-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaThe Power of Gene-Based Rare Variant Methods to Detect Disease-Associated Variation and Test Hypotheses About Complex Disease
Re-sequencing technologies allow for a more complete interrogation of the role of human variation in complex disease. The inadequate power of single variant methods to assess the role of less common variation has led to the development of numerous statistical methods for testing aggregate groups of variants for association with disease. Such endeavors pose substantial analytical challenges, however, due to the diverse array of genetic hypotheses that need to be considered. In this work, we systematically quantify and compare the performance of a panel of commonly used gene-based association methods under a range of allelic architectures, significance thresholds, locus effect sizes, sample sizes, and filters for neutral variation. We find that MiST, SKAT-O, and KBAC have the highest mean power across simulated datasets. Across all methods, however, the power to detect even loci of relatively large effect is very low at exome-wide significance thresholds for sample sizes comparable with those of ongoing sequencing studies; as such, the absence of signal in studies of a few thousand individuals does not exclude a role for rare variation in complex traits. Finally, we directly compare the results reported by different gene-based methods in order to identify their comparative advantages and disadvantages under distinct locus architectures. Our findings have implications for meaningful interpretation of both positive and negative findings in ongoing and future sequencing studies.
Published in the journal: . PLoS Genet 11(4): e32767. doi:10.1371/journal.pgen.1005165
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1005165Summary
Re-sequencing technologies allow for a more complete interrogation of the role of human variation in complex disease. The inadequate power of single variant methods to assess the role of less common variation has led to the development of numerous statistical methods for testing aggregate groups of variants for association with disease. Such endeavors pose substantial analytical challenges, however, due to the diverse array of genetic hypotheses that need to be considered. In this work, we systematically quantify and compare the performance of a panel of commonly used gene-based association methods under a range of allelic architectures, significance thresholds, locus effect sizes, sample sizes, and filters for neutral variation. We find that MiST, SKAT-O, and KBAC have the highest mean power across simulated datasets. Across all methods, however, the power to detect even loci of relatively large effect is very low at exome-wide significance thresholds for sample sizes comparable with those of ongoing sequencing studies; as such, the absence of signal in studies of a few thousand individuals does not exclude a role for rare variation in complex traits. Finally, we directly compare the results reported by different gene-based methods in order to identify their comparative advantages and disadvantages under distinct locus architectures. Our findings have implications for meaningful interpretation of both positive and negative findings in ongoing and future sequencing studies.
Introduction
To assess whether a single variant at a locus contributes to disease risk, the statistical analysis framework is relatively straightforward: compare the frequencies of alleles or genotypes at the site in relation to phenotype. To assess whether multiple variants in the same gene contribute to disease, a much larger array of potential genetic models must be considered. If the causal alleles are rare (defined here as MAF<1%), then power to detect each variant’s effect individually is limited. For example, power to detect a variant with MAF = 0.5% and relative risk (RR) = 3 in 3K case-control samples (1.5K cases and 1.5K controls) at α = 5×10-8 is ~5% [1]. Variants that are private to individuals, as some deleterious mutations are hypothesized to be, present greater challenges yet. As a result, numerous statistical methods have been developed in recent years to test aggregate groups of rare variants for association to disease [2–4].
Re-sequencing experiments have identified a handful of rare variants which modulate risk for common, complex diseases. Examples include variants in NOD2 for Crohn’s disease (4 variants with MAF 0.1–0.8%, ORs 1.4–4.0, detected by single variant association)[5], PCSK9 for coronary heart disease (2 variants with MAF 0.8 and 1.8%, OR ~0.1, detected by single variant association)[6], LPL for hypertriglyceridemia (154 missense variants with MAF<1%, present in cases, detected using the T1 gene-based association method)[7], and MTNR1B for type 2 diabetes (13 functionally-screened variants with MAF<0.1%, collective OR ~5.5, detected using the KBAC gene-based method)[8]. Each of these disease loci is characterized by different numbers, frequencies, and effect sizes of rare variants, but in each of these examples, the estimated proportion of phenotypic variance explained per locus is ~0.5–1.5%.
As large-scale (e.g. genome-wide or exome-wide) studies are now being conducted in hundreds and thousands of individuals, several questions emerge. If loci similar to LPL or MTNR1B exist undiscovered across the genome, what is the power of different gene-based methods to detect them? What effect sizes are studies of a given sample size well-powered to detect? To what extent does power depend on the underlying architecture of causal allelic variation, and how should researchers navigate through the ensemble of available gene-based tests? To interpret the results of gene-based association methods in sequencing studies, it is critical to quantify the power of each method to detect signals under a range of hypothesized locus architectures.
Although the introduction of each novel gene-based association test has typically been accompanied by evaluation of the test’s performance alongside alternatives, each such analysis has compared different subsets of tests, made different assumptions about locus architecture and study design, and employed different simulation approaches. Comparative studies on the relative power of different methods [9–11], while informative, have used small sample sizes, simulated limited locus architectures (e.g., with fixed numbers of causal variants) that may not be representative of complex diseases, and considered only nominal levels of significance (α>0.01). Thus, further work is required to determine how different gene-based tests perform under different genetic models of complex disease.
In this study, we systematically explore the power of eleven currently available and widely-used gene-based association methods to detect rare variant signals drawn from a range of principled genetic architectures of disease, in sample sizes consistent with those of ongoing re-sequencing studies. We assess the impact of locus architecture, effect size, and functional variant filters on the power of each method at stringent levels of significance. By evaluating all tests together at loci simulated under a range of continuous frequency-effect size distributions, we characterize each method’s success and failure modes, and describe genetic hypotheses for which particular methods may be better powered than others.
Results
We first developed a simulation approach to evaluate the power of each gene-based method. We assumed two key requirements for simulations to be informative: 1) simulated genetic variation must approximate the site frequency spectrum (SFS) and haplotype structure of empirical data, and 2) the distribution of relative risks by frequency class should correspond to hypotheses about the genetic architecture of disease that are compatible with observation.
To achieve these objectives, we employed the program HAPGEN2 [12] to simulate variation across the full SFS in thousands of individuals and build a phased reference panel with more individuals than are publicly available at present for a single ethnic group. We started with phased haplotypes from 379 European individuals (1000G Project Phase 1, release 3) [13]. To expand this reference panel to a larger number of individuals, we applied a staged, iterative approach which preserves linkage disequilibrium structure between relatively common variants while introducing new low-frequency variants upon the original haplotypes to match the empirical SFS observed at exonic regions of 202 genes in a study of >12K individuals of European ancestry [14] (S1 Text and Figs 1B, 1C, 1D and S1 and S2). All simulations were performed on 24 human genes of average coding length on chromosome 10 (Fig 1A and S1 Table). While gene coding length does likely contribute to the power to detect association signals, the selection of genes with average length in this study enabled us to conduct controlled characterization of the effects of locus architecture on power.
