-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Kariérní portál
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaA Review of Bacteria-Animal Lateral Gene Transfer May Inform Our Understanding of Diseases like Cancer
Lateral gene transfer (LGT) from bacteria to animals occurs more frequently than was appreciated prior to the advent of genome sequencing. In 2007, LGT from bacterial Wolbachia endosymbionts was detected in ∼33% of the sequenced arthropod genomes using a bioinformatic approach. Today, Wolbachia/host LGT is thought to be widespread and many other cases of bacteria-animal LGT have been described. In insects, LGT may be more frequently associated with endosymbionts that colonize germ cells and germ stem cells, like Wolbachia endosymbionts. We speculate that LGT may occur from bacteria to a wide variety of eukaryotes, but only becomes vertically inherited when it occurs in germ cells. As such, LGT may happen routinely in somatic cells but never become inherited or fixed in the population. Lack of inheritance of such mutations greatly decreases our ability to detect them. In this review, we propose that such noninherited bacterial DNA integration into chromosomes in human somatic cells could induce mutations leading to cancer or autoimmune diseases in a manner analogous to mobile elements and viral integrations.
Published in the journal: . PLoS Genet 9(10): e32767. doi:10.1371/journal.pgen.1003877
Category: Review
doi: https://doi.org/10.1371/journal.pgen.1003877Summary
Lateral gene transfer (LGT) from bacteria to animals occurs more frequently than was appreciated prior to the advent of genome sequencing. In 2007, LGT from bacterial Wolbachia endosymbionts was detected in ∼33% of the sequenced arthropod genomes using a bioinformatic approach. Today, Wolbachia/host LGT is thought to be widespread and many other cases of bacteria-animal LGT have been described. In insects, LGT may be more frequently associated with endosymbionts that colonize germ cells and germ stem cells, like Wolbachia endosymbionts. We speculate that LGT may occur from bacteria to a wide variety of eukaryotes, but only becomes vertically inherited when it occurs in germ cells. As such, LGT may happen routinely in somatic cells but never become inherited or fixed in the population. Lack of inheritance of such mutations greatly decreases our ability to detect them. In this review, we propose that such noninherited bacterial DNA integration into chromosomes in human somatic cells could induce mutations leading to cancer or autoimmune diseases in a manner analogous to mobile elements and viral integrations.
Introduction
Many eukaryotic chromosomes contain DNA of microbial origin that arose via lateral gene transfer (LGT). One such example is the directed transfer of Agrobacterium tumefaciens DNA that results in crown gall disease in plants and that has been used to create transgenic crops. A. tumefaciens specifically transfers 10–30 kbp of DNA from its 200–800 kbp tumor-inducing (Ti) plasmid to plants via the bacterial type IV secretion system [1]. The DNA from the Ti plasmid (T-DNA) is targeted to the nucleus, incorporated into the plant chromosome by illegitimate recombination, and transcribed from eukaryotic promoters in the T-DNA [2], [3]. LGT events are not limited to occurring between microbes and plants, but can also occur between microbes and animals. In this review, we synthesize our current understanding of the potential for LGT from bacteria to the somatic human genome by examining (a) LGT in animals with a particular emphasis on bacteria-animal LGT, (b) insertional mutagenesis in the human genome, and (c) the role of microbes in oncogenesis. Current and future work is then presented through two hypotheses about such integrations and their potential role in bacteria-associated chronic human diseases like cancer. Such transfers in the human genome may have been missed previously because they would not be inherited, and until recently, most LGT research has focused solely on the inherited consensus genome (e.g., [4]–[6]). However, important mutations are not limited to merely the vertically inherited genome. For example, several recent studies (e.g., [7]–[10]) have shown that the somatic genome can carry important novel mutations related to disease.
LGT in Animals
An Overview of LGT in Animals
Since plant germ cells are not physically protected, A. tumefaciens–mediated LGT in plants may be expected to occur more frequently when compared to inherited LGT in vertebrate animals where germ cells are protected. Yet, LGT can be detected in many animals, including vertebrates [11], [12]. For example, phylogenetically related antifreeze proteins in fish are scattered across disparate fish taxa, indicating a role for lateral gene transfer [13]. These proteins allow fish to survive at temperatures below freezing by preventing the formation of ice crystals [13]. Pea aphids and the two-spotted spider mite have both been found to synthesize carotenoids from an LGT that may have originated from fungi [14], [15]. In aphids, the resulting red-green color polymorphism changes the insect's susceptibility to natural enemies [14], while in the spider mite these carotenoid biosynthetic genes are differentially expressed in diapause [15], the arthropod equivalent to hibernation.
Overview of Functional Bacteria-Animal LGT
In addition to the eukaryote-eukaryote transfers described above, several functional LGTs have been described between bacteria and invertebrate animals. For example, in mealybugs, LGTs from at least three different bacterial lineages have resulted in hybrid biosynthetic pathways [16]. These pathways are composed of genes of eukaryotic ancestry in the mealybug genome, genes of bacterial ancestry in the mealybug genome, and genes of bacterial ancestry in the obligate endosymbiont bacterial genome [16]. For example, riboflavin biosynthesis requires one endosymbiont genome–encoded protein and three insect genome–encoded proteins that arose via LGT that have α-Proteobacteria and Bacteroidetes ancestry [16]. The bacterial donors of these bacteria-eukaryote LGTs are proposed to be facultative bacterial symbionts, not the primary obligate endosymbionts [16].
Bacteria-animal LGTs have also been observed in multiple agricultural pests where the LGT facilitates parasitization of an agricultural crop. For example, Hypothenemus hampei, the coffee berry borer, acquired a Bacillus mannanase gene, which allowed it to exploit coffee berries as a new ecological niche relative to the insect's sister taxa [17]. Several plant parasitic nematodes have acquired plant cell wall–degrading enzymes from bacteria that allow the nematodes to invade plant tissues and exploit plants as a new ecological niche. More specifically, LGT followed by duplication resulted in 60 genes of bacterial origin in the nematode Meloidogyne incongita genome, including cellulases, pectate lyases, and expansin-like proteins that degrade or modify plant cell walls and are not typically found in other animals [18]. These proteins have been biochemically characterized, are secreted into plant tissues, and are involved in parasitism [18]. M. incongita also acquired cellulase genes via an additional independent LGT by Pristionchus nematodes [19], [20], necromenic nematodes that live in association with beetles.
