-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaRumors of Its Disassembly Have Been Greatly Exaggerated: The Secret Life of the Synaptonemal Complex at the Centromeres
article has not abstract
Published in the journal: . PLoS Genet 8(6): e32767. doi:10.1371/journal.pgen.1002807
Category: Perspective
doi: https://doi.org/10.1371/journal.pgen.1002807Summary
article has not abstract
The synaptonemal complex (SC) and the centromeres have long been thought to play their roles at quite different, non-overlapping periods during meiosis. The SC is viewed as a hallmark structure that connects homologous chromosomes and is functionally critical for early prophase (zygotene and pachytene), while centromeres are known to assemble the kinetochore, act as a target for microtubules, and direct chromosome movement during anaphase I and II.
However, several recent studies have suggested much earlier functions for centromeres (at least in some organisms) and detailed post-pachytene functions of the SC. First, studies in budding yeast, Drosophila, and higher plants provide evidence that centromeres play an important role in the initiation of synapsis in these systems [1]–[5]. Second, building on earlier work [6], [7], recent studies have clearly demonstrated that in both yeast and flies the SC persists at the centromeres long after the end of pachytene—at least until late prophase [1], [8], [9]—and that these regions of centromeric pairing and synapsis play important roles in mediating segregation at anaphase I (see below). In this issue of PLoS Genetics, two studies of mammalian spermatocytes [10], [11] demonstrate that while the centromeres are not involved in synaptic initiation in mammals, stretches of the SC at paired centromeres and at sites of crossing over do indeed persist beyond the breakdown of synapsis along the chromosome arms at the end of pachytene (see Figure 1).
Fig. 1. Synapsis and centromere pairing in mouse spermatocytes. 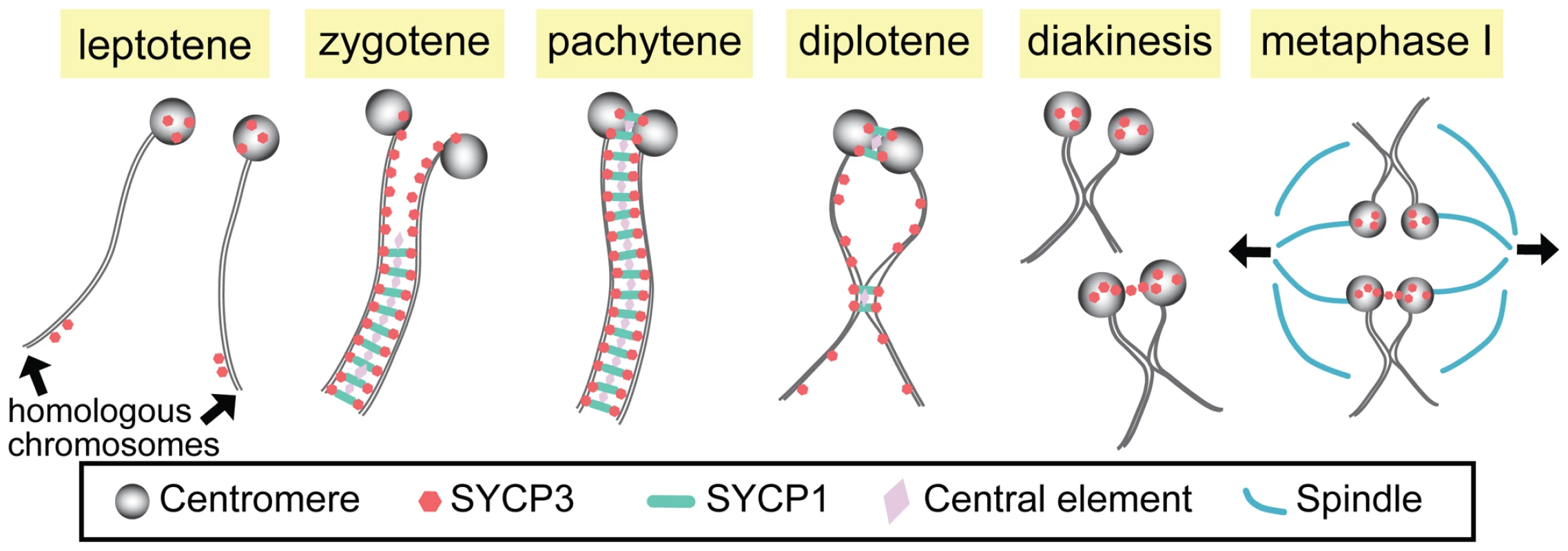
Homologous chromosomes are not paired in leptotene, but SYCP3 starts to localize to centromeres and arms. In zygotene, synapsis initiates from the non-centromeric regions and progresses as SCYP1 and central element proteins zip up SYCP3-containing chromosome axes. At late zygotene, centromeres are at last paired and synapsed, and full SC is formed along the entire chromosome in pachytene. In diplotene, the SC disassembles except at the sites where centromere pairing persists and chiasmata are formed. At diakinesis/metaphase I, SYCP3 proteins are accumulated on the centromere and a small fraction of homologous centromeres are linked by SYCP3-containing stretches, which may function in bi-orientation of homologous centromeres and attachment to the spindles for the first meiotic division. The SC is a proteinaceous railroad-track–like structure that assembles in early meiotic prophase immediately following or concomitant with homolog pairing. This assembly process is often referred to as homolog synapsis. Although the SC is structurally complex, it may be thought of as consisting primarily of three components: lateral elements that run along the entire length of each homolog (and in mammals contain the SYCP3 protein); transverse filaments that, like the teeth of a zipper, serve to connect the two lateral elements of the SC (and include the mammalian SYCP1 protein); and a set of proteins located at the center of the SC (cleverly referred to as central element proteins).