Fig. 1. Generation of simulated genotype data at human gene loci in large sample sizes with HAPGEN2. 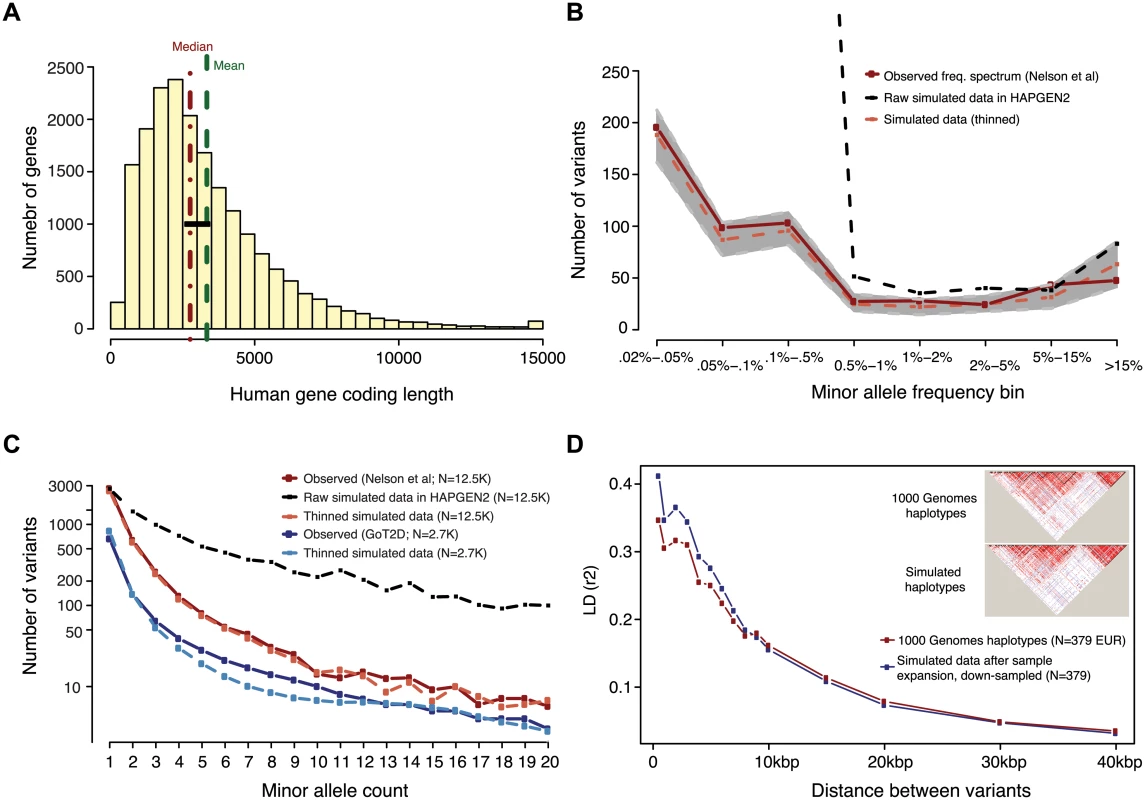
Haplotypes were simulated at ‘average’ human protein-coding genes drawn from the center of the distribution of RefSeq gene total exon length (A). Vertical dotted lines in red and green indicate the median and mean values of exon length, respectively. Black bar represents the 24 genes selected for simulation. (B,C) Site frequency spectrum of simulated data, as compared to observed human data. Data were simulated via staged expansion of 1000 Genomes Project haplotypes using the HAPGEN2 software; the mutation parameter was fit to match the site frequency spectrum of protein-coding variation observed in exome sequencing studies, e.g. as reported Nelson et al 2012. Raw simulated data from HAPGEN2 in large sample sizes produced an excess of rare sites; these were down-sampled to match observed data. The grey area in (B) represents the [5%, 95%] interval across all simulated genes, obtained using bootstrapping. The site frequency spectrum of simulated data in a smaller sample size (N = 2.7K) also matched an independent set of observed exome sequencing data from the GoT2D consortium (C). Haplotype structure, as measured by linkage disequilibrium between variants, was also preserved in the simulated data after sample expansion (D). The inset shows a representative example of simulations at the GATA3 gene locus. We modeled the complex disease type 2 diabetes (T2D, assuming prevalence 8%), and introduced phenotypic effects (relative risk per variant, assuming additive effects) by sampling up to 35 exonic causal variants per locus (variant cap imposed due to software limitations, see Methods) from six different joint distributions of causal variant frequencies and effect sizes (S3 and S4 Figs and Tables 1 and S2). These distributions were obtained from forward simulations of global genetic architecture under different disease models that are consistent with properties of empirical sequence variation and the observed prevalence and heritability of T2D [15]. The three main architectures assume strong (AR1), moderate (AR2), or weak (AR3) purifying selection against causal alleles. Broadly, AR1 results in a sharp inverse correlation between variant frequency and effect size, AR2 produces modest correlation, and AR3 is characterized by rare and common alleles that have similar additive effects on phenotype. AR4 and AR5 are variations of AR1 and AR2, respectively, in which only rare (MAF<1%) variants at a locus contribute to disease. AR6 assumes a frequency-effect size map identical to AR2, but assigns a 50%-50% mixture of risk and protective effects; this represents the hypothesis that some variants in a gene increase disease risk, while other variants in the same gene have protective effects.
Tab. 1. Locus architectures modeled at simulated loci. 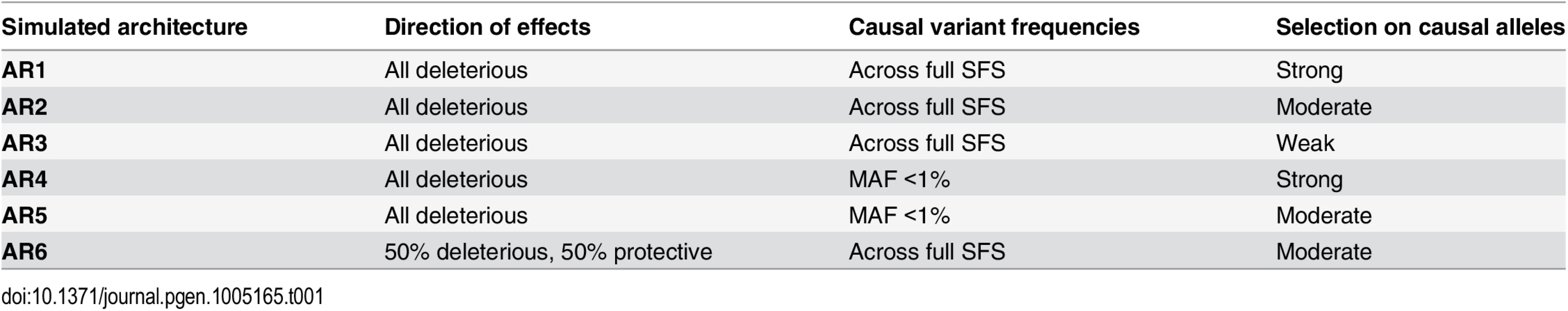
We evaluated a set of eleven gene-based association methods (CMC [16], VT [17], FRQWGT [18], WILCOX-WSS [19], KBAC [20], BURDEN [18], UNIQ [18], C-ALPHA [21], SKAT [22], SKAT-O [23], and MiST [24]; see Table 2) on these simulated datasets. The tests we applied can be broadly categorized as unidirectional ‘burden’ tests, bidirectional variance-component tests (SKAT, C-ALPHA), and linear combinations of these two classes (SKAT-O, MiST). The unidirectional tests can be further sub-divided into collapsing regression methods (CMC), weighted sum methods (FRQWGT, KBAC, WILCOX-WSS, VT), and permutation-based summary count methods (BURDEN, UNIQ). We selected this set of tests because they represent a broad range of analytical approaches, most of which are readily available in the widely-used software packages PLINK/Seq [18] and EPACTS [25]. Before further evaluation, we confirmed that all tests were well-calibrated, at α = 0.05 and 10-4, in (null) datasets where no variants were assigned any causal effects (S5 Fig).
Tab. 2. Published gene-based rare variant association methods evaluated. 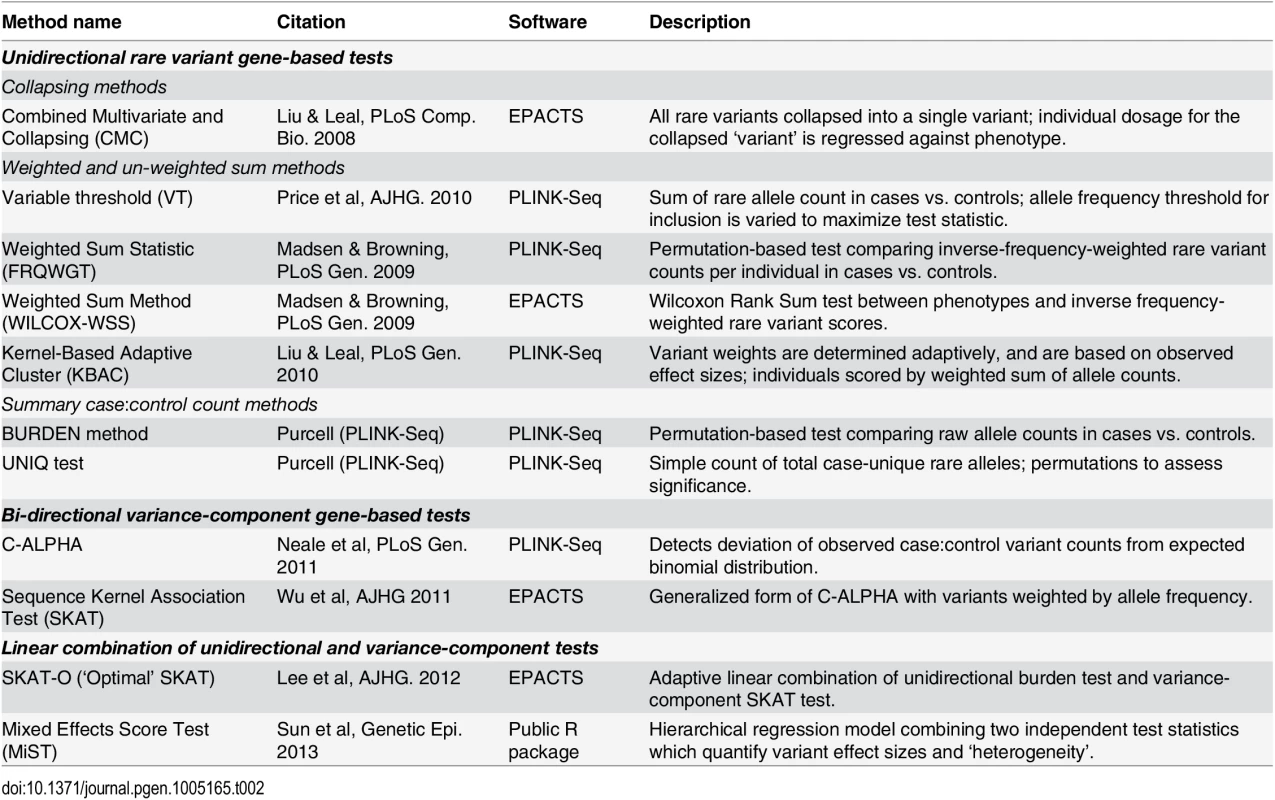
Relative and absolute power of gene-based association methods under different locus architectures
A key question for re-sequencing studies is: what is the power of gene-based association methods to detect causal loci at stringent levels of significance? To address this, we ran each gene-based test at simulated loci explaining 1% of the variance in T2D liability [26, 27] (see Methods) in 1500 cases and 1500 controls (sample size comparable to several recent or ongoing complex trait sequencing studies [28, 29]). Each gene-based test was run on all exonic variants (causal and non-causal) with MAF<1%, unless otherwise stated. The power of each test is shown as a function of significance threshold (α) and architecture in Figs 2, S6, and S7.