A Case Study: Wolbachia Endosymbionts Have Recently Transferred DNA to Multiple Invertebrate Host Genomes
In contrast to the examples above, where single genes were transferred, are functional, and have persisted over time, Wolbachia-insect LGTs span many hundreds of kilobases of DNA, have no evidence of being functional with no obvious change in insect phenotype, and are likely recent given that they have minimal divergence from the likely donor. Of the systems being studied today, recent LGT from bacteria to animals seems to be most prevalent between Wolbachia endosymbionts and their invertebrate hosts. In this case, the term gene is used loosely since a phenotype has not been ascribed to the transferred DNA. Wolbachia endosymbionts colonize a wide range of arthropods and filarial nematodes, including 20–70% of insect species, and are maternally inherited through the egg cytoplasm [21], [22]. Transmission through the egg cytoplasm provides ample opportunity for Wolbachia DNA to transfer to the host genome [11]. In addition, Wolbachia endosymbionts can sometimes colonize their host's germ stem cell [23]. Increased prevalence of LGT may be expected in hosts with colonized germ stem cells since a transfer in the germ stem cell will be inherited by a larger number of progeny in comparison with a transfer in a single gamete, like an ovum or spermatozoon.
Recent LGT from Wolbachia endosymbionts to their hosts has been characterized in diverse invertebrate hosts, including beetles [24]–[26], fruit flies [27], wasps [27], [28], tsetse flies [29], and filarial nematodes [27], [30], [31], and has been reviewed recently [11], [12]. In 2007, most of the genome sequencing projects (8/11) containing Wolbachia endosymbiont sequences showed evidence of having recent LGT between the endosymbiont genome and the host chromosome [27]. We successfully characterized LGT in all five of the hosts that we examined [27]. Therefore, we estimated that ∼70% of sequenced Wolbachia-infected hosts may have at least one Wolbachia-host LGT [27].
The high prevalence of Wolbachia-host LGT is not reflective of all microbe-host relationships. Many characteristics of their association have an effect on the frequency of LGT between the two organisms. One of the major factors limiting the occurrence of LGT is the proximity of the two organisms [32], which helps explain the increased prevalence of LGT between obligate intracellular endosymbionts and their hosts [11]. However, other factors are also likely at work, as evidenced in aphids where functional LGTs are not detected that arise from Buchnera aphidicola, the primary obligate endosymbionts of aphids [33], [34]. However, in aphids, functional LGT is detected from relatives of Wolbachia endosymbionts [33], [34]. Likewise, in the mealybug, LGT is attributed to facultative symbionts instead of the primary obligate endosymbionts [16]. This may suggest that LGT from Wolbachia endosymbionts and other similar bacteria (e.g., bacteria in the Arsenophonus and Cardinium clades) is more abundant for other reasons. For example, with Wolbachia endosymbionts, it may be because they do not merely pass through the germ cell, but rather colonize the germ cell and the germ stem cell. However, the characterization of transfers from Wolbachia endosymbionts to their hosts highlights that LGT is an ongoing process in at least some animals and that LGT can result in DNA transfers that are not functional and may never become functional.
An Open Question: Does Somatic LGT from Bacteria Occur in Humans?
Despite extensive microbe-animal LGT in invertebrates, bacterial LGT to humans and other mammals has rarely been described. One barrier to inherited LGT in humans is the segregation of gametes. Unlike in insects and plants, the human germ line is both physically and immunologically well protected from bacteria. Nevertheless, there are 10× more bacterial cells than human cells in the human body [35]. Therefore, somatic human cells can be bathed in bacteria and have ample opportunity to be mutagenized by bacterial DNA through LGT. Such LGT will not become inherited by offspring of the human, but may be propagated through the individual's lifetime if the cell is capable of undergoing clonal expansion. LGT to somatic tissues acting as a mutagen may therefore be important in bacteria-associated diseases like cancer, chronic inflammatory diseases, and autoimmune disease. Under this scenario, somatic LGT events would not enable adaptation to a new niche, but rather have the potential to be disruptive to normal gene function. While this review focuses on the potential for integrations of bacterial DNA in the somatic human genome, it is possible that DNA may also get integrated from the food we consume.
To protect human cells from bacteria, the innate immune system uses pattern recognition receptors (PRRs) to recognize pathogen-associated molecular patterns, or PAMPs, including LPS, flagellin, lipoteichoic acid, lipoproteins, and peptidoglycan [36]. Both microbial DNA and mRNA are also PAMPs in humans [37], [38], while a subset of microbial rRNA does not elicit an immune response [38]. Therefore, we might expect that bacterial rRNA is more likely to mutagenize the human somatic genome. This is a particularly interesting proposition considering some mobile elements in animals have rRNA as their origin and are mobilized by the LINE-1 machinery [39], [40].
We hypothesize that bacterial rRNA may integrate into the human somatic genome and induce disease through random mutagenesis, as do other insertion-creating mutagens, like mobile elements and viruses. Should this integrated DNA mutagenize a gene by disrupting the coding region or causing deregulation, the consequences could potentially be severe. For example, such insertional mutagenesis by mobile elements and viruses has been associated with cancer.
Insertional Mutagenesis of the Human Genome
The Role of Mobile Elements in Cancer Progression
Since mobile elements are already known to mutagenize the human genome, including cancer genomes, they provide a useful comparison for thinking about mutagenesis via LGT. Only the non–long terminal repeat (LTR) retrotransposons, including LINE and Alu elements, have been shown to actively jump throughout the human genome and thus cause ongoing mutagenesis [41]. LINE-1 (L1) is the only non-LTR retrotransposon that has the machinery to move itself in the human genome, and it also mobilizes Alu elements [41], [42]. L1s and Alu elements move to new genomic locations through germ line retrotransposition [42], [43], but both elements tend to be inactive in somatic tissues [41]. However, their reactivation may contribute to tumorigenesis [44].
L1s have been implicated in both colon and lung cancer. An L1 insertion was found in the APC gene in colon tumor cells but not in normal cells [45]. In colorectal cancer, tumors can be identified that have been mutagenized by L1s while matched normal DNA samples lack such mutations, and some insertions disrupt genes with known cancer driver functions [46]. Nine L1 insertions were found to be only in lung cancer tumors, but not normal adjacent tissue, with 30% of tumors having between one and three new insertions [47]. However, it is not clear if somatic L1 insertions are passenger mutations, or if they are directly related to tumor formation [47].
With >1 million copies in the human genome, it is not surprising that Alu insertions are also related to carcinogenesis. Alu insertions have been reported in the MLV12 and MLL genes associated with leukemia, the BRCA1 and BRCA2 genes associated with breast cancer, and the MLH1 and MSH2 genes leading to hereditary nonpolyposis colorectal cancer syndrome (HNPCC), among others [42]. The number of Alu and L1 integrations detected that are related to disease shows the potential importance of such integration events.