The question thus arises as to the function of these short regions of SC that survive the end of pachytene. Studies in yeast have clearly shown that the maintenance of centromeric pairing by short regions of the SC plays a role in mediating the segregation of both exchange and non-exchange chromosome segregation at meiosis I [8], [9]. Similar data demonstrate that persistent pericentromeric pairings also play a critical role in mediating the segregation of non-exchange homologs in Drosophila [12], [13].
In this issue of PLoS Genetics, the Pezza and Hunter groups show that the SC at mammalian centromeres also persists beyond the breakdown of synapsis along the chromosome arms at the end of pachytene in mouse spermatocytes [10], [11]. Indeed, the centromeres are the last sites at which SC is observed. Both groups also demonstrate that the persistent pairing of centromeric regions is dependent on the transverse filament protein SYCP1. Moreover, Qiao et al. [11] characterize the structure of the SC at centromeres by structured illumination microscopy (SIM), revealing that the SC stretches at the centromere represent bona fide tripartite SC in which transverse filaments connect two lateral elements.
Unlike SC breakdown along the chromosome arms following the end of pachytene, the much later dissolution of the SC at the centromeres may be a temporally complex process in which at least one SC component remains at the centromeres. Consistent with previous observations, both Bisig et al. [10] and Qiao et al. [11] show that while SYCP1 becomes undetectable prior to Nuclear Envelope Breakdown, SYCP3 remains associated with the centromeres until at least anaphase I in males. Qiao et al. also suggest that in some cases the persistence of SYCP3 at the centromere may provide a “bridge between the two homologous kinetochores” at diakinesis/metaphase I. Studies in other organisms support the view that such SYCP3 bridges may play important roles in facilitating segregation of achiasmate sex chromosomes [14].
However, centromeres do not appear to function as synaptic initiation sites in mammals. Indeed, both Bisig et al. and Qiao et al. demonstrate that centromeres are the last to pair, doing so only at zygotene–pachytene transition. Qiao et al. also use Rnf212 (Zip3 homolog) knock-out mice to explore whether or not centromeres can be “forced” to become synaptic initiation sites. In yeast zip3 mutants, synapsis occurs predominantly at the centromeres [2]. In Rnf212−/− spermatocytes, however, synapsis does not initiate at centromeres even though SC-associated centromere pairing at diplotene in the mutants was similar to that observed in wild-type. These data show clearly that, unlike the situations in yeast, flies, and higher plants, centromeres in the mouse do not have the capacity to initiate synapsis—indeed, they are extremely delayed in terms of their ability to initiate synapsis.