Fig. 2. Power of different gene-based rare variant association methods at simulated disease loci. 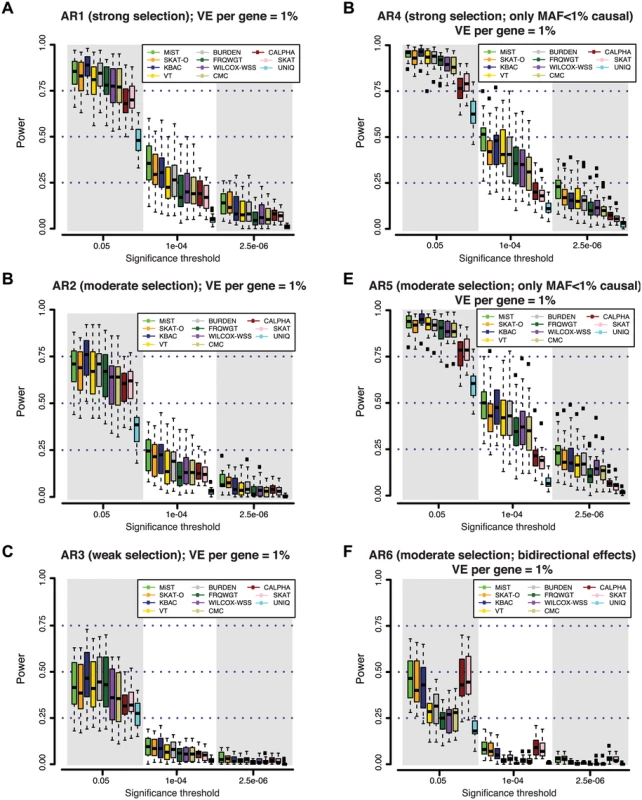
At each gene locus, one hundred independent simulations of phenotypic effects were generated in a sample size of 3K individuals (1.5K cases / 1.5K controls). Variant effects were drawn from varied models of genetic architecture (A-F), hypothesizing different degrees of purifying selection against disease alleles (see Methods). Under models with strong selection, there is a strong inverse correlation between variant frequency and effect size; under weak selection rare variant effects are less skewed. At all loci, genetic variants together contribute 1% of the phenotypic variance underlying a trait with common prevalence (8%; modeled as type 2 diabetes). Power is measured as the fraction out of 100 simulations of each gene in which a gene-based test reported a p-value lower than the significance threshold. In (A-C), causal variants span the full frequency spectrum (including common alleles), and thus rare alleles account for only a fraction of the locus heritability; in (D-E), all causal variants are rare (MAF<1%). In (F), causal variants have bi-directional effects (some increase risk of disease, while others reduce risk). In the context of an exome-wide sequencing study, where an appropriate threshold may be α = 2.5×10-6 (α = 0.05, after Bonferroni correction for ~20K genes), we found that power is very low (<20%) across all architectures and tests considered. At a less stringent threshold of α = 10-4, which might be used to nominate loci for follow-up (under the null, only ~2 such genes would be expected exome-wide), power of the best performing tests across AR1-AR5 remained low (10–50%). This was true irrespective of the allele frequency threshold used for variant inclusion; results for a MAF threshold of 0.5% and 5% are shown in S8 Fig.
We noted that at a nominal level of significance (α = 0.05), many methods had high power (~75%-95%) to detect loci at which deleterious variants (AR1-AR5) explain ~1% of phenotypic variance (Figs 2 and S7). KBAC was consistently the most sensitive method to detect deleterious effects at less stringent levels of significance (up to 95% power at α = 0.05, under AR4). This high sensitivity could be useful in identifying putative signals when only a small number of hypotheses are being tested (e.g. sequencing across only a few targeted loci), or to exclude rare variant models at candidate loci.
Next, we asked whether any of the gene-based methods appear to be uniformly more powerful than others, across the various locus architectures we considered. Under simulated architectures where causal variants all have unidirectional (deleterious) effects (Fig 2A, 2B, 2C, 2D, and 2E), we found that MiST, SKAT-O, and KBAC consistently achieve highest power, while UNIQ is least-powered. However, we did observe differential behavior of these tests depending on the significance threshold: MiST and SKAT-O retained greater power than unidirectional alternatives at stringent thresholds (α<10-5), while at less conservative thresholds (α>10-3), KBAC was more sensitive (Figs 2A, 2B, 2C, 2D, 2E, 2F and S7).
We next sought to understand how power is influenced by locus architecture. Unsurprisingly, we found that power is higher when the majority of the locus’ total phenotypic effect is due to rare variants included in the association test (e.g. those with MAF<1%). This is evidenced by the gain in power under models with a greater contribution of rare variants: the power of MiST, for example, increased from AR3 (10% at α = 10-4 in 3K individuals) to AR2 (23%) to AR1 (36%). Power was higher still under architectures where variants with MAF<1% (i.e. those variants tested) contributed all of the locus’ effect (AR4 and AR5): here, the power of MiST was ~50% at α = 10-4. Power also depends on the direction of causal effects at a locus: under AR6 (where both risk and protective effects are present), the variance-component tests (SKAT and C-ALPHA) and combined tests (MiST and SKAT-O) were least affected (by design) [21–24] and outperformed all the other methods, retaining ~10% power at α = 10-4, while that of unidirectional tests was reduced to <5% (Figs 2F and S7). Finally, we find that power is inversely related to the degree of linkage disequilibrium between causal variants at a locus (S9 Fig).
We next queried the overlap between signals detected by gene-based methods versus those detected by single variant association. In direct contrast to gene-based methods, the power of single variant association decreased as the contribution of rare variants increased: power at a genome-wide threshold of α = 5×10-8 for single variants was ~20%, ~10%, and ~7% under AR3, AR2, and AR1, respectively (blue bars in Fig 3A, 3C, and 3E). However, in all cases, the combined application of gene-based and single variant methods yielded greater sensitivity than single variant association alone (yellow bars in Figs 3A, 3C, 3E, and S10). This occurred because the association tests detect distinct subsets of loci: gene-based methods uniquely identified loci where the signal was driven by groups of rare variants for which single variant association test statistics were not individually significant (pink loci in Fig 3B, 3D, and 3F).
Fig. 3. Power of best-performing gene-based rare variant method as compared to single variant association. 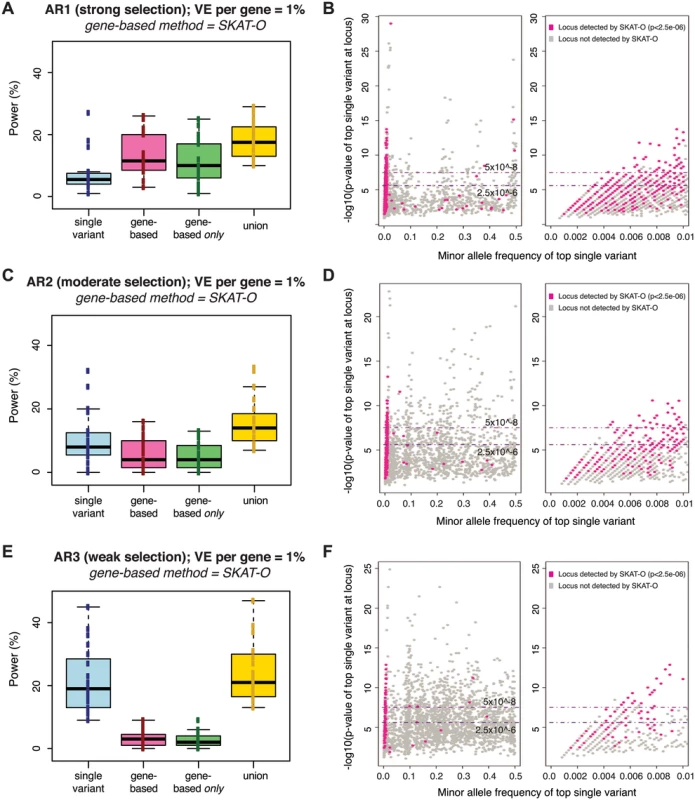
Power is measured across one hundred simulations of phenotypic effects at each of 24 human gene loci in N = 3K samples (as in Fig 2). Under each architecture (AR1, AR2, AR3), the power of the best-performing gene-based test at alpha = 2.5e-06 (SKAT-O) is compared to single variant association (Fisher’s exact) at alpha = 5e-08 (panels A, C, E). No MAF threshold was applied to the single variant association tests; gene-based tests included only variants with MAF<1%. Blue boxplot shows range of power for single variant association across genes simulated; pink shows power of the gene-based test alone; green shows the fraction of loci detected only by gene-based test (and not single variant association); yellow shows the combined power of both gene-based and single variant association. Next to each boxplot (panels B, D, F) are scatterplots on which each simulated locus (under AR1, AR2, and AR3, respectively) is represented as a point based on the minor allele frequency (x-axis) and association p-value (y-axis) of the single most-associated variant (the top individual signal) across the locus. Single variant association detects loci plotted above the upper dotted line (at 5e-08), while gene-based association identifies a distinct subset of loci (those highlighted in pink, where the SKAT-O p-value is <2.5e-06). This latter group of loci are those where the top single variant is preferentially rare (and no common variant association signal exists); right-most scatterplots zoom into this portion of the x-axis (MAF<1%). Similar plots for AR4, AR5, and AR6 are shown in S10 Fig. As expected, the comparative advantage of gene-based tests was most evident under architectures where there is strong purifying selection against causal alleles (under AR4, for example, the power of single-variant tests at α = 5×10-8 was <5%, while gene-based tests achieved ~50% power at α = 10-4, and ~20% power even at α = 2.5×10-6; S10A and S10B Fig). Under both AR2 and AR3 (where limited purifying selection made causal alleles more common), the power of single variant association (~20% at α = 5×10-8 under AR3) exceeded that of the best gene-based test (<5% at α = 2.5×10-6 under AR3), though each method detected unique loci. These results confirm that single variant and gene-based association methods should be jointly employed for maximal power across divergent locus architectures.