Viral Integrations and Oncogenesis
Viruses are also known to integrate into the somatic human genome and cause mutations that are associated with cancer. In 2008, it was estimated that 15–20% of cancers worldwide were linked to infections by viruses, parasites, or bacteria [48]. The integration of human papillomavirus (HPV) is a critical event leading to HPV-associated tumorigenesis and may be an important biomarker of invasive cervical cancer [49]. In the absence of integration, the virus replicates but the cell maintains control of its own proliferation. However, cellular proliferation becomes deregulated when HPV integrates in a manner that results in the loss of E1 and E2, viral proteins required for transcriptional control and replication. Absence of these regulator proteins leads to subsequent deregulation of E6 causing downregulation of the p53 pathway, which increases cell proliferation [50]. As many as 80–100% of cervical carcinoma tumors have an integration of HPV-16 or HPV-18 [51], [52] and these integrations are clonal within tumors [53], providing evidence that HPV acts as a carcinogen.
Hepatitis B virus (HBV) was linked to hepatocellular carcinoma (HCC) in 1981 [54], prompting its vaccine to be the first approved for cancer prevention [55]. HBV's exact role in formation of HCC is unclear despite the fact that its integrations can be found clonally in HCC tumors. Regardless, levels of HBV integration in tumor cells can predict patient survival. Individuals with >3 integrations have decreased survival compared to those with <3 integrations [10]. Seven human oncogenes and tumor suppressor genes have repeatedly been shown to contain HBV integrations, including TERT, ITPR1, IRAK2, MAPK1, MLL2, MLL4, and CCNE1 [10], [56]–[59]. This vast number of genes disrupted by HBV illustrates the multitude of ways that HBV can contribute to HCC carcinogenesis.
Mitochondrial Insertions and Disease
Integrations into the nuclear genome of human cells are not limited to mobile element and viral integrations. Transfers of mitochondrial DNA (mtDNA) into the nuclear genome have long been described and are called nuclear mitochondrial DNA segments (numts) [60]. There are various methods that have been suggested for how the mtDNA exits the mitochondria and enters the nucleus, including mitochondrial lysis, direct contact between the nucleus and the mitochondria, degradation of abnormal mitochondria, and mitochondrial DNA encapsulation inside the nucleus [61]. Once entrance into the nucleus has occurred, the mtDNA can integrate into the nuclear genome through nonhomologous end joining of double-strand break repair [61], [62].
While various numts have been reported, only a handful are implicated in disease. In one case, a reciprocal translocation occurred between chromosomes 9 and 11 resulting in a 41-bp insertion of mtDNA linking the breakpoint of chromosome 9 to the translocated portion of chromosome 11 [63]. There has been one report of a numt from the mitochondrial coxIII gene inserted in the c-myc gene of HeLa cells forming a chimeric RNA, but it is unknown if this numt contributed to carcinogenesis [64]. Other examples that link numts to human diseases are a 251-bp numt insertion into the human plasma factor VII gene causing severe factor VII deficiency [65], a 72-bp numt insertion in the GLI3 gene associated with Pallister-Hall syndrome [66], a 93-bp insertion into MCOLN1 related to mucolipidosis IV [67], and a 36-bp insertion into the USH1C gene implicated in Usher syndrome type IC [68], [69]. Thus far, most research on numts is focused on inherited mutations. While it has been noted that numts confound analyses of the mitochondrial genome in cancer samples [70], we are not aware of a published analysis of numts in the human somatic genome or cancer genomes, although it would be informative. An increase in somatic numts has been associated with aging in rats [71], and increases in both somatic numts and nuclear plastid–derived DNA, or nupts, have been associated with environmental stress in plants [72].
The Role of Microbes in Oncogenesis
Microbial Involvement in Tumorigenesis
Many bacterial associations with cancer have been characterized, including Helicobacter pylori with gastric carcinoma and gastric mucosa–associated lymphoid tissue (MALT) lymphoma [73], Escherichia coli with colorectal cancer [74], Schistosoma haematobium with bladder carcinoma [75], Bacteroides fragilis with colon cancer [76], and Fusobacterium spp. with colorectal cancer [77]. Many of these associations have been disputed, as it is often difficult to determine if an existing microbial infection is a symptom or a cause of cancer. Additionally, a single microbial species may merely be a marker for a more complex microbiome whose rare members contribute most substantially to oncogenesis, as has been observed in other systems [78]. Despite these associations between microbes and cancer, and the association of viral and mobile element integrations with cancer, microbial DNA integration has not been described for microbe-associated cancers. Prior to the widespread use of whole genome sequencing, the large size of microbial genomes made microbial integrations more difficult to detect. For example, viral integrations can be detected with Southern blots using viral-specific probes (e.g., [79]). The relatively larger genome size and thus higher complexity of microbial genomes likely precludes identifying integrations in the same manner without knowing something about the specific insert a priori. Sequencing reads that resemble bacterial DNA are sometimes removed from eukaryotic genome projects, further preventing identification of legitimate bacterial sequences in eukaryotic genomes. Instead, the microbial contribution to carcinogenesis is generally thought to occur via increased inflammation leading to DNA damage and secretion of bacterial effector proteins like toxins [80], [81].
Current and Future Directions
Hypothesis 1: Bacterial DNA Integration into Somatic Cells Could Induce Oncogenic Mutations
Bacterial integration that disrupts and mutagenizes proto-oncogenes or tumor suppressor genes could provide an additional avenue for bacteria-associated oncogenesis, beyond inflammation-induced damage. Such integrations could arise through a directed mechanism, as has been observed with Agrobacterium-induced crown gall disease. Alternatively, these integrations may merely result from the release of nucleic acids following lysis of bacteria. Integrations could occur specifically with particular integration sites as is observed with mobile elements or randomly in a manner analogous to mutations induced by exposure to known carcinogens, like UV radiation and cigarette smoke. Because bacterial DNA and mRNA are recognized by the human immune system [37], [38], we anticipate that most bacterial integrations will arise from nucleic acids that are not recognized as PAMPs, like some rRNA [38]. Given that L1 machinery can mobilize Alu elements as well as cellular genes [42], we hypothesize that the L1 machinery may also play a role in integrating bacterial rRNA into the somatic human genome. Although these events could lead to any type of genetic disorder, LGT events in cancer are easier to detect due to clonal expansion within the tumor (Figure 1).
