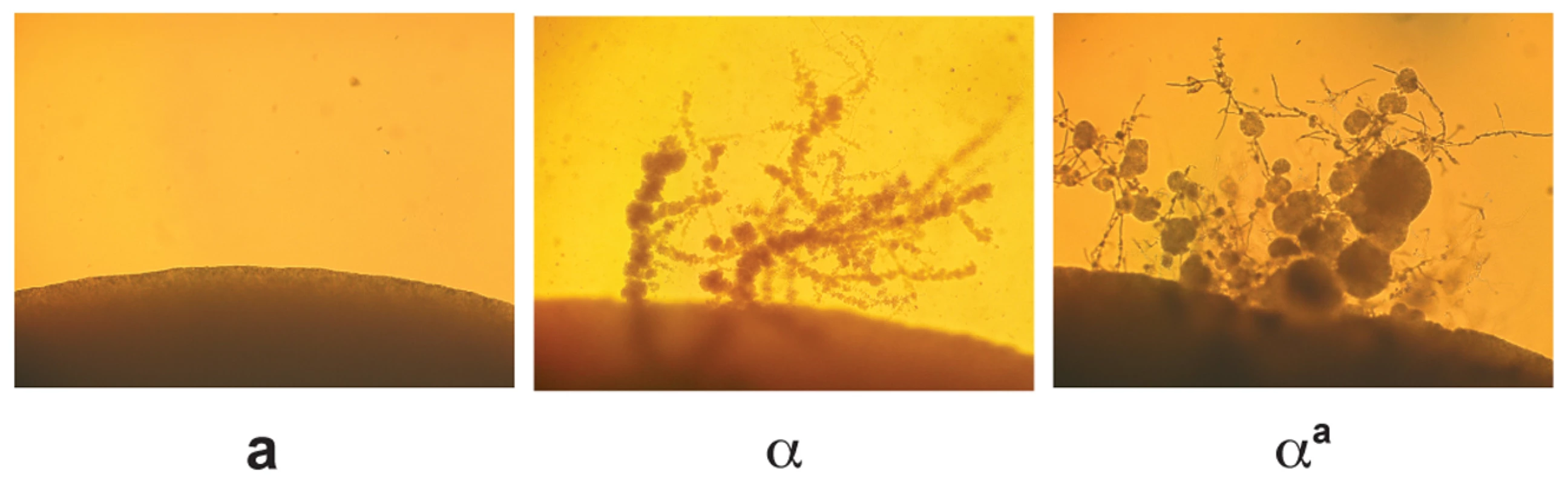
Fruiting assays were carried out on filament agar for 14 days at room temperature in the dark. The peripheries of colonies are shown at 100× magnification. Left panel: wild type a cells. Middle panel: wild type α cells. Right panel: αa cells. Discussion
Sexual development is an essential part of the life cycle for most eukaryotic organisms and requires mechanisms to ensure that development takes place between compatible partners. In both ascomycete and basidiomycete fungi, mating type is specified by components of the MAT locus [20]; however, the mechanisms by which MAT components act to confer cell identity vary widely among fungi [21],[22]. In this study we discovered that the basidiomycete C. neoformans establishes haploid cell type through the expression of mating type-specific pheromones and pheromone receptors. By replacing the pheromone and the pheromone receptor genes of α cells with those from a cells, we created an α cell with the haploid identity of an a. This altered strain exhibited the mating behaviors of an a cell, including the ability to fuse with an α mating partner. This result shows that no additional mating type-specific factors are necessary to denote haploid cell type. Furthermore, the addition of the a-specific factor SXI2a was then sufficient to facilitate filamentation and spore formation. Through these studies we have identified the factors that determine haploid cell identity (pheromones and their receptors), thus answering the question of how the MAT locus of C. neoformans controls cell type-specification.