To characterize the impact of locus effect size on the power of gene-based tests, we simulated loci where the phenotypic variance explained (VE) by genetic variants is 0.5%, 1% (as in Figs 2 and 3), and 2% (all under AR2). At loci where VE = 2%, power increased to nearly 40% (at α = 10-4), as compared to ~23% when VE = 1% (Figs 4A, S11A, and S11B). When VE = 0.5%, power was extremely low (<8% at α = 10-4 in 3K individuals), indicating that exome-wide sequencing studies of this size are substantially under-powered to interrogate genes for weaker effects (S11A Fig).
Fig. 4. Power of gene-based methods as a function of sample size, locus effect size, and neutral variation. 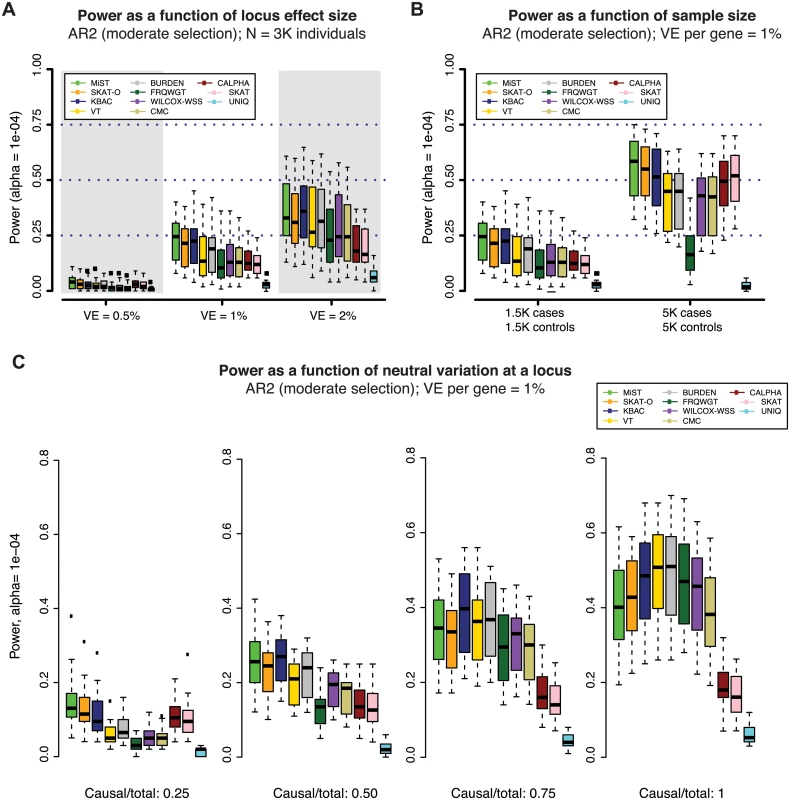
Power was measured across one hundred simulations at each of 24 gene loci (as in Figs 2 and 3). Across all panels above, variant effects were drawn from the architecture model AR2 (assuming moderate selection against causal variants, and thus modest inverse correlation between variant frequency and effect size). In (A), variant effects were sampled at each locus such that the total fraction of phenotypic variance explained by the locus was ~0.5%, 1% (as in Figs 2 and 3) or 2%. In (B), loci were simulated to explain 1% of phenotypic variance in sample sizes of 1.5K cases/1.5K controls (as in Figs 2 and 3) and 5K cases/5K controls. In both (A) and (B), all exonic variants with MAF < 1% were included in the burden test (both causal and non-causal variants, resulting in a fewer than 50% of all tested variants being causal). In (C), non-causal (neutral) variants were selectively removed such that the ratio of causal variants to total variants tested ranged from 0.25 to 1 (only causal variants tested). The gene-based methods each have varied performance under different locus effect sizes, sample sizes, and causal variant filtering scenarios. Impact of sample size and neutral variation on power of gene-based association
The relatively modest power of gene-based tests at stringent levels of significance across the architectures considered here presents challenges to investigators seeking to discover novel disease-associated loci in studies of this size. Thus, we next investigated the extent to which power could be improved by a) increasing sample size, or b) excluding neutral variation at a locus.
We found that gene-based methods exhibit differential gains in power as sample size increases from 3K to 10K individuals (Fig 4B). The median power of MiST, for example, increased from ~23% to ~60% (at α = 10-4, under AR2) in 10K samples and was largely retained (~50%) even at α = 2.5×10-6 (S11C Fig). However, the increase in power was not uniform across methods. This occurred, in part, because (unlike for single variant tests) the relationship between sample size and power is not straightforward for gene-based tests: as sample size increases, causal alleles are observed more times, but the number of (rare) non-causal alleles also grows sharply. Thus, methods that up-weight all rare alleles regardless of their observed effect (e.g., FRQWGT) may benefit least from increases in sample size (S11–S13 Fig).
As the number of observations of rare alleles increases with sample size, the performance of single variant association tests will certainly improve, but our analysis suggests that gene-based tests will still uniquely identify loci at which the aggregate signal is driven by variants too rare to be individually detected. When the top single variant in our simulated datasets had MAF < = 0.4%, the locus was rarely detected by single variant association in a sample of 3K individuals (Fig 3B, 3D, and 3F). Single variant tests would have <80% power to detect an effect at a variant of that frequency (at α = 5×10-8) even in 10K samples, unless the RR of that variant was over 3. Moreover, as sample sizes increase, the threshold required to assess significance for gene-based methods will remain the same (as the number of independent tests performed will not change), while that for single variant association tests will need to become more stringent as more novel variants are discovered. Hence, we expect the joint application of single variant and gene-based methods to remain beneficial even as sample sizes increase.
Our study also confirmed that gene-based tests are highly sensitive to the fraction of neutral variation at a locus (Figs 4C and S13), as has been previously described [10, 11, 23]. We additionally found that unidirectional burden tests exhibit the sharpest increases in power as the fraction of neutral variation decreases. Under AR2 in 3K individuals, KBAC power at α = 10-4 exceeded 50% when only disease-causing variants were included (increasing from ~22% prior to variant filtering). These tests may therefore be most powerful for testing targeted hypotheses at loci where rich functional annotation enables exclusion of a subset of neutral variants. Conversely, variance-component tests (C-ALPHA, SKAT) as well as combined methods (MiST, SKAT-O) are characterized by a relative immunity to neutral variation. This latter group of methods, then, are attractive options for jointly testing large numbers of less strictly filtered variants (e.g. in a pathway-based analysis).
Concordance between the results of different gene-based association methods
We next investigated the degree of overlap between signals detected by each gene-based method. For each pair of association methods, we computed Pearson’s correlation coefficients between their reported p-values on a logarithmic scale (Figs 5A, 5B, and S14). We found that tests with similar design characteristics (e.g., SKAT and C-ALPHA, R2 = 0.99) exhibit very high correlation, as expected (Fig 5C). Some methods were highly correlated, but there was variability in the p-values reported (e.g., MiST and SKAT-O, R2 = 0.92), while others were much less related or even uncorrelated (e.g., SKAT-O and UNIQ, R2 = 0.02). While in this latter case low correlation was driven by the lower mean power of UNIQ relative to SKAT-O, it is worth noting that there did exist a set of true causal loci (where many case-private singletons segregate) at which UNIQ reported p<10-4, but SKAT-O reported p>0.01 (Fig 5C).