Fig. 1. A scenario for bacteria-human LGT and carcinogenesis. 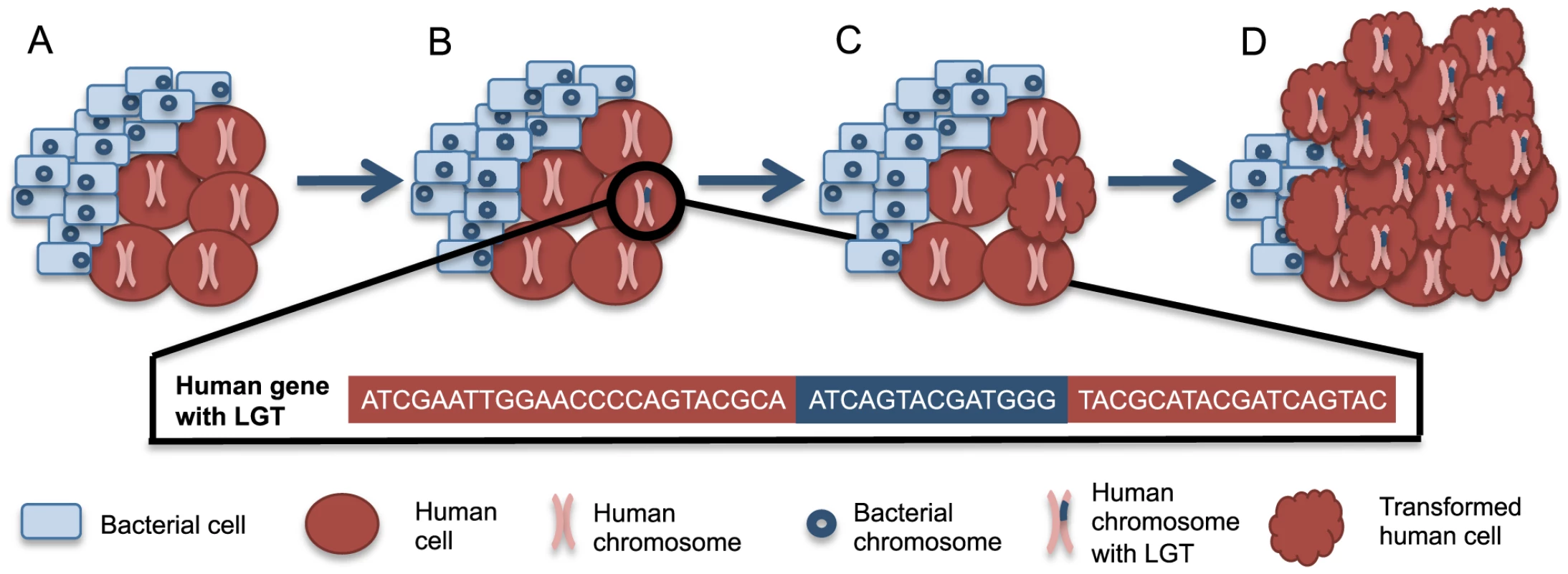
(A) Normally, human cells (red circles) can coexist with a population of microbial cells (blue rectangles), which are not drawn to scale. (B) Occasionally, bacteria-human LGT may occur from one bacterial cell to one human cell as depicted by the human chromosome with bacterial DNA within it (pink chromosome with blue insertion). The bacterial sequence (blue) inserted into the human sequence (red) is illustrated below. (C) The cell with the LGT can undergo transformation to a cancerous phenotype, represented by the scalloped red cell. Such transformation may be related to the integrations or may instead be related to some other alteration in the cell. (D) The now cancerous cell clonally expands and forms a tumor where the majority of cancer cells share the original LGT. The availability of large cancer genome datasets like The Cancer Genome Atlas (TCGA) facilitates testing this hypothesis across a wide variety of cancer types. Evidence has recently been presented supporting bacterial LGTs of Acinetobacter-like DNA in acute myeloid leukemia samples and of Pseudomonas-like DNA in stomach adenocarcinoma samples [82]. There was a higher frequency of LGT in the tumor samples when compared to the available normal samples [82]. The integrations found in stomach adenocarcinoma samples were in known oncogenes and tumor suppressor genes, while the integrations in acute myeloid leukemia samples were in the mitochondrial genome or numts [82]. It was not possible with this analysis to determine if bacterial integrations contributed to carcinogenesis or occurred as passenger mutations during cancer progression. For example, cancer cells may become more permissive to mutations during carcinogenesis and thus receptive to LGT. Regardless, the clonal expansion of tumor cells containing LGTs, as depicted in Figure 1, likely facilitated this discovery. Integrations were found only from specific members of the microbiome, suggesting that LGT is limited to a subset of bacteria [82]. In these cases of mutagenesis involving explicit carcinogenic bacteria, the transfers and any resulting etiology could be prevented through the development of vaccines. Hopefully this new evidence supporting the existence of bacteria-human LGT will prompt more investigation of this topic.
Hypothesis 2: Bacterial Integration into Somatic Cells Could Yield a Protein or Epitope
With all frequency-based interactions, even the most unlikely event can still occur, albeit very infrequently. Integration of bacterial DNA is more likely to disrupt gene function than to result in expression of transferred genes in the new host. However, following integration of bacterial DNA there is the potential for a bacterial gene to become transcribed and a protein or peptide to be synthesized. Expression of some bacterial genes, like the vitamin K biosynthetic gene, could be beneficial in intestinal cells. In contrast, the synthesis of a protein or peptide with a particular bacterial pathogenicity epitope could elicit an adverse immune reaction to a human cell. Such a reaction could in turn lead to an immune reaction to human epitopes. In combination with the well-established mechanism of imperfect removal of autoreactive lymphocytes [83], this could lead to an autoimmune response and possibly an autoimmune disease. The autoimmune response may persist for a lifetime since it is based on the human epitopes. But, the DNA integration that began the chain reaction may not persist or be detected since the human cell expressing the bacterial epitope may be destroyed in the initial immune reaction. It is important to consider that this is only a hypothesis and no data has been presented to demonstrate that this occurs. While this hypothesis may be more unlikely than the first, it is an idea that should be considered.
Conclusions
Extensive LGT has been detected between bacteria and animals, particularly between endosymbionts and their hosts. Recent LGT may be associated specifically with endosymbionts that colonize germ cells and the germ stem cell of their respective hosts. The extensive LGT observed between Wolbachia endosymbionts and their invertebrate hosts suggests that LGT involving bacteria and animals may occur more frequently than was thought a decade ago. While vertebrates have an immune system and segregated gametes that may prevent transfers like those seen in invertebrates, transfers to the vertebrate somatic genome have not been appreciated and warrant further examination. Bacterial DNA integration may be a mutagen associated with noninherited genetic diseases, like cancer, as described in a recent paper demonstrating LGT from Acinetobacter spp. in leukemia samples and from Pseudomonas spp. in stomach cancer samples.
Zdroje
1. GelvinSB (2003) Agrobacterium-mediated plant transformation: the biology behind the “gene-jockeying” tool. Microbiol Mol Biol Rev 67 : 16–37.