Finally, Qiao et al. demonstrate that remnants of SC also persist beyond pachytene at the sites of chiasma formation where they may regulate the necessary local remodeling of the homolog axes. In the absence of SYCP1, chiasma-like structures are still observed at diplotene (despite the inability of this genotype to generate normal crossovers), but these aberrant structures are more numerous and often exhibit fused axial elements as well as fusions of telomeric axes. The authors interpret these data to mean that prior to chiasma formation, the SYCP3-based axes are stabilized by the presence of the central region of the SC.
The take-home messages from these stories are that the SC's functions don't end with the disappearance of the majority of the structure at the end pachytene, nor are they limited to synapsis and recombination. Such findings are a complement to the demonstration that in at least some organisms (although not in mammals) the centromeres act as sites of the synaptic initiation. Moreover, SC structure at the centromeres may be somewhat different from that of the so-called normal euchromatic SC, and at least some SC components (such as SYCP3) may have functions that persist even after dissolution of the complete SC. Thus the SC needs to be considered as a rather flexible and adaptable structure that functions far more widely (both temporally and mechanistically) than previously thought. Given their roles in synaptic initiation in some organisms, the same may be true of centromeres. It's a fascinating partnership.
Zdroje
1. TakeoSLakeCMMorais-de-SáESunkelCEHawleyRS 2011 Synaptonemal complex-dependent centromeric clustering and the initiation of synapsis in Drosophila oocytes. Curr Biol 21 1845 1851
2. TsubouchiTMacqueenAJRoederGS 2008 Initiation of meiotic chromosome synapsis at centromeres in budding yeast. Genes Dev 22 3217 3226
3. TannetiNSLandyKJoyceEFMcKimKS 2011 A pathway for synapsis initiation during zygotene in Drosophila oocytes. Curr Biol 21 1852 1857
4. Martínez-PérezEShawPReaderSAragón-AlcaideLMillerT 1999 Homologous chromosome pairing in wheat. J Cell Sci 112 1761 1769
5. StewartMNDawsonDS 2008 Changing partners: moving from non-homologous to homologous centromere pairing in meiosis. Trends Genet 24 564 573
6. HolmPBRasmussenSWZicklerDLuBCSageJ 1981 Chromosome pairing, recombination nodules and chiasma formation in the basidiomycete coprinus cinereus. Carlsberg Research Communications 46 305 346
7. ZicklerDKlecknerN 1999 Meiotic chromosomes: integrating structure and function. Annu Rev Genet 33 603 754
8. GladstoneMNObesoDChuongHDawsonDS 2009 The synaptonemal complex protein Zip1 promotes bi-orientation of centromeres at meiosis I. PLoS Genet 5 e1000771 doi:10.1371/journal.pgen.1000771
9. NewnhamLJordanPRockmillBRoederGSHoffmannE 2010 The synaptonemal complex protein, Zip1, promotes the segregation of nonexchange chromosomes at meiosis I. Proc Natl Acad Sci U S A 107 781 785
10. BisigCGGuiraldelliMFKouznetsovaAScherthanHHöögC 2012 Synaptonemal complex components persist at centromeres and are required for homologous centromere pairing in mouse spermatocytes. PLoS Genet 8 e1002701 doi:10.1371/journal.pgen.1002701
11. QiaoHChenJKReynoldsAHöögCPaddyM 2012 Interplay between synaptonemal complex, homologous recombination and centromeres during mammalian meiosis. PLoS Genet 8 e1002790 doi:10.1371/journal.pgen.1002790.
12. HughesSEGillilandWDCotittaJLTakeoSCollinsKA 2009 Heterochromatic threads connect oscillating chromosomes during prometaphase I in Drosophila oocytes. PLoS Genet 5 e1000348 doi:10.1371/journal.pgen.1000348.