Fig. 5. Concordance between results of different gene-based methods. 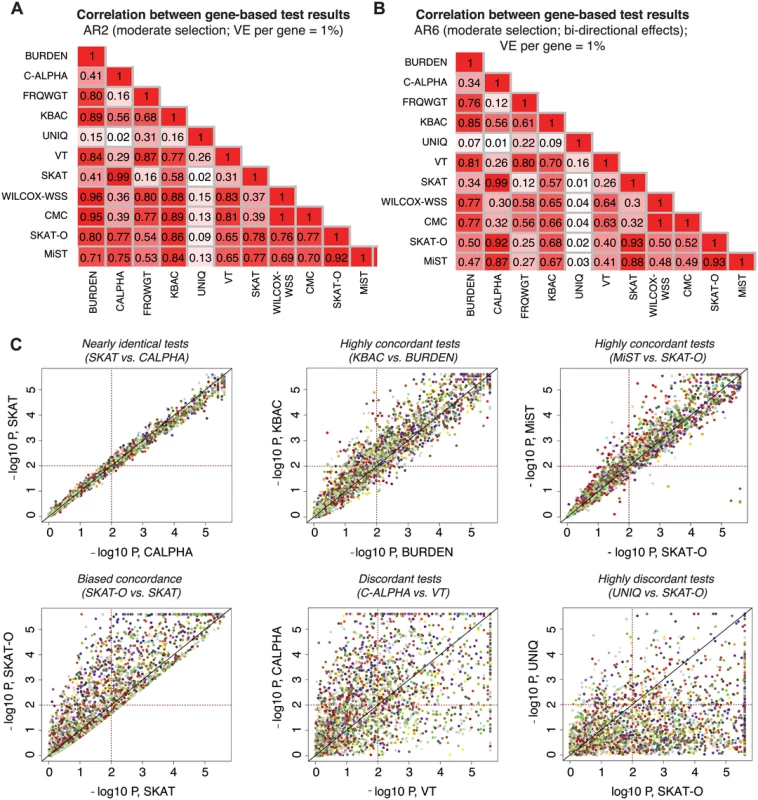
Pairwise correlation coefficients (R2) between the p-values reported by different gene-based association methods under AR2 (moderate selection; shown in (A) and under AR6 (moderate selection and bi-directional phenotypic effects, shown in (B)). P-values above 0.1 are excluded in computation of the correlation. In (C), scatter plots show the results (-log10 of the p-values) reported by a pair of gene-based tests under AR2; p-values below 5e-06 are plotted at 5e-06. Each point represents an individual locus at which both gene-based methods were applied (2400 total points); points of the same color represent different simulations at the same gene loci (e.g. same gene and haplotype structure, but different variant phenotypic effects). Dotted lines mark p = 0.01, such that points above the horizontal line or to the right of the vertical line represent loci at which nominally significant results are reported by the gene-based method. All data above generated in 3K samples (1.5K cases, 1.5K controls). Other methods, such as SKAT and SKAT-O, showed asymmetric concordance (R2 = 0.78): SKAT-O detected a set of causal loci entirely undetected by SKAT, but was more conservative on the whole, reporting p-values up to an order of magnitude higher than those reported by SKAT at the majority of loci tested. These correlations were also architecture-dependent: under AR2 (where there are only deleterious effects), for example, SKAT-O exhibited high concordance with KBAC (R2 = 0.86), while under AR6 (where bidirectional effects are present), SKAT-O was most concordant with C-ALPHA and SKAT (R2 = 0.93). MiST shared this behavior, reflecting the ‘unified’ design of these tests as combinations of a unidirectional burden test and a bidirectional variance-based method [23, 24].
To understand the drivers of such differences and identify scenarios where certain tests may be more powerful than others, we conducted pairwise comparisons between KBAC (one of the highest performing methods at α = 10-4 across AR1-AR5) and the other gene-based methods. We focused here on loci where VE = 1%, simulated under AR2. For each comparison, we characterized the properties of loci at which KBAC (but not the other method) reports p<0.01, and vice-versa. In the comparison between KBAC and C-ALPHA (Fig 6A), we found that loci at which only KBAC detected signal were characterized by a higher aggregate skew in case to control counts (often driven by singletons, which do not contribute to the variance component tests’ dispersion statistic). Loci at which only C-ALPHA detected signal, on the other hand, were characterized by a relatively common single variant of large effect (in the background of many variants with balanced case to control counts).
Fig. 6. Properties of loci at which gene-based methods report discordant results. 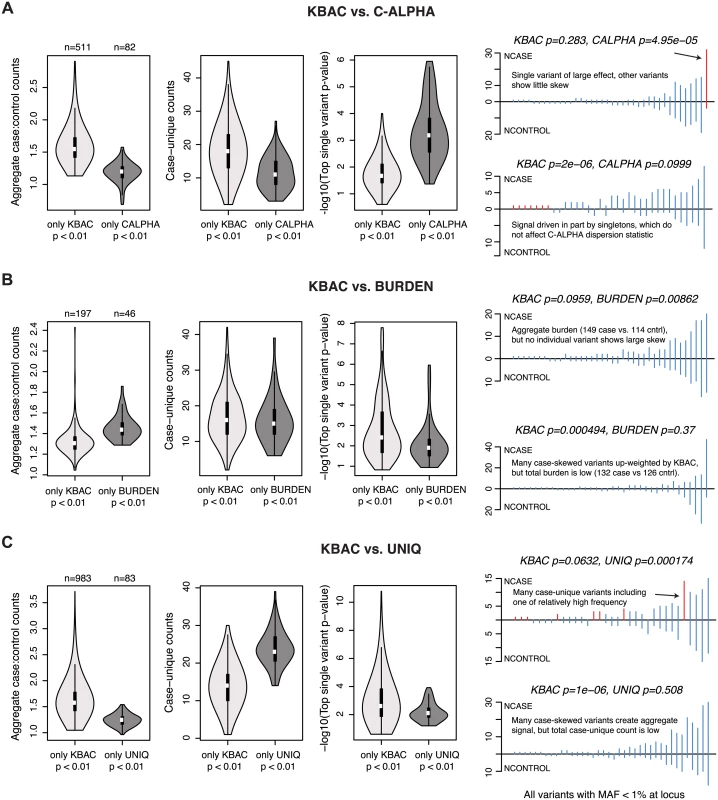
Characteristics of causal loci at which KBAC (the method with highest mean power at nominal levels of significance) produces discordant results as compared to another gene-based method. Results are shown above for the simulated architecture AR2 in 3K samples. KBAC is compared to the (A) C-ALPHA, (B) BURDEN, and (C) UNIQ gene-based methods. In each comparison, loci are identified at which KBAC (but not the other method) reports a p-value < 0.01, or at which the other method (but not KBAC) reports a p-value < 0.01. For each group of loci, leftmost vioplot shows the distribution of aggregate case:control counts (number of minor alleles observed in cases divided by number of minor alleles observed in controls, for variants with MAF<1%). Middle vioplot shows distribution of case-unique counts (number of observations of alleles that are only present in cases and absent from controls). Rightmost vioplot shows distribution of the top single variant p-value observed for an exonic variant at the locus (log10 scale). Line plots at right show the distribution of variants (MAF < 1%) at representative simulated loci where the methods are discordant. Each line represents a variant; height above line measures the variant’s case counts, while height below measures control counts. Red lines highlight variants which drive the difference in test performance. For loci where the ratio of aggregate case to control counts is high, but no individual variants/genotypes show any substantial skew, the BURDEN test may be more powerful than KBAC (Fig 6B). This makes sense: KBAC adaptively weights multi-site genotype counts by their observed case-bias, and if all variants have low weights, the maximum achievable KBAC statistic is low, whereas BURDEN quantifies the significance of the observed signal in aggregate. Finally, UNIQ (unsurprisingly) more readily detected loci at which signal is driven by either many rare variants private to cases, or by a single relatively frequent case-unique (or control-unique) variant (Fig 6C). Taken together, these data indicate that although a given method may exhibit high mean power across divergent architectures, it may not be optimal for testing specific genetic hypotheses.
Given the observation that different methods capture different signals, we wondered whether a strategy in which subsets of methods are collectively applied to a locus might be informative in an exome-wide setting (e.g., to test multiple hypotheses about locus architecture at once). To test this, we employed a stepwise forward selection approach, starting with each of the three best-performing gene-based methods across architectures (MiST, SKAT-O and KBAC) and using the degree of difference (in orders of magnitude) between additional methods’ reported p-values as the inclusion criterion (see Methods, S1 Text).