2. GelvinSB (2000) Agrobacterium and plant genes involved in T-DNA transfer and integration. Annu Rev Plant Physiol Plant Mol Biol 51 : 223–256.
3. TzfiraT, RheeY, ChenMH, KunikT, CitovskyV (2000) Nucleic acid transport in plant-microbe interactions: the molecules that walk through the walls. Annu Rev Microbiol 54 : 187–219.
4. LanderES, LintonLM, BirrenB, NusbaumC, ZodyMC, et al. (2001) Initial sequencing and analysis of the human genome. Nature 409 : 860–921.
5. SalzbergSL, WhiteO, PetersonJ, EisenJA (2001) Microbial genes in the human genome: lateral transfer or gene loss? Science 292 : 1903–1906.
6. SkaarEP, SeifertHS (2002) The misidentification of bacterial genes as human cDNAs: was the human D-1 tumor antigen gene acquired from bacteria? Genomics 79 : 625–627.
7. PleasanceED, CheethamRK, StephensPJ, McBrideDJ, HumphraySJ, et al. (2010) A comprehensive catalogue of somatic mutations from a human cancer genome. Nature 463 : 191–196.
8. LeeE, IskowR, YangL, GokcumenO, HaseleyP, et al. (2012) Landscape of somatic retrotransposition in human cancers. Science 337 : 967–971.
9. BassAJ, LawrenceMS, BraceLE, RamosAH, DrierY, et al. (2011) Genomic sequencing of colorectal adenocarcinomas identifies a recurrent VTI1A-TCF7L2 fusion. Nat Genet 43 : 964–968.
10. SungWK, ZhengH, LiS, ChenR, LiuX, et al. (2012) Genome-wide survey of recurrent HBV integration in hepatocellular carcinoma. Nat Genet 44 : 765–769.
11. Dunning HotoppJC (2011) Horizontal gene transfer between bacteria and animals. Trends Genet 27 : 157–163.
12. Dunning Hotopp JC (2013) Lateral gene transfer in multicellular organisms. In: Gophna U, editor. Lateral gene transfer in evolution. Springer Science. pp. 161–179.
13. GrahamLA, LougheedSC, EwartKV, DaviesPL (2008) Lateral transfer of a lectin-like antifreeze protein gene in fishes. PLoS ONE 3: e2616 doi:10.1371/journal.pone.0002616
14. MoranNA, JarvikT (2010) Lateral transfer of genes from fungi underlies carotenoid production in aphids. Science 328 : 624–627.
15. AltincicekB, KovacsJL, GerardoNM (2012) Horizontally transferred fungal carotenoid genes in the two-spotted spider mite Tetranychus urticae. Biol Lett 8 : 253–257.
16. HusnikF, NikohN, KogaR, RossL, DuncanRP, et al. (2013) Horizontal gene transfer from diverse bacteria to an insect genome enables a tripartite nested mealybug symbiosis. Cell 153 : 1567–1578.
17. AcunaR, PadillaBE, Florez-RamosCP, RubioJD, HerreraJC, et al. (2012) Adaptive horizontal transfer of a bacterial gene to an invasive insect pest of coffee. Proc Natl Acad Sci U S A 109 : 4197–4202.
18. DanchinEGJ, RossoaM-N, VieiraaP, Almeida-EngleraJd, CoutinhobPM, et al. (2010) Multiple lateral gene transfers and duplications have promoted plant parasitism ability in nematodes. Proc Natl Acad Sci U S A 107 : 17651–17656.
19. MayerWE, SchusterLN, BartelmesG, DieterichC, SommerRJ (2011) Horizontal gene transfer of microbial cellulases into nematode genomes is associated with functional assimilation and gene turnover. BMC Evol Biol 11 : 13.
20. DieterichC, CliftonSW, SchusterLN, ChinwallaA, DelehauntyK, et al. (2008) The Pristionchus pacificus genome provides a unique perspective on nematode lifestyle and parasitism. Nat Genet 40 : 1193–1198.
21. StouthamerR, BreeuwerJA, HurstGD (1999) Wolbachia pipientis: microbial manipulator of arthropod reproduction. Annu Rev Microbiol 53 : 71–102.
22. WerrenJH (1997) Biology of Wolbachia. Annu Rev Entomol 42 : 587–609.
23. FastEM, ToomeyME, PanaramK, DesjardinsD, KolaczykED, et al. (2011) Wolbachia enhance Drosophila stem cell proliferation and target the germline stem cell niche. Science 334 : 990–992.
24. KondoN, NikohN, IjichiN, ShimadaM, FukatsuT (2002) Genome fragment of Wolbachia endosymbiont transferred to X chromosome of host insect. Proc Natl Acad Sci U S A 99 : 14280–14285.
25. NikohN, TanakaK, ShibataF, KondoN, HizumeM, et al. (2008) Wolbachia genome integrated in an insect chromosome: evolution and fate of laterally transferred endosymbiont genes. Genome Res 18 : 272–280.
26. AikawaT, AnbutsuH, NikohN, KikuchiT, ShibataF, et al. (2009) Longicorn beetle that vectors pinewood nematode carries many Wolbachia genes on an autosome. Proc Biol Sci 276 : 3791–3798.
27. Dunning HotoppJC, ClarkME, OliveiraDC, FosterJM, FischerP, et al. (2007) Widespread lateral gene transfer from intracellular bacteria to multicellular eukaryotes. Science 317 : 1753–1756.
28. WerrenJH, RichardsS, DesjardinsCA, NiehuisO, GadauJ, et al. (2010) Functional and evolutionary insights from the genomes of three parasitoid Nasonia species. Science 327 : 343–348.
29. DoudoumisV, AlamU, AksoyE, Abd-AllaA, TsiamisG, et al. (2012) Tsetse-Wolbachia symbiosis: comes of age and has great potential for pest and disease control. J Invertebr Pathol 112 Suppl 1: S94–S103.
30. FennK, ConlonC, JonesM, QuailMA, HolroydNE, et al. (2006) Phylogenetic relationships of the Wolbachia of nematodes and arthropods. PLoS Pathog 2: e94 doi:10.1371/journal.ppat.0020094
31. McNultySN, FosterJM, MitrevaM, Dunning HotoppJC, MartinJ, et al. (2010) Endosymbiont DNA in endobacteria-free filarial nematodes indicates ancient horizontal genetic transfer. PLoS ONE 5: e11029 doi:10.1371/journal.pone.0011029
32. BeikoRG, HarlowTJ, RaganMA (2005) Highways of gene sharing in prokaryotes. Proc Natl Acad Sci U S A 102 : 14332–14337.