13. DernburgAFSedatJWHawleyRS 1996 Direct evidence of a role for heterochromatin in meiotic chromosome segregation. Cell 86 135 146
14. de la FuenteRParraMTVieraACalventeAGomezR 2007 Meiotic pairing and segregation of achiasmate sex chromosomes in eutherian mammals: the role of SYCP3 protein. PLoS Genet 3 e198 doi:10.1371/journal.pgen.0030198.
Štítky
Genetika Reprodukční medicína
Článek Uracil-Containing DNA in : Stability, Stage-Specific Accumulation, and Developmental InvolvementČlánek Fuzzy Tandem Repeats Containing p53 Response Elements May Define Species-Specific p53 Target GenesČlánek Preferential Genome Targeting of the CBP Co-Activator by Rel and Smad Proteins in Early EmbryosČlánek Protective Coupling of Mitochondrial Function and Protein Synthesis via the eIF2α Kinase GCN-2Článek Cohesin Proteins Promote Ribosomal RNA Production and Protein Translation in Yeast and Human CellsČlánek TERRA Promotes Telomere Shortening through Exonuclease 1–Mediated Resection of Chromosome EndsČlánek Attenuation of Notch and Hedgehog Signaling Is Required for Fate Specification in the Spinal CordČlánek Genome-Wide Functional Profiling Identifies Genes and Processes Important for Zinc-Limited Growth ofČlánek MicroRNA93 Regulates Proliferation and Differentiation of Normal and Malignant Breast Stem Cells
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2012 Číslo 6
-
Všechny články tohoto čísla
- Rumors of Its Disassembly Have Been Greatly Exaggerated: The Secret Life of the Synaptonemal Complex at the Centromeres
- Mimetic Butterflies Introgress to Impress
- Tipping the Balance in the Powerhouse of the Cell to “Protect” Colorectal Cancer
- Selection-Driven Gene Loss in Bacteria
- Decreased Mitochondrial DNA Mutagenesis in Human Colorectal Cancer
- Parallel Evolution of Auditory Genes for Echolocation in Bats and Toothed Whales
- Diverse CRISPRs Evolving in Human Microbiomes
- The Rad4 ATR-Activation Domain Functions in G1/S Phase in a Chromatin-Dependent Manner
- Stretching the Rules: Monocentric Chromosomes with Multiple Centromere Domains
- Uracil-Containing DNA in : Stability, Stage-Specific Accumulation, and Developmental Involvement
- Fuzzy Tandem Repeats Containing p53 Response Elements May Define Species-Specific p53 Target Genes
- Adaptive Introgression across Species Boundaries in Butterflies
- G Protein Activation without a GEF in the Plant Kingdom
- Synaptonemal Complex Components Persist at Centromeres and Are Required for Homologous Centromere Pairing in Mouse Spermatocytes
- An Engineering Approach to Extending Lifespan in
- Incompatibility and Competitive Exclusion of Genomic Segments between Sibling Species
- Effects of Histone H3 Depletion on Nucleosome Occupancy and Position in
- Patterns of Evolutionary Conservation of Essential Genes Correlate with Their Compensability
- Interplay between Synaptonemal Complex, Homologous Recombination, and Centromeres during Mammalian Meiosis
- Preferential Genome Targeting of the CBP Co-Activator by Rel and Smad Proteins in Early Embryos
- A Mouse Model of Acrodermatitis Enteropathica: Loss of Intestine Zinc Transporter ZIP4 (Slc39a4) Disrupts the Stem Cell Niche and Intestine Integrity
- Protective Coupling of Mitochondrial Function and Protein Synthesis via the eIF2α Kinase GCN-2
- Geographic Differences in Genetic Susceptibility to IgA Nephropathy: GWAS Replication Study and Geospatial Risk Analysis
- Cohesin Proteins Promote Ribosomal RNA Production and Protein Translation in Yeast and Human Cells
- TERRA Promotes Telomere Shortening through Exonuclease 1–Mediated Resection of Chromosome Ends