In 3K individuals, under AR2 (where MiST power is ~23% at α = 10-4), we found that particular combinations of tests (e.g., KBAC+MiST+VT+UNIQ+FRQWGT) could jointly achieve ~31% sensitivity at α = 10-4 (using the single minimum p-value reported across all three tests). However, this gain came at the cost of a higher false positive rate (FPR): after adjusting the p-value significance threshold to correct for the increase in FPR, we found negligible gains in power compared to the application of a single test (S3 Table). Joint application of gene-based tests may still be useful, however, in settings where a higher FPR is tolerable, e.g., to increase sensitivity in a ‘discovery’ exome-wide sequencing scan which precedes large-scale targeted follow-up.
Discussion
Given the wide array of aggregate rare variant association methods now available for application in re-sequencing or genotyping studies of complex traits [30], it is critical to characterize and quantify the statistical power of each method to test heterogeneous genetic hypotheses. In this study, we conducted a comparative analysis of a panel of commonly used gene-based rare variant association tests under a broad range of realistic allelic architectures, significance thresholds, locus effect sizes, sample sizes, and filters for neutral variation.
In sample sizes comparable to those of many contemporary sequencing studies (3K case-control individuals), we find that while gene-based association methods augment the power of single variant tests by preferentially detecting loci at which rare variants drive the causal architecture, their absolute power is low. All gene-based methods evaluated in this study have limited power, even to detect loci explaining as much as 1% of the variance in phenotypic liability underlying a common trait such as type 2 diabetes (mean power across architectures is ~5–20% at α = 2.5×10-6). Even in 10K case-control samples, power remains modest (~60% at α = 2.5×10-6). Based on estimates of variance explained by known rare and common variant signals (the strongest single common variant association for T2D, mapping near TCF7L2, explains ~1% of phenotypic variance), it seems probable that for any given complex trait, at best a handful of loci will have effects on this scale. The full potential of exome sequencing to provide biological insights into disease, then, will depend largely on the detection of loci of smaller aggregate effects, and will require far larger sample sizes than these.
The low mean power to detect disease-associated loci prompted the question of whether some methods are better powered than others to discover novel signals under specific hypothesized locus architectures. We find that at more stringent significance thresholds (α<10-4), MiST and SKAT-O have the highest power across architectures simulated here, especially when rare variants have bidirectional effects on disease. Thus, for investigators looking to discover signals across thousands of loci (e.g., in exome-wide scans), these tests are likely to maximize sensitivity.
Weighted sum methods (and KBAC in particular), on the other hand, are consistently best-powered to detect rare variants of deleterious effect at less stringent levels of significance, and also show the greatest gains in power when neutral variation can be filtered out. These attributes may be useful in various scenarios: to test a small number of biological hypotheses (e.g. at only a few loci, especially if functional annotations are available), to prioritize signals for further follow-up from a discovery scan, or to place bounds (e.g., after an exome-wide sequencing study) on the total number of genes harboring rare variants of a given effect size that are likely to exist.
In addition to MiST, SKAT-O and KBAC, we find that other methods may have individual strengths under particular scenarios (e.g., UNIQ to test whether a gene harbors an excess of highly penetrant rare variants, or BURDEN to detect a collection of variants each of very weak effect); these methods may be optimal for testing such specific genetic hypotheses. Finally, in larger sample sizes (n = 10K case-control individuals), our simulations demonstrate that the increasing number of neutral (non-causal) rare variants may limit gains in the power of some methods (e.g. FRQWGT). Here, MiST is best-powered at stringent significance thresholds. Taken together, these results suggest that the interpretation of novel signal discovery (or the lack thereof) in sequencing studies may vary based on the specific gene-based methods that are used.
This study has a number of limitations. It is based on simulated data (albeit data consistent with available empirical information on genetic variation and disease epidemiology [15]). It does not explore the effects of properties such as demographic history, gene size, mutation rate, haplotype length, or degree of linkage disequilibrium between causal variants on the power of gene-based association methods. Moreover, it does not characterize the performance of these methods at non-coding regions, where causal variant frequencies and effect sizes may be different, and where there is likely a higher proportion of neutral variation. This simulation approach, however, enabled us to undertake a controlled, quantitative characterization of the performance of gene-based association methods under a range of scenarios. Future work should characterize these methods in study populations of different ethnicities, where different site frequency spectra and linkage disequilibrium patterns between causal variants may alter power (S9 Fig). Architectures we simulated assumed a common binary trait; power to detect loci explaining phenotypic variance for less prevalent traits is likely higher, but we did not study this relationship. The tools available on our website (http://mccarthy.well.ox.ac.uk/publications/2014/moutsianas_simulations/) allow the investigation of this question for any complex trait by generating simulated data using a custom, user-specified RR-by-allele frequency heat-map and disease prevalence.
In summary, we find that specific gene-based association methods are best deployed in the setting of particular experimental study designs, and when testing for particular genetic models of disease. Such an approach will likely enable meaningful interpretation of both positive and negative findings in ongoing sequencing studies, and is bound to remain important even as sample sizes increase and new statistical methods for aggregate testing of rare variants are developed.
Methods
Generation of simulated reference panels
Simulated datasets were generated using HAPGEN2 [12]. HAPGEN2 generates case-control data using a haplotype reshuffling approach based on the Li & Stephens model [31]. Under this model, simulated (unobserved) haplotypes are assumed to be an imperfect mosaic of actual (observed) haplotypes and are simulated using a Hidden Markov Model with recombination and mutation rates as parameters. Case and control samples are generated by over-sampling haplotype segments which contain alleles at which phenotypic effects are introduced (based on the relative risks assigned to them). A phased reference panel of haplotypes from 379 European (98 TSI, 89 GBR, 85 CEU, 14 IBS, and 93 FIN) individuals from the 1000 Genomes Project (1000G Project Phase 1, release 3) [13] was augmented to 12,514 individuals by iteratively simulating haplotypes (with no phenotypic effects) and adding them to the original reference panel, in increments of 300 individuals per iteration.
An excess of rare variation was introduced to the data using an empirically selected value of θ = 0.08 for the mutation parameter in HAPGEN2, so as to match the singleton count observed in empirical re-sequencing data in a sample of this size. We used the SFS reported by Nelson et al [14], which was based on sequencing 351kb of coding sequence in 12,514 samples of European descent. The resulting dataset was subsequently thinned using a rejection sampling approach, to match the full site frequency spectrum observed in real data. This two-step approach (matching for singletons, and then thinning the dataset) was necessary to model the excess in rarer variation observed in whole exome sequencing datasets while preserving the LD structure of the reference panel. In order to validate that this approach led to a realistic SFS when sub-sampled to smaller sizes, we compared the SFS observed in the simulated, thinned panel, in subsets of 2,738 individuals, to that of empirical exome-wide sequencing data on the same number of individuals, from the GoT2D project (dark and light blue lines, Fig 1).
Forward simulation of population-scale data to model genetic architecture
Forward population genetic simulations of global complex disease architecture (specifically, for type 2 diabetes, a disease of prevalence 8% and heritability ~45%) were conducted across a range of disease models varying in mutational target size and coupling to purifying selection [15]. By varying only these two parameters, a wide range of continuous joint frequency and effect size distributions were generated; under models with strong coupling to selection, rare variants explain the bulk of heritability and have large effects, while under models with weak coupling to selection, common variants explain the bulk of heritability and rare variants have weaker effects. For the HAPGEN2 simulations conducted here, we sampled variant effect sizes from the distributions observed in the forward simulated datasets at loci explaining ~1% of phenotypic variance underlying T2D (S3 Fig).
Assignment of variant phenotypic effects within HAPGEN2 simulations
Variant effects were selected from the frequency-effect size distributions described above. We simulated these effects at randomly selected exonic variants across each gene. We used variant frequencies measured in the augmented reference panel of 12,514 individuals. In unidirectional architectures, all rare variants were assumed to increase risk of disease (RR>1). In bidirectional architectures, protective effects were sampled in the same way, but the relative risks were inverted. Variant effects were sampled until the cumulative variance explained (VE) on the liability scale by each locus reached the desired threshold (e.g. VE = 0.5%, 1%, or 2%). The following procedure was followed for introducing variation at each locus:
Pick an exonic variant at random
Introduce an effect by sampling from the frequency-RR distribution of the respective architecture
If the cumulative variance explained (on the liability scale, %VE) by variants at the locus is below of the specified threshold, go to step (i) and repeat
If the variance is above the specified threshold, remove one of the introduced effects (at random) and go to step (i)
If the cumulative variance explained is close enough to the specified threshold (0.95*VE,1.05*VE), then
If the number of introduced variants is over 35, quit and restart, else:
Accept the sampling and simulate data using the variants and effect sizes chosen, using HAPGEN2.
The upper bound of 35 on the total number of causal variants introduced per locus was imposed due to instability in HAPGEN2 behavior above this threshold; this limit was rarely reached in 3K samples, but it did restrict architectures simulated in 10K samples (S11C and S12 Figs).
The calculation of variance explained at each locus was conducted using the method described by So et al., which is available online as an R script [26]. This calculation requires three parameters as input (per variant): the prevalence of the trait (in this case assumed to be 8%, to model type 2 diabetes), the population frequency of the risk allele, and the genotype relative risk. We assumed independence between risk variants at a given locus, and thus estimated the total percentage of variance explained as the sum of the variance explained by each individual variant.