33. NikohN, McCutcheonJP, KudoT, MiyagishimaS-y, MoranNA, et al. (2010) Bacterial genes in the aphid genome: absence of functional gene transfer from Buchnera to its host. PLoS Genet 6: e1000827 doi:10.1371/journal.pgen.1000827
34. NikohN, NakabachiA (2009) Aphids acquired symbiotic genes via lateral gene transfer. BMC Biol 7 : 12.
35. LuckeyTD (1970) Introduction to the ecology of the intestinal flora. Am J Clin Nutr 23 : 1430–1432.
36. MogensenTH (2009) Pathogen recognition and inflammatory signaling in innate immune defenses. Clin Microbiol Rev 22 : 240–273.
37. HackerG, RedeckeV, HackerH (2002) Activation of the immune system by bacterial CpG-DNA. Immunology 105 : 245–251.
38. SanderLE, DavisMJ, BoekschotenMV, AmsenD, DascherCC, et al. (2011) Detection of prokaryotic mRNA signifies microbial viability and promotes immunity. Nature 474 : 385–389.
39. WeinerAM (1980) An abundant cytoplasmic 7S RNA is complementary to the dominant interspersed middle repetitive DNA sequence family in the human genome. Cell 22 : 209–218.
40. KapitonovVV, JurkaJ (2003) A novel class of SINE elements derived from 5S rRNA. Mol Biol Evol 20 : 694–702.
41. Roy-EngelAM (2012) LINEs, SINEs and other retroelements: do birds of a feather flock together? Front Biosci 17 : 1345–1361.
42. HancksDC, KazazianHHJr (2012) Active human retrotransposons: variation and disease. Curr Opin Genet Dev 22 : 191–203.
43. MillsRE, BennettEA, IskowRC, DevineSE (2007) Which transposable elements are active in the human genome? Trends Genet 23 : 183–191.
44. KonkelMK, BatzerMA (2010) A mobile threat to genome stability: the impact of non-LTR retrotransposons upon the human genome. Semin Cancer Biol 20 : 211–221.
45. MikiY, NishishoI, HoriiA, MiyoshiY, UtsunomiyaJ, et al. (1992) Disruption of the APC gene by a retrotransposal insertion of L1 sequence in a colon cancer. Cancer Res 52 : 643–645.
46. SolyomS, EwingAD, RahrmannEP, DoucetT, NelsonHH, et al. (2012) Extensive somatic L1 retrotransposition in colorectal tumors. Genome Res 22 : 2328–2338.
47. IskowRC, McCabeMT, MillsRE, ToreneS, PittardWS, et al. (2010) Natural mutagenesis of human genomes by endogenous retrotransposons. Cell 141 : 1253–1261.
48. de MartelC, FerlayJ, FranceschiS, VignatJ, BrayF, et al. (2012) Global burden of cancers attributable to infections in 2008: a review and synthetic analysis. Lancet Oncol 13 : 607–615.
49. SchiffmanM, CastlePE, JeronimoJ, RodriguezAC, WacholderS (2007) Human papillomavirus and cervical cancer. Lancet 370 : 890–907.
50. FaridiR, ZahraA, KhanK, IdreesM (2011) Oncogenic potential of Human Papillomavirus (HPV) and its relation with cervical cancer. Virol J 8 : 269.
51. CordenSA, Sant-CassiaLJ, EastonAJ, MorrisAG (1999) The integration of HPV-18 DNA in cervical carcinoma. Mol Pathol 52 : 275–282.
52. MelsheimerP, VinokurovaS, WentzensenN, BastertG, von Knebel DoeberitzM (2004) DNA aneuploidy and integration of human papillomavirus type 16 E6/E7 oncogenes in intraepithelial neoplasia and invasive squamous cell carcinoma of the cervix uteri. Clin Cancer Res 10 : 3059–3063.
53. Van TineBA, KappesJC, BanerjeeNS, KnopsJ, LaiL, et al. (2004) Clonal selection for transcriptionally active viral oncogenes during progression to cancer. J Virol 78 : 11172–11186.
54. BeasleyRP, HwangLY, LinCC, ChienCS (1981) Hepatocellular carcinoma and hepatitis B virus. A prospective study of 22,707 men in Taiwan. Lancet 2 : 1129–1133.
55. MoorePS, ChangY (2010) Why do viruses cause cancer? Highlights of the first century of human tumour virology. Nat Rev Cancer 10 : 878–889.
56. MurakamiY, SaigoK, TakashimaH, MinamiM, OkanoueT, et al. (2005) Large scaled analysis of hepatitis B virus (HBV) DNA integration in HBV related hepatocellular carcinomas. Gut 54 : 1162–1168.
57. Paterlini-BrechotP, SaigoK, MurakamiY, ChamiM, GozuacikD, et al. (2003) Hepatitis B virus-related insertional mutagenesis occurs frequently in human liver cancers and recurrently targets human telomerase gene. Oncogene 22 : 3911–3916.
58. GozuacikD, MurakamiY, SaigoK, ChamiM, MugnierC, et al. (2001) Identification of human cancer-related genes by naturally occurring Hepatitis B Virus DNA tagging. Oncogene 20 : 6233–6240.
59. SaigoK, YoshidaK, IkedaR, SakamotoY, MurakamiY, et al. (2008) Integration of hepatitis B virus DNA into the myeloid/lymphoid or mixed-lineage leukemia (MLL4) gene and rearrangements of MLL4 in human hepatocellular carcinoma. Hum Mutat 29 : 703–708.
60. LopezJV, YuhkiN, MasudaR, ModiW, O'BrienSJ (1994) Numt, a recent transfer and tandem amplification of mitochondrial DNA to the nuclear genome of the domestic cat. J Mol Evol 39 : 174–190.
61. Hazkani-CovoE, ZellerRM, MartinW (2010) Molecular poltergeists: mitochondrial DNA copies (numts) in sequenced nuclear genomes. PLoS Genet 6: e1000834 doi:10.1371/journal.pgen.1000834
62. Hazkani-CovoE, CovoS (2008) Numt-mediated double-strand break repair mitigates deletions during primate genome evolution. PLoS Genet 4: e1000237 doi:10.1371/journal.pgen.1000237
63. Willett-BrozickJE, SavulSA, RicheyLE, BaysalBE (2001) Germ line insertion of mtDNA at the breakpoint junction of a reciprocal constitutional translocation. Hum Genet 109 : 216–223.
64. ShayJW, BabaT, ZhanQM, KamimuraN, CuthbertJA (1991) HeLaTG cells have mitochondrial DNA inserted into the c-myc oncogene. Oncogene 6 : 1869–1874.
65. BorensztajnK, ChafaO, Alhenc-GelasM, SalhaS, ReghisA, et al. (2002) Characterization of two novel splice site mutations in human factor VII gene causing severe plasma factor VII deficiency and bleeding diathesis. Br J Haematol 117 : 168–171.