- Stimulation of Host Immune Defenses by a Small Molecule Protects from Bacterial Infection
- A Broad Requirement for TLS Polymerases η and κ, and Interacting Sumoylation and Nuclear Pore Proteins, in Lesion Bypass during Embryogenesis
- Genome-Wide Identification of Ampicillin Resistance Determinants in
- The CCR4-NOT Complex Is Implicated in the Viability of Aneuploid Yeasts
- Gustatory Perception and Fat Body Energy Metabolism Are Jointly Affected by Vitellogenin and Juvenile Hormone in Honey Bees
- Genetic Variants on Chromosome 1q41 Influence Ocular Axial Length and High Myopia
- Is a Key Regulator of Pancreaticobiliary Ductal System Development
- The NSL Complex Regulates Housekeeping Genes in
- Attenuation of Notch and Hedgehog Signaling Is Required for Fate Specification in the Spinal Cord
- Dual-Level Regulation of ACC Synthase Activity by MPK3/MPK6 Cascade and Its Downstream WRKY Transcription Factor during Ethylene Induction in Arabidopsis
- Genome-Wide Functional Profiling Identifies Genes and Processes Important for Zinc-Limited Growth of
- Base-Pair Resolution DNA Methylation Sequencing Reveals Profoundly Divergent Epigenetic Landscapes in Acute Myeloid Leukemia
- MicroRNA93 Regulates Proliferation and Differentiation of Normal and Malignant Breast Stem Cells
- Phylogenomic Analysis Reveals Dynamic Evolutionary History of the Drosophila Heterochromatin Protein 1 (HP1) Gene Family
- Found: The Elusive ANTAR Transcription Antiterminator
- The Mutation in Chickens Constitutes a Structural Rearrangement Causing Both Altered Comb Morphology and Defective Sperm Motility
- Control of CpG and Non-CpG DNA Methylation by DNA Methyltransferases
- Polymorphisms in the Mitochondrial Ribosome Recycling Factor Compromise Cell Respiratory Function and Increase Atorvastatin Toxicity
- Gene Expression Profiles in Parkinson Disease Prefrontal Cortex Implicate and Genes under Its Transcriptional Regulation
- Global Regulatory Functions of the Endoribonuclease III in Gene Expression
- Extensive Evolutionary Changes in Regulatory Element Activity during Human Origins Are Associated with Altered Gene Expression and Positive Selection
- The Regulatory Network of Natural Competence and Transformation of
- Brain Expression Genome-Wide Association Study (eGWAS) Identifies Human Disease-Associated Variants
- Quantifying the Adaptive Potential of an Antibiotic Resistance Enzyme
- Divergence of the Yeast Transcription Factor Affects Sulfite Resistance
- The Histone Demethylase Jhdm1a Regulates Hepatic Gluconeogenesis
- RNA Methylation by the MIS Complex Regulates a Cell Fate Decision in Yeast
- Rare Copy Number Variants Observed in Hereditary Breast Cancer Cases Disrupt Genes in Estrogen Signaling and Tumor Suppression Network
- Genome-Wide Location Analysis Reveals Distinct Transcriptional Circuitry by Paralogous Regulators Foxa1 and Foxa2
- A Mutation Links a Canine Progressive Early-Onset Cerebellar Ataxia to the Endoplasmic Reticulum–Associated Protein Degradation (ERAD) Machinery
- The Mechanism for RNA Recognition by ANTAR Regulators of Gene Expression
- Limits to the Rate of Adaptive Substitution in Sexual Populations
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Rumors of Its Disassembly Have Been Greatly Exaggerated: The Secret Life of the Synaptonemal Complex at the Centromeres
- The NSL Complex Regulates Housekeeping Genes in
- Tipping the Balance in the Powerhouse of the Cell to “Protect” Colorectal Cancer
- Interplay between Synaptonemal Complex, Homologous Recombination, and Centromeres during Mammalian Meiosis
Přihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