Cell identity in fungi
There is a long history of investigating pheromone/pheromone receptor function in fungi, particularly in S. cerevisiae, and myriad experiments have been carried out to understand the roles of two phylogenetically unrelated receptors STE2 (α factor receptor) and STE3 (a factor receptor). Through altered expression experiments it was found that mating type is controlled by the identity of the expressed surface receptor. Experiments where the receptor expression patterns were reversed in a and α cells led to changes in mating behaviors. For example, α cells that express only STE2 (and not STE3), can sense and respond to α pheromone and therefore mate as a cells and vice versa. Thus, in S. cerevisiae, both a and α cells contain the STE2 and STE3 genes (and both pheromone genes), and it is the actions of transcription factors that impart differential expression patterns that determine cell identity [23],[24].
This is in sharp contrast to other fungi, like U. maydis, in which different cell types encode distinct pheromones and pheromone receptors. Experiments in U. maydis where the pheromones and pheromone receptors have been “swapped” between mating types demonstrate that mis-expression of pheromones and pheromone receptors is sufficient to alter cell identity. In this case, the pheromone receptors have distinct binding specificities but have descended from a common ancestor [4]. This is also the case for other basidiomycetes, including S. commune and C. cinerea. Within these fungi, different mating types encode distinct alleles of pheromones and pheromone receptors; pheromone and pheromone receptors do not govern mating, and cell fusion occurs independently of pheromone signaling. Instead, pheromones and pheromone receptors control post-fusion sexual development [25]–[27]. These examples speak to the wide variety of strategies employed by pheromones and pheromone receptors to control sexual development in fungi and emphasize the need to investigate pheromone/pheromone receptor function on a case-by-case basis.
C. neoformans pheromones and pheromone receptors
In C. neoformans, both a and α cells contain mating type-specific pheromone genes (MFa1, 2, and 3 in a cells, and MFα1, 2, and 3 in α cells), but the regulation of these genes differs between mating types. In a cells, MFa pheromone is highly induced on V8 agar (or under conditions of nutrient limitation) in the absence of a mating partner. In addition to induction under nutrient limiting conditions, MFα pheromone expression is also activated by factors secreted by a cells (MFa pheromone) [13], and expression of the MFα genes is dramatically upregulated upon exposure to synthetic MFa pheromone [12],[13].
In the experiments carried out here, expression of the MFa1 gene in αa cells is driven by the MFα1 promoter. The MFa1 construct was originally engineered in this fashion to avoid any cell type-specific regulation that might be imposed upon an a-specific promoter in an α cell. However, in αa strains where MFa1 gene expression was driven by the MFa1 promoter, identical phenotypes and expression patterns were observed.
Attempts to test the roles of the pheromone receptors were not as straight forward, and initially, the STE3a cassette was transformed into a wild type α strain to replace the STE3α gene at its native locus. However, after multiple attempts, no transformants were recovered (Ekena, Giles, and Hull, unpublished data). Transformants were recovered only from strains in which the pheromone genes were deleted previously. These findings, while far from conclusive, intimate that cells expressing both STE3a and α pheromone may engage in autocrine signaling that affects cell growth.
A similar observation has been made in S. cerevisiae, in which a strain that expresses both a pheromone and its cognate receptor undergoes G1 cell cycle arrest [24]. In S. cerevisiae this arrest is transient; however, if a similar arrest is occurring in C. neoformans, the cells do not appear to recover. This is in contrast to U. maydis where cells that have been engineered to express both receptor alleles do not arrest, are fully viable, and are mating competent. It remains to be determined if and how autocrine signaling may be functioning in C. neoformans.
Receptor identity influences mating morphology but not fruiting morphology
In C. neoformans, a and α cells carry out distinct functions to initiate the mating process. For example, in response to nitrogen limitation, a cells secrete MFa pheromone, and α cells respond to the presence of the MFa pheromone by formation of germ tubes. This morphological response is specific to α cells, and only these cells (and not a cells) form germ tubes as an initial mating response [11],[28],[29]. Interestingly, αa cells not only recognize and fuse with α cells (as would be expected for an a cell), but they also exhibit the morphology of an a cell, and do not produce germ tubes. This finding indicates that the morphological response to pheromone is mediated by the pheromone receptor (not downstream signaling events) and is independent of other mating type-specific factors. Therefore, pheromones and pheromone receptors alone dictate both the detection of a mating partner and the initial morphological response of haploid cells in C. neoformans. Interestingly, however, changes in mating identity did not influence the α-specific process of α fruiting.