66. TurnerC, KilloranC, ThomasNS, RosenbergM, ChuzhanovaNA, et al. (2003) Human genetic disease caused by de novo mitochondrial-nuclear DNA transfer. Hum Genet 112 : 303–309.
67. GoldinE, StahlS, CooneyAM, KaneskiCR, GuptaS, et al. (2004) Transfer of a mitochondrial DNA fragment to MCOLN1 causes an inherited case of mucolipidosis IV. Hum Mutat 24 : 460–465.
68. AhmedZM, SmithTN, RiazuddinS, MakishimaT, GhoshM, et al. (2002) Nonsyndromic recessive deafness DFNB18 and Usher syndrome type IC are allelic mutations of USHIC. Hum Genet 110 : 527–531.
69. ChenJM, ChuzhanovaN, StensonPD, FerecC, CooperDN (2005) Meta-analysis of gross insertions causing human genetic disease: novel mutational mechanisms and the role of replication slippage. Hum Mutat 25 : 207–221.
70. SchonEA, DiMauroS, HiranoM (2012) Human mitochondrial DNA: roles of inherited and somatic mutations. Nat Rev Genet 13 : 878–890.
71. CaroP, GomezJ, ArduiniA, Gonzalez-SanchezM, Gonzalez-GarciaM, et al. (2010) Mitochondrial DNA sequences are present inside nuclear DNA in rat tissues and increase with age. Mitochondrion 10 : 479–486.
72. WangD, LloydAH, TimmisJN (2012) Nuclear genome diversity in somatic cells is accelerated by environmental stress. Plant Signal Behav 7 : 595–597.
73. KustersJG, van VlietAH, KuipersEJ (2006) Pathogenesis of Helicobacter pylori infection. Clin Microbiol Rev 19 : 449–490.
74. MaddocksODK, ShortAJ, DonnenbergMS, BaderS, HarrisonDJ (2009) Attaching and effacing Escherichia coli downregulate DNA mismatch repair protein in vitro and are associated with colorectal adenocarcinomas in humans. PLoS ONE 4: e5517 doi:10.1371/journal.pone.0005517
75. FriedB, ReddyA, MayerD (2011) Helminths in human carcinogenesis. Cancer Lett 305 : 239–249.
76. SearsCL, IslamS, SahaA, ArjumandM, AlamNH, et al. (2008) Association of enterotoxigenic Bacteroides fragilis infection with inflammatory diarrhea. Clin Infect Dis 47 : 797–803.
77. KosticAD, OjesinaAI, PedamalluCS, JungJ, VerhaakRG, et al. (2011) PathSeq: software to identify or discover microbes by deep sequencing of human tissue. Nat Biotechnol 29 : 393–396.
78. ArumugamM, RaesJ, PelletierE, Le PaslierD, YamadaT, et al. (2011) Enterotypes of the human gut microbiome. Nature 473 : 174–180.
79. JeonS, Allen-HoffmannBL, LambertPF (1995) Integration of human papillomavirus type 16 into the human genome correlates with a selective growth advantage of cells. J Virol 69 : 2989–2997.
80. SchwabeRF, WangTC (2012) Cancer. Bacteria deliver a genotoxic hit. Science 338 : 52–53.
81. ChangAH, ParsonnetJ (2010) Role of bacteria in oncogenesis. Clin Microbiol Rev 23 : 837–857.
82. RileyDR, SieberKB, RobinsonKM, WhiteJR, GanesanA, et al. (2013) Bacteria-human somatic cell lateral gene transfer is enriched in cancer samples. PLoS Comput Biol 9: e1003107 doi:10.1371/journal.pcbi.1003107
83. SalinasGF, BrazaF, BrouardS, TakPP, BaetenD (2013) The role of B lymphocytes in the progression from autoimmunity to autoimmune disease. Clin Immunol 146 : 34–45.
Štítky
Genetika Reprodukční medicína
Článek Defending Sperm FunctionČlánek How to Choose the Right MateČlánek Conserved Translatome Remodeling in Nematode Species Executing a Shared Developmental TransitionČlánek Genome-Wide and Cell-Specific Epigenetic Analysis Challenges the Role of Polycomb in SpermatogenesisČlánek The Integrator Complex Subunit 6 (Ints6) Confines the Dorsal Organizer in Vertebrate EmbryogenesisČlánek Multiple bHLH Proteins form Heterodimers to Mediate CRY2-Dependent Regulation of Flowering-Time inČlánek Playing the Field: Sox10 Recruits Different Partners to Drive Central and Peripheral MyelinationČlánek A Minimal Nitrogen Fixation Gene Cluster from sp. WLY78 Enables Expression of Active Nitrogenase inČlánek Evolutionary Tuning of Protein Expression Levels of a Positively Autoregulated Two-Component System
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2013 Číslo 10
-
Všechny články tohoto čísla
- Defending Sperm Function
- How to Choose the Right Mate
- A Mutation in the Gene in Labrador Retrievers with Hereditary Nasal Parakeratosis (HNPK) Provides Insights into the Epigenetics of Keratinocyte Differentiation
- Conserved Translatome Remodeling in Nematode Species Executing a Shared Developmental Transition
- A Novel Actin mRNA Splice Variant Regulates ACTG1 Expression
- Tracking Proliferative History in Lymphocyte Development with Cre-Mediated Sister Chromatid Recombination
- Correlated Occurrence and Bypass of Frame-Shifting Insertion-Deletions (InDels) to Give Functional Proteins
- Chimeric Protein Complexes in Hybrid Species Generate Novel Phenotypes
- Loss of miR-10a Activates and Collaborates with Activated Wnt Signaling in Inducing Intestinal Neoplasia in Female Mice
- Both Rare and Copy Number Variants Are Prevalent in Agenesis of the Corpus Callosum but Not in Cerebellar Hypoplasia or Polymicrogyria
- Reverse PCA, a Systematic Approach for Identifying Genes Important for the Physical Interaction between Protein Pairs
- Partial Deletion of Chromosome 8 β-defensin Cluster Confers Sperm Dysfunction and Infertility in Male Mice
- Genome-Wide and Cell-Specific Epigenetic Analysis Challenges the Role of Polycomb in Spermatogenesis
- Coordinate Regulation of Mature Dopaminergic Axon Morphology by Macroautophagy and the PTEN Signaling Pathway
- Cooperation between RUNX1-ETO9a and Novel Transcriptional Partner KLF6 in Upregulation of in Acute Myeloid Leukemia
- Mobility of the Native Conjugative Plasmid pLS20 Is Regulated by Intercellular Signaling
- FliZ Is a Global Regulatory Protein Affecting the Expression of Flagellar and Virulence Genes in Individual Bacterial Cells
- Specific Tandem Repeats Are Sufficient for Paramutation-Induced Trans-Generational Silencing
- Condensin II Subunit dCAP-D3 Restricts Retrotransposon Mobilization in Somatic Cells