Running gene-based tests of association on simulated data
The latest releases of the PLINK/SEQ (v0.09) [18] and EPACTS (v3.2.3) [25] software packages were used to run ten of the gene-based methods evaluated in this study. MiST was run using a publicly available R package (http://cran.r-project.org/web/packages/MiST/index.html) [24]. All exonic variants (causal and non-causal) below varying minor allele frequency thresholds (1% for all analyses discussed in the main text, unless otherwise stated) were included in the tests, except when the fraction of neutral variation was varied. In this case, the proportion of causal variants included in the test was fixed to 0.25, 0.50, 0.75, or 1 (Fig 4C).
Selecting a subset of tests for joint application to the data
The subsets of tests chosen for inclusion into composite tests were selected using a stepwise forward selection approach. Starting with a single test (three runs per architecture, each starting with one of the top three performing tests across architectures, MiST, SKAT-O and KBAC), the next test to be included at each step was the one which reported the greatest number of novel signals, i.e. not previously detected by the tests already included. Novel signals were defined as loci for which the p-value reported by the candidate test for inclusion was lower by a specified multiplicative “margin” (factor) than the lowest p-value reported by tests already included in the composite test. Three margins were used (100, 10, and 1); a margin of 100, for example, implies that for signals to be considered novel, they p-value of the candidate test needs to be two orders of magnitude lower than the lowest of the ones already included in the composite test.
Datasets and software
All datasets discussed in this study, together with the scripts used to generate them and results of both single variant association and gene-based methods across all architectures, are available on the website http://mccarthy.well.ox.ac.uk/publications/2014/moutsianas_simulations/. The website also contains the software used for the script generation (a wrapper for HAPGEN2 [12]), which can be used to generate analogous simulated data for the genes we included in the manuscript under alternative scenarios/architectures.
Supporting Information
Zdroje
1. Purcell S., Cherny S.S. & Sham P.C. Genetic Power Calculator: design of linkage and association genetic mapping studies of complex traits. Bioinformatics 19, 149–50 (2003). 12499305
2. Kiezun A. et al. Exome sequencing and the genetic basis of complex traits. Nature Genetics 44, 623–30 (2012). doi: 10.1038/ng.2303 22641211
3. Asimit J. & Zeggini E. Rare variant association analysis methods for complex traits. Annual Review of Genetics 44, 293–308 (2010). doi: 10.1146/annurev-genet-102209-163421 21047260
4. Stitziel N.O., Kiezun A. & Sunyaev S. Computational and statistical approaches to analyzing variants identified by exome sequencing. Genome Biology 12, 227 (2011). doi: 10.1186/gb-2011-12-9-227 21920052
5. Rivas M. et al. Deep resequencing of GWAS loci identifies independent rare variants associated with inflammatory bowel disease. Nature Genetics 43, 1066–73 (2011). doi: 10.1038/ng.952 21983784
6. Cohen J.C., Boerwinkle E., Mosley T.H. & Hobbs H.H. Sequence variations in PCSK9, low LDL, and protection against coronary heart disease. New England Journal of Medicine 1264–1272 (2006). 16554528
7. Johansen C.T. et al. Excess of rare variants in genes identified by genome-wide association study of hypertriglyceridemia. Nature Genetics 42, 684–7 (2010). doi: 10.1038/ng.628 20657596
8. Bonnefond A. et al. Rare MTNR1B variants impairing melatonin receptor 1B function contribute to type 2 diabetes. Nature Genetics 44, 297–301 (2012). doi: 10.1038/ng.1053 22286214
9. Bansal V., Libiger O., Torkamani A.L.I. & Schork N.J. An application and empirical comparison of statistical analysis methods for associating rare variants to a complex phenotype. Pac Symp Biocomput 76–87 (2011). 21121035
10. Ladouceur M., Dastani Z., Aulchenko Y.S., Greenwood C.M.T. & Richards J.B. The empirical power of rare variant association methods: results from Sanger sequencing in 1,998 individuals. PLoS Genetics 8, e1002496 (2012). doi: 10.1371/journal.pgen.1002496 22319458
11. Basu S. & Pan W. Comparison of statistical tests for disease association with rare variants. Genetic Epidemiology 35, 606–619 (2011). doi: 10.1002/gepi.20609 21769936
12. Su Z., Marchini J. & Donnelly P. HAPGEN2: simulation of multiple disease SNPs. Bioinformatics 27, 2304–5 (2011). doi: 10.1093/bioinformatics/btr341 21653516
13. The 1000 Genomes Project Consortium A map of human genome variation from population-scale sequencing. Nature 467, 1061–73 (2010). doi: 10.1038/nature09534 20981092
14. Nelson M. et al. An abundance of rare functional variants in 202 drug target genes sequenced in 14,002 People. Science 337, 100–104 (2012). doi: 10.1126/science.1217876 22604722
15. Agarwala V., Flannick J., Sunyaev S. & Altshuler D. Evaluating empirical bounds on complex disease genetic architecture. Nature Genetics 45, 1418–27 (2013). doi: 10.1038/ng.2804 24141362
16. Li B. & Leal S.M. Methods for detecting associations with rare variants for common diseases: Application to analysis of sequence data. The American Journal of Human Genetics 83(3)311–321 (2008).
17. Price A.L. et al. Pooled association tests for rare variants in exon-resequencing studies. American Journal of Human Genetics 86, 832–8 (2010). doi: 10.1016/j.ajhg.2010.04.005 20471002
18. PLINK/SEQ: A library for the analysis of genetic variation data. at <http://atgu.mgh.harvard.edu/plinkseq/>
19. Madsen B.E. & Browning S.R. A groupwise association test for rare mutations using a weighted sum statistic. PLoS Genetics 5, e1000384 (2009). doi: 10.1371/journal.pgen.1000384 19214210
20. Liu D.J. & Leal S.M. A novel adaptive method for the analysis of next-generation sequencing data to detect complex trait associations with rare variants due to gene main effects and interactions. PLoS Genetics 6, e1001156 (2010). doi: 10.1371/journal.pgen.1001156 20976247
21. Neale B.M. et al. Testing for an unusual distribution of rare variants. PLoS Genetics 7, e1001322 (2011). doi: 10.1371/journal.pgen.1001322 21408211
22. Wu S. et al. Rare variant association testing for sequencing data using the Sequence Kernel Association Test (SKAT). American Journal of Human Genetics 89, 82–93 (2011). doi: 10.1016/j.ajhg.2011.05.029 21737059
23. Lee S. et al. Optimal unified approach for rare-variant association testing with application to small-sample case-control whole-exome sequencing studies. American Journal of Human Genetics 91, 224–37 (2012). doi: 10.1016/j.ajhg.2012.06.007 22863193
24. Sun J., Zheng Y. & Hsu L. A unified mixed-effects model for rare-variant association in sequencing studies. Genetic Epidemiology 37, 334–44 (2013). doi: 10.1002/gepi.21717 23483651