Monokaryotic (or α) fruiting is a process that is specific to α cells and was recently shown to be a form of sexual development [19]. While the mechanisms underlying α fruiting are relatively undefined, it is well established that the α fruiting pathway is independent of Sxi1α and Sxi2a, and that this process is attenuated in strains in which STE3α or the MFα genes have been deleted [12],[13],[19]. Our data reveal that although α fruiting is primarily an α-specific process, it occurs regardless of whether pheromones and pheromone receptor are derived from a or α cells.
In summary, while the α fruiting process occurs despite the identity of pheromones and their receptors, these factors are wholly responsible for determining the initial mating response and haploid cell identity. The findings presented here establish a complete cell identity determination profile for C. neoformans and enhance our understanding of the myriad strategies by which fungi specify mating types and undergo sexual development.
Materials and Methods
Constructing modified α strains
To generate the αmfαΔ, STE3a strain (CHY1900), the αmfαΔ (WSC17) strain was biolistically transformed with a fragment containing STE3a and a nourseothricin resistance (NATR) cassette that was flanked by ∼1 kb of sequence from upstream and downstream of the STE3α open reading frame (ORF) [13]. Nourseothricin resistant transformants were screened by PCR for proper integration of the deletion/insertion construct into the STE3α locus, and putative deletion/insertion strains were confirmed by Southern blot analysis [30]. To generate the αa strain (CHY1901), the αmfαΔ, STE3a strain was biolistically transformed with a randomly integrating fragment of DNA containing MFa1 (under the control of the MFα1 promoter) and a neomycin resistance (NEOR) cassette. Neomycin resistant transformants were screened for copy number of the insertion using Southern blot analysis [30]. The αa/α strain (CHY2049) was generated by crossing the 5-FOA resistant αa strain (CHY1517) by wild type α cells (JEC21) and selecting for fusants at 37°C on minimal medium containing neomycin. To generate the αa/αSXI2a strain (CHY2096), 5-FOA resistant αa/α cells were biolistically transformed with a randomly integrating fragment of DNA containing the SXI2a (under the control of the GPD1 promoter) and URA5 genes.
Northern blot analysis
RNA was prepared from C. neoformans cells using a hot phenol extraction [30]. Strains were grown on solid V8 medium for 24 hours at room temperature. Northern blots were carried out according to standard protocols with 10 µg of total RNA used for each sample [30]. The glycerol-3-phosphate dehydrogenase gene (GPD1) probe was PCR-generated using CHO651 (CGTCGTTGAATCTACCGGTG) & CHO652 (CACCAGCAATGTAAGAGATG). All other probes were generated by PCR using the following oligonucleotides: CHO2030 (CCCCGACTATCCCTTTTGGAATCTCACTGC) & CHO2031 (GGCGAACAGTTCTTCGGGATATTGTGATACC) for STE3a, CHO2032 (GCACGACCTCAGCCTCGTCATTTTCAGCGG) & CHO2033 (CCGTATCCAGCAGCAATGATCGTCAGC) for STE3α, CHO2052 (GACTTACTCTTGCGTTATTGCTTAAAGTGGG) & CHO1917 (GAAAAGAGGTACGAGTAGAT) for MFa1, and CHO2053 (TTGTGTCATCGCCTAGACCCAACGTCCCC) & CHO2054 (CCATCTAAACAAGTCCCATACGCTTCGTTACC) for MFα1. Radiolabeled probes (Decaprime II kit; Ambion) were used in hybridization reactions as described previously [31]. Hybridizations and washes were carried out at 65°C as described previously [30].
Strain manipulations and media
All strains used were of the serotype D background (Table 1) and were handled using standard techniques and media as described previously [32],[33]. Sexual development assays were conducted on 5% V8 medium at room temperature in the dark for 2–4 days and were evaluated by observing the periphery of test spots. The mating tester strains used were JEC20 (a) and JEC21 (α), and crosses were photographed at 100X magnification [18]. Spores were photographed at 400X magnification. Fruiting assays were carried out by growing cells on filament agar (0.67% yeast nitrogen base (without amino acids or ammonium sulfate) containing 100 nM (NH4)2SO4 at room temperature in the dark for 14 days. The periphery of test spots were photographed at 100X magnification.