- Dominant Mutations in Identify the Mlh1-Pms1 Endonuclease Active Site and an Exonuclease 1-Independent Mismatch Repair Pathway
- The Insulator Homie Promotes Expression and Protects the Adjacent Gene from Repression by Polycomb Spreading
- Human Intellectual Disability Genes Form Conserved Functional Modules in
- Coordination of Cell Proliferation and Cell Fate Determination by CES-1 Snail
- ORFs in Drosophila Are Important to Organismal Fitness and Evolved Rapidly from Previously Non-coding Sequences
- Different Roles of Eukaryotic MutS and MutL Complexes in Repair of Small Insertion and Deletion Loops in Yeast
- The Spore Differentiation Pathway in the Enteric Pathogen
- Acceleration of the Glycolytic Flux by Steroid Receptor Coactivator-2 Is Essential for Endometrial Decidualization
- The Human Nuclear Poly(A)-Binding Protein Promotes RNA Hyperadenylation and Decay
- Genome Wide Analysis Reveals Zic3 Interaction with Distal Regulatory Elements of Stage Specific Developmental Genes in Zebrafish
- Xbp1 Directs Global Repression of Budding Yeast Transcription during the Transition to Quiescence and Is Important for the Longevity and Reversibility of the Quiescent State
- The Integrator Complex Subunit 6 (Ints6) Confines the Dorsal Organizer in Vertebrate Embryogenesis
- Incorporating Motif Analysis into Gene Co-expression Networks Reveals Novel Modular Expression Pattern and New Signaling Pathways
- The Bacterial Response Regulator ArcA Uses a Diverse Binding Site Architecture to Regulate Carbon Oxidation Globally
- Direct Monitoring of the Strand Passage Reaction of DNA Topoisomerase II Triggers Checkpoint Activation
- Multiple bHLH Proteins form Heterodimers to Mediate CRY2-Dependent Regulation of Flowering-Time in
- A Reversible Histone H3 Acetylation Cooperates with Mismatch Repair and Replicative Polymerases in Maintaining Genome Stability
- ALS-Associated Mutations Result in Compromised Alternative Splicing and Autoregulation
- Robust Demographic Inference from Genomic and SNP Data
- Preferential Binding to Elk-1 by SLE-Associated Risk Allele Upregulates Expression
- Rad52 Sumoylation Prevents the Toxicity of Unproductive Rad51 Filaments Independently of the Anti-Recombinase Srs2
- The Serum Resistome of a Globally Disseminated Multidrug Resistant Uropathogenic Clone
- Identification of 526 Conserved Metazoan Genetic Innovations Exposes a New Role for Cofactor E-like in Neuronal Microtubule Homeostasis
- SUMO Localizes to the Central Element of Synaptonemal Complex and Is Required for the Full Synapsis of Meiotic Chromosomes in Budding Yeast
- Integrated Enrichment Analysis of Variants and Pathways in Genome-Wide Association Studies Indicates Central Role for IL-2 Signaling Genes in Type 1 Diabetes, and Cytokine Signaling Genes in Crohn's Disease
- Genome-Wide High-Resolution Mapping of UV-Induced Mitotic Recombination Events in
- Genome-Wide Analysis of Cell Type-Specific Gene Transcription during Spore Formation in
- Playing the Field: Sox10 Recruits Different Partners to Drive Central and Peripheral Myelination
- Two Portable Recombination Enhancers Direct Donor Choice in Fission Yeast Heterochromatin
- Mining the Human Phenome Using Allelic Scores That Index Biological Intermediates
- Yeast Tdh3 (Glyceraldehyde 3-Phosphate Dehydrogenase) Is a Sir2-Interacting Factor That Regulates Transcriptional Silencing and rDNA Recombination
- A Minimal Nitrogen Fixation Gene Cluster from sp. WLY78 Enables Expression of Active Nitrogenase in
- A Review of Bacteria-Animal Lateral Gene Transfer May Inform Our Understanding of Diseases like Cancer
- High Throughput Sequencing Reveals Alterations in the Recombination Signatures with Diminishing Spo11 Activity
- Partitioning the Heritability of Tourette Syndrome and Obsessive Compulsive Disorder Reveals Differences in Genetic Architecture
- Eleven Candidate Susceptibility Genes for Common Familial Colorectal Cancer
- A GDF5 Point Mutation Strikes Twice - Causing BDA1 and SYNS2
- Systematic Unraveling of the Unsolved Pathway of Nicotine Degradation in
- Natural Genetic Variation of Integrin Alpha L () Modulates Ischemic Brain Injury in Stroke
- Evolutionary Tuning of Protein Expression Levels of a Positively Autoregulated Two-Component System
- Evolutionary Change within a Bipotential Switch Shaped the Sperm/Oocyte Decision in Hermaphroditic Nematodes
- Limiting of the Innate Immune Response by SF3A-Dependent Control of MyD88 Alternative mRNA Splicing
- Multiple Signaling Pathways Coordinate to Induce a Threshold Response in a Chordate Embryo
- Distinct Regulatory Mechanisms Act to Establish and Maintain Pax3 Expression in the Developing Neural Tube
- Genome Wide Analysis of Narcolepsy in China Implicates Novel Immune Loci and Reveals Changes in Association Prior to Versus After the 2009 H1N1 Influenza Pandemic
- Mismatch Repair Genes and Modify CAG Instability in Huntington's Disease Mice: Genome-Wide and Candidate Approaches
- The Histone H3 K27 Methyltransferase KMT6 Regulates Development and Expression of Secondary Metabolite Gene Clusters
- Hsp70-Hsp40 Chaperone Complex Functions in Controlling Polarized Growth by Repressing Hsf1-Driven Heat Stress-Associated Transcription
- Function and Evolution of DNA Methylation in
- Stimulation of mTORC1 with L-leucine Rescues Defects Associated with Roberts Syndrome
- Transcription Termination and Chimeric RNA Formation Controlled by FPA
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- A GDF5 Point Mutation Strikes Twice - Causing BDA1 and SYNS2
- Dominant Mutations in Identify the Mlh1-Pms1 Endonuclease Active Site and an Exonuclease 1-Independent Mismatch Repair Pathway
- Eleven Candidate Susceptibility Genes for Common Familial Colorectal Cancer
- The Histone H3 K27 Methyltransferase KMT6 Regulates Development and Expression of Secondary Metabolite Gene Clusters
Přihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