25. EPACTS: Efficient and Parallelizable Association Container Toolbox. <http://genome.sph.umich.edu/wiki/EPACTS>.
26. So H.-C., Gui A.H.S., Cherny S.S. & Sham P.C. Evaluating the heritability explained by known susceptibility variants: a survey of ten complex diseases. Genetic Epidemiology 35, 310–7 (2011). doi: 10.1002/gepi.20579 21374718
27. Falconer D.S. The inheritance of liability to diseases with variable age of onset, with particular reference to diabetes mellitus. Annals of Human Genetics 31, 1–20 (1967). 6056557
28. Lohmueller KE, Sparso T et al. Whole-exome sequencing of 2,000 Danish individuals and the role of rare coding variants in type 2 diabetes. Am. J. Human Genetics. 93, 1072–86 (2013). doi: 10.1016/j.ajhg.2013.11.005 24290377
29. Fu W, O’Connor TD, et al. Analysis of 6,515 exomes reveals the recent origin of most human protein-coding variants. Nature. 493, 216–220 (2013). doi: 10.1038/nature11690 23201682
30. Lee S., Abecasis G.R., Boehnke M., & Lin X. Rare-Variant Association Analysis: Study Designs and Statistical Tests. American Journal of Human Genetics 95, 5–23 (2014). doi: 10.1016/j.ajhg.2014.06.009 24995866
31. Li N & Stephens M. Modeling linkage disequilibrium and identifying recombination hotspots using single-nucleotide polymorphism data. Genetics 165 (4): 2213–33 (2003). 14704198
Štítky
Genetika Reprodukční medicína
Článek Retraction: Astakine 2—the Dark Knight Linking Melatonin to Circadian Regulation in CrustaceansČlánek Adventures in WonderlandČlánek Genomic Location of the Major Ribosomal Protein Gene Locus Determines Global Growth and InfectivityČlánek Spatial Fluctuations in Expression of the Heterocyst Differentiation Regulatory Gene in FilamentsČlánek Genome-Wide Negative Feedback Drives Transgenerational DNA Methylation Dynamics in ArabidopsisČlánek Systematic Dissection of the Sequence Determinants of Gene 3’ End Mediated Expression ControlČlánek The Chromatin Remodeler CHD8 Is Required for Activation of Progesterone Receptor-Dependent EnhancersČlánek Selection against Heteroplasmy Explains the Evolution of Uniparental Inheritance of MitochondriaČlánek The DNA Helicase Recql4 Is Required for Normal Osteoblast Expansion and Osteosarcoma FormationČlánek Dual-Specificity Anti-sigma Factor Reinforces Control of Cell-Type Specific Gene Expression in
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2015 Číslo 4
-
Všechny články tohoto čísla
- Retraction: Astakine 2—the Dark Knight Linking Melatonin to Circadian Regulation in Crustaceans
- Adventures in Wonderland
- Experimental Swap of 's Assortative Mating Preferences Demonstrates Key Role of X-Chromosome Divergence Island in Incipient Sympatric Speciation
- Chromosome Replacement and Deletion Lead to Clonal Polymorphism of Berry Color in Grapevine
- The Protein Quality Control Machinery Regulates Its Misassembled Proteasome Subunits
- Genome-Wide Association Study Identifies as a Critical Gene for Susceptibility to Noise-Induced Hearing Loss
- Genomic Location of the Major Ribosomal Protein Gene Locus Determines Global Growth and Infectivity
- Viable Neuronopathic Gaucher Disease Model in Medaka () Displays Axonal Accumulation of Alpha-Synuclein
- Multi-locus Analysis of Genomic Time Series Data from Experimental Evolution
- The Genetic Legacy of the Expansion of Turkic-Speaking Nomads across Eurasia
- Lack of GDAP1 Induces Neuronal Calcium and Mitochondrial Defects in a Knockout Mouse Model of Charcot-Marie-Tooth Neuropathy
- The Pif1 Helicase, a Negative Regulator of Telomerase, Acts Preferentially at Long Telomeres
- Inhibiting K63 Polyubiquitination Abolishes No-Go Type Stalled Translation Surveillance in
- SYD-1C, UNC-40 (DCC) and SAX-3 (Robo) Function Interdependently to Promote Axon Guidance by Regulating the MIG-2 GTPase
- Spatial Fluctuations in Expression of the Heterocyst Differentiation Regulatory Gene in Filaments
- Synergistic and Independent Actions of Multiple Terminal Nucleotidyl Transferases in the 3’ Tailing of Small RNAs in Arabidopsis
- Host Genetic Variation Influences Gene Expression Response to Rhinovirus Infection
- Contribution of Large Region Joint Associations to Complex Traits Genetics
- Volatility of Mutator Phenotypes at Single Cell Resolution
- Proteolysis of Virulence Regulator ToxR Is Associated with Entry of into a Dormant State
- Genome-Wide Negative Feedback Drives Transgenerational DNA Methylation Dynamics in Arabidopsis
- A Multi-layered Protein Network Stabilizes the FtsZ-ring and Modulates Constriction Dynamics
- Systematic Dissection of the Sequence Determinants of Gene 3’ End Mediated Expression Control
- Genome Sequencing of the Perciform Fish Provides Insights into Molecular and Genetic Mechanisms of Stress Adaptation
- Natural Variant E610G Is a Semi-dominant Suppressor of IAP-Induced RNA Processing Defects
- The Alkaline Response Pathway: Identification of a Novel Rim Pathway Activator
- Transgenerational Inheritance of Diet-Induced Genome Rearrangements in Drosophila
- A Single Nucleotide Polymorphism Uncovers a Novel Function for the Transcription Factor Ace2 during Hyphal Development
- DNA Damage Response and Spindle Assembly Checkpoint Function throughout the Cell Cycle to Ensure Genomic Integrity
- The Functional Interplay Between the t(9;22)-Associated Fusion Proteins BCR/ABL and ABL/BCR in Philadelphia Chromosome-Positive Acute Lymphatic Leukemia
- Extreme Recombination Frequencies Shape Genome Variation and Evolution in the Honeybee,
- Beyond Glycolysis: GAPDHs Are Multi-functional Enzymes Involved in Regulation of ROS, Autophagy, and Plant Immune Responses
- Comprehensive Profiling of Amino Acid Response Uncovers Unique Methionine-Deprived Response Dependent on Intact Creatine Biosynthesis
- Windpipe Controls Intestinal Homeostasis by Regulating JAK/STAT Pathway via Promoting Receptor Endocytosis and Lysosomal Degradation
- Ataxin-2 Regulates Translation in a New BAC-SCA2 Transgenic Mouse Model
- Cross-Population Joint Analysis of eQTLs: Fine Mapping and Functional Annotation
- The Power of Gene-Based Rare Variant Methods to Detect Disease-Associated Variation and Test Hypotheses About Complex Disease
- The Chromatin Remodeler CHD8 Is Required for Activation of Progesterone Receptor-Dependent Enhancers
- Competition between VanU Repressor and VanR Activator Leads to Rheostatic Control of Vancomycin Resistance Operon Expression
- A Missense Change in the Gene Links Aberrant Autophagy to a Neurodegenerative Vacuolar Storage Disease
- Simultaneous Discovery, Estimation and Prediction Analysis of Complex Traits Using a Bayesian Mixture Model
- Selection against Heteroplasmy Explains the Evolution of Uniparental Inheritance of Mitochondria
- Genome-Destabilizing Effects Associated with Top1 Loss or Accumulation of Top1 Cleavage Complexes in Yeast
- Imputation-Based Population Genetics Analysis of Malaria Parasites
- Heterozygosity for a Hypomorphic Polβ Mutation Reduces the Expansion Frequency in a Mouse Model of the Fragile X-Related Disorders
- Neto-Mediated Intracellular Interactions Shape Postsynaptic Composition at the Neuromuscular Junction
- Ndd1 Turnover by SCF Is Inhibited by the DNA Damage Checkpoint in
- Frameshift Variant Associated with Novel Hoof Specific Phenotype in Connemara Ponies
- The DNA Helicase Recql4 Is Required for Normal Osteoblast Expansion and Osteosarcoma Formation
- Spastin Binds to Lipid Droplets and Affects Lipid Metabolism
- Maintenance of Glia in the Optic Lamina Is Mediated by EGFR Signaling by Photoreceptors in Adult Drosophila
- Auxin Influx Carriers Control Vascular Patterning and Xylem Differentiation in
- Dual-Specificity Anti-sigma Factor Reinforces Control of Cell-Type Specific Gene Expression in
- The Lowe Syndrome Protein OCRL1 Is Required for Endocytosis in the Zebrafish Pronephric Tubule
- Postnatal Loss of Hap1 Reduces Hippocampal Neurogenesis and Causes Adult Depressive-Like Behavior in Mice
- CAPER Is Vital for Energy and Redox Homeostasis by Integrating Glucose-Induced Mitochondrial Functions via ERR-α-Gabpa and Stress-Induced Adaptive Responses via NF-κB-cMYC
- Distinct and Cooperative Activities of HESO1 and URT1 Nucleotidyl Transferases in MicroRNA Turnover in
- The Evolutionary Origination and Diversification of a Dimorphic Gene Regulatory Network through Parallel Innovations in and
- MAPK Signaling Pathway Alters Expression of Midgut ALP and ABCC Genes and Causes Resistance to Cry1Ac Toxin in Diamondback Moth
- Spatio-temporal Remodeling of Functional Membrane Microdomains Organizes the Signaling Networks of a Bacterium
- Asymmetric Transcript Discovery by RNA-seq in . Blastomeres Identifies , a Gene Important for Anterior Morphogenesis
- A Stress-Induced Small RNA Modulates Alpha-Rhizobial Cell Cycle Progression
- Systematic Profiling of Poly(A)+ Transcripts Modulated by Core 3’ End Processing and Splicing Factors Reveals Regulatory Rules of Alternative Cleavage and Polyadenylation
- The UPR Branch IRE1- in Plants Plays an Essential Role in Viral Infection and Is Complementary to the Only UPR Pathway in Yeast
- A Non-canonical RNA Silencing Pathway Promotes mRNA Degradation in Basal Fungi
- Co-chaperone p23 Regulates . Lifespan in Response to Temperature
- Re-replication of a Centromere Induces Chromosomal Instability and Aneuploidy
- Shade Avoidance Components and Pathways in Adult Plants Revealed by Phenotypic Profiling
- Lipid-Induced Epigenomic Changes in Human Macrophages Identify a Coronary Artery Disease-Associated Variant that Regulates Expression through Altered C/EBP-Beta Binding
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Lack of GDAP1 Induces Neuronal Calcium and Mitochondrial Defects in a Knockout Mouse Model of Charcot-Marie-Tooth Neuropathy
- Proteolysis of Virulence Regulator ToxR Is Associated with Entry of into a Dormant State
- Frameshift Variant Associated with Novel Hoof Specific Phenotype in Connemara Ponies
- Ataxin-2 Regulates Translation in a New BAC-SCA2 Transgenic Mouse Model
Přihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