Microscopy and staining
Fluorescent microscopy was carried out on a Zeiss Axioskop 2 fluorescent microscope fitted with an Axiocam MRM REV3 digital camera and corresponding AV4 software. Light microscopy was carried out using a Zeiss Axioplan microscope fitted with a 10X long working distance objective. Photographs were taken with a Nikon Coolpix 5400 camera mounted on the microscope. To label fusants, a and α cells were grown to stationary phase in yeast extract peptone dextrose (YPD) liquid medium and were suspended in 1X phosphate buffered saline (PBS). Carboxylic acid, succinimidyl-ester Alexa Fluor 488 or 594 dyes (Invitrogen, Carlsbad CA) were coupled to a or α yeast cells in 100 mM potassium phosphate buffer pH 7.5 for 30 min at room temperature. After washing, differentially labeled cells were mixed in 1X PBS and spotted onto V8 agar. Spots were incubated at room temperature for 6 hours and were then resuspended into a mounting solution of 1X PBS containing 0.4 µg/ml calcofluor white MR2 (Sigma-Aldrich). For filament staining, crosses were incubated on a thin layer of V8 media on a microscope slide and incubated at room temperature for 24 hours. Cells were fixed with 2.5% gluteraldehyde/4% paraformaldehyde for 30 minutes. Cells were washed twice with 1X PBS containing 0.1% Triton X-100, once with 1X PBS, and incubated in 1X PBS containing 0.4 µg/mL calcofluor white to stain septa, and 1nM Sytox green (Invitrogen) to stain nuclei for 20 minutes. Cells were washed in 5% glycerol with 20% DABCO antifade (Sigma) prior to visualization.
Zdroje
1. HaqqCM
DonahoePK
1998 Regulation of sexual dimorphism in mammals. Physiol Rev 78 1 33
2. JohnsonAD
1995 Molecular mechanisms of cell-type determination in budding yeast. Curr Opin Genet Dev 5 552 558
3. HerskowitzI
1989 A regulatory hierarchy for cell specialization in yeast. Nature 342 749 757
4. BolkerM
UrbanM
KahmannR
1992 The a mating type locus of U. maydis specifies cell signaling components. Cell 68 441 450
5. BanuettF
1992 Ustilago maydis, the delightful blight. Trends Genet 8 174 180
6. BanuettF
HerskowitzI
1989 Different a alleles of Ustilago maydis are necessary for maintenance of filamentous growth but not for meiosis. Proc Natl Acad Sci U S A 86 5878 5882
7. LengelerKB
FoxDS
FraserJA
AllenA
ForresterK
2002 Mating-type locus of Cryptococcus neoformans: a step in the evolution of sex chromosomes. Eukaryot Cell 1 704 718
8. HsuehYP
FraserJA
HeitmanJ
2008 Transitions in sexuality: recapitulation of an ancestral tri - and tetrapolar mating system in Cryptococcus neoformans. Eukaryot Cell 7 1847 1855
9. HullCM
BoilyMJ
HeitmanJ
2005 Sex-specific homeodomain proteins Sxi1α and Sxi2a coordinately regulate sexual development in Cryptococcus neoformans. Eukaryot Cell 4 526 535
10. HullCM
DavidsonRC
HeitmanJ
2002 Cell identity and sexual development in Cryptococcus neoformans are controlled by the mating type-specific homeodomain protein Sxi1α. Genes & Dev 16 3046 3060
11. MooreTDE
EdmanJC
1993 The α-mating type locus of Cryptococcus neoformans contains a peptide pheromone gene. Mol Cell Biol 13 1962 1970
12. ChungS
KarosM
ChangYC
LukszoJ
WickesBL
2002 Molecular analysis of CPRα, a MATα-specific pheromone receptor gene of Cryptococcus neoformans. Eukaryot Cell 1 432 439
13. ShenWC
DavidsonRC
CoxGM
HeitmanJ
2002 Pheromones stimulate mating and differentiation via paracrine and autocrine signaling in Cryptococcus neoformans. Eukaryot Cell 1 366 377
14. McClellandCM
FuJ
WoodleeGL
SeymourTS
WickesBL
2002 Isolation and characterization of the Cryptococcus neoformans MATa pheromone gene. Genetics 160 935 947
15. DavidsonRC
MooreTD
OdomAR
HeitmanJ
2000 Characterization of the MFα pheromone of the human fungal pathogen Cryptococcus neoformans. Mol Microbiol 38 1017 1026
16. ChangYC
MillerGF
Kwon-ChungKJ
2003 Importance of a developmentally regulated pheromone receptor of Cryptococcus neoformans for virulence. Infect Immun 71 4953 4960
17. SiaRA
LengelerKB
HeitmanJ
2000 Diploid strains of the pathogenic basidiomycete Cryptococcus neoformans are thermally dimorphic. Fungal Genet Biol 29 153 163
18. WickesBL
MayorgaME
EdmanU
EdmanJC
1996 Dimorphism and haploid fruiting in Cryptococcus neoformans: association with the α-mating type. Proc Natl Acad Sci U S A 93 7327 7331
19. LinX
HullCM
HeitmanJ
2005 Sexual reproduction between partners of the same mating type in Cryptococcus neoformans. Nature 434 1017 1021
20. FraserJA
HeitmanJ
2003 Fungal mating-type loci. Curr Biol 13 R792 795
21. VaillancourtLJ
RaperCA
1996 Pheromones and pheromone receptors as mating-type determinants in basidiomycetes. Genet Eng (N Y) 18 219 247
22. KronstadJW
StabenC
1997 Mating type in filamentous fungi. Annu Rev Genet 31 245 276
23. BenderA
Sprague JGF
1986 Yeast peptide pheromones, a-factor and α-factor, activate a common response mechanism in their target cells. Cell 47 929 937
24. NakayamaN
MiyajimaA
AraiK
1987 Common signal transduction system shared by STE2 and STE3 in haploid cells of Saccharomyces cerevisiae: autocrine cell-cycle arrest results from forced expression of STE2. Embo J 6 249 254
25. O'SheaSF
ChaurePT
HalsallJR
OlesnickyNS
LeibbrandtA
1998 A large pheromone and receptor gene complex determines multiple B mating type specificities in Coprinus cinereus. Genetics 148 1081 1090
26. VaillancourtLJ
RaudaskoskiM
SpechtCA
RaperCA
1997 Multiple genes encoding pheromones and a pheromone receptor define the Bβ1 mating-type specificity in Schizophyllum commune. Genetics 146 541 551
27. WendlandJ
VaillancourtLJ
HegnerJ
LengelerKB
LaddisonKJ
1995 The mating-type locus Bα1 of Schizophyllum commune contains a pheromone receptor gene and putative pheromone genes. EMBO J 14 5271 5278
28. WangP
PerfectJR
HeitmanJ
2000 The G-protein beta subunit GPB1 is required for mating and haploid fruiting in Cryptococcus neoformans. Mol Cell Biol 20 352 362
29. AlspaughJA
DavidsonRC
HeitmanJ
2000 Morphogenesis of Cryptococcus neoformans. Contrib Microbiol 5 217 238
30. AusubelF
BrentR
KingstonR
MooreD
SeidmanJ
1997 Current Protocols in Molecular Biology. Boston, MA John Wiley and Sons, Inc
31. ChurchGM
GilbertW
1984 Genomic sequencing. Proc Natl Acad Sci USA 81 1991 1995
32. AlspaughJA
PerfectJR
HeitmanJ
1998 Signal transduction pathways regulating differentiation and pathogenicity of Cryptococcus neoformans. Fungal Genet Biol 25 1 14
33. ShermanF
FinkGR
HicksJB
1986 Laboratory Course Manual for Methods in Yeast Genetics: Cold Spring Harbor Laboratory. 186
Štítky
Genetika Reprodukční medicína
Článek Nuclear Pore Proteins Nup153 and Megator Define Transcriptionally Active Regions in the GenomeČlánek Deletion of the Huntingtin Polyglutamine Stretch Enhances Neuronal Autophagy and Longevity in MiceČlánek Analysis of the Genome and Transcriptome Uncovers Unique Strategies to Cause Legionnaires' DiseaseČlánek Population Genomics of Parallel Adaptation in Threespine Stickleback using Sequenced RAD TagsČlánek Wing Patterns in the Mist
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2010 Číslo 2
-
Všechny články tohoto čísla
- Nuclear Pore Proteins Nup153 and Megator Define Transcriptionally Active Regions in the Genome
- The Scale of Population Structure in
- Allelic Exchange of Pheromones and Their Receptors Reprograms Sexual Identity in
- Genetic and Functional Dissection of and in Age-Related Macular Degeneration
- A Single Nucleotide Polymorphism within the Acetyl-Coenzyme A Carboxylase Beta Gene Is Associated with Proteinuria in Patients with Type 2 Diabetes
- The Genetic Interpretation of Area under the ROC Curve in Genomic Profiling
- Genome-Wide Association Study in Asian Populations Identifies Variants in and Associated with Systemic Lupus Erythematosus
- Cdk2 Is Required for p53-Independent G/M Checkpoint Control
- Uncoupling of Satellite DNA and Centromeric Function in the Genus
- Genomic Hotspots for Adaptation: The Population Genetics of Müllerian Mimicry in the Clade
- Use of DNA–Damaging Agents and RNA Pooling to Assess Expression Profiles Associated with and Mutation Status in Familial Breast Cancer Patients
- Cheating by Exploitation of Developmental Prestalk Patterning in
- Replication and Active Demethylation Represent Partially Overlapping Mechanisms for Erasure of H3K4me3 in Budding Yeast
- Cdk1 Targets Srs2 to Complete Synthesis-Dependent Strand Annealing and to Promote Recombinational Repair
- A Genome-Wide Association Study Identifies Susceptibility Variants for Type 2 Diabetes in Han Chinese
- Genome-Wide Identification of Susceptibility Alleles for Viral Infections through a Population Genetics Approach
- Transcriptional Rewiring of the Sex Determining Gene Duplicate by Transposable Elements
- Genomic Hotspots for Adaptation: The Population Genetics of Müllerian Mimicry in
- Proteasome Nuclear Activity Affects Chromosome Stability by Controlling the Turnover of Mms22, a Protein Important for DNA Repair
- Deletion of the Huntingtin Polyglutamine Stretch Enhances Neuronal Autophagy and Longevity in Mice
- Structure, Function, and Evolution of the spp. Genome
- Human and Non-Human Primate Genomes Share Hotspots of Positive Selection
- A Kinase-Independent Role for the Rad3-Rad26 Complex in Recruitment of Tel1 to Telomeres in Fission Yeast
- Analysis of the Genome and Transcriptome Uncovers Unique Strategies to Cause Legionnaires' Disease
- Molecular Evolution and Functional Characterization of Insulin-Like Peptides
- Molecular Poltergeists: Mitochondrial DNA Copies () in Sequenced Nuclear Genomes
- Population Genomics of Parallel Adaptation in Threespine Stickleback using Sequenced RAD Tags
- Wing Patterns in the Mist
- DNA Binding of Centromere Protein C (CENPC) Is Stabilized by Single-Stranded RNA
- Genome-Wide Association Study Reveals Multiple Loci Associated with Primary Tooth Development during Infancy
- Mutations in , Encoding an Equilibrative Nucleoside Transporter ENT3, Cause a Familial Histiocytosis Syndrome (Faisalabad Histiocytosis) and Familial Rosai-Dorfman Disease
- Genome-Wide Identification of Binding Sites Defines Distinct Functions for PHA-4/FOXA in Development and Environmental Response
- Ku Regulates the Non-Homologous End Joining Pathway Choice of DNA Double-Strand Break Repair in Human Somatic Cells
- Nucleoporins and Transcription: New Connections, New Questions
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Genome-Wide Association Study in Asian Populations Identifies Variants in and Associated with Systemic Lupus Erythematosus
- Nucleoporins and Transcription: New Connections, New Questions
- Nuclear Pore Proteins Nup153 and Megator Define Transcriptionally Active Regions in the Genome
- The Genetic Interpretation of Area under the ROC Curve in Genomic Profiling
Přihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání




