-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaA Newly Uncovered Group of Distantly Related Lysine Methyltransferases Preferentially Interact with Molecular Chaperones to Regulate Their Activity
Methylation is a post-translational modification that can affect numerous features of proteins, notably cellular localization, turnover, activity, and molecular interactions. Recent genome-wide analyses have considerably extended the list of human genes encoding putative methyltransferases. Studies on protein methyltransferases have revealed that the regulatory function of methylation is not limited to epigenetics, with many non-histone substrates now being discovered. We present here our findings on a novel family of distantly related putative methyltransferases. Affinity purification coupled to mass spectrometry shows a marked preference for these proteins to associate with various chaperones. Based on the spectral data, we were able to identify methylation sites in substrates, notably trimethylation of K135 of KIN/Kin17, K561 of HSPA8/Hsc70 as well as corresponding lysine residues in other Hsp70 isoforms, and K315 of VCP/p97. All modification sites were subsequently confirmed in vitro. In the case of VCP, methylation by METTL21D was stimulated by the addition of the UBX cofactor ASPSCR1, which we show directly interacts with the methyltransferase. This stimulatory effect was lost when we used VCP mutants (R155H, R159G, and R191Q) known to cause Inclusion Body Myopathy with Paget's disease of bone and Fronto-temporal Dementia (IBMPFD) and/or familial Amyotrophic Lateral Sclerosis (ALS). Lysine 315 falls in proximity to the Walker B motif of VCP's first ATPase/D1 domain. Our results indicate that methylation of this site negatively impacts its ATPase activity. Overall, this report uncovers a new role for protein methylation as a regulatory pathway for molecular chaperones and defines a novel regulatory mechanism for the chaperone VCP, whose deregulation is causative of degenerative neuromuscular diseases.
Published in the journal: . PLoS Genet 9(1): e32767. doi:10.1371/journal.pgen.1003210
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1003210Summary
Methylation is a post-translational modification that can affect numerous features of proteins, notably cellular localization, turnover, activity, and molecular interactions. Recent genome-wide analyses have considerably extended the list of human genes encoding putative methyltransferases. Studies on protein methyltransferases have revealed that the regulatory function of methylation is not limited to epigenetics, with many non-histone substrates now being discovered. We present here our findings on a novel family of distantly related putative methyltransferases. Affinity purification coupled to mass spectrometry shows a marked preference for these proteins to associate with various chaperones. Based on the spectral data, we were able to identify methylation sites in substrates, notably trimethylation of K135 of KIN/Kin17, K561 of HSPA8/Hsc70 as well as corresponding lysine residues in other Hsp70 isoforms, and K315 of VCP/p97. All modification sites were subsequently confirmed in vitro. In the case of VCP, methylation by METTL21D was stimulated by the addition of the UBX cofactor ASPSCR1, which we show directly interacts with the methyltransferase. This stimulatory effect was lost when we used VCP mutants (R155H, R159G, and R191Q) known to cause Inclusion Body Myopathy with Paget's disease of bone and Fronto-temporal Dementia (IBMPFD) and/or familial Amyotrophic Lateral Sclerosis (ALS). Lysine 315 falls in proximity to the Walker B motif of VCP's first ATPase/D1 domain. Our results indicate that methylation of this site negatively impacts its ATPase activity. Overall, this report uncovers a new role for protein methylation as a regulatory pathway for molecular chaperones and defines a novel regulatory mechanism for the chaperone VCP, whose deregulation is causative of degenerative neuromuscular diseases.
Introduction
Methyltransferases catalyze the transfer of a methyl group (CH3) from a donor, generally S-adenosyl-L-methionine (AdoMet), to various acceptor molecules such as proteins, DNA, RNA, lipids, and small metabolites [1]–[3]. Protein N-methylation predominantly targets the side chains of two amino acids, lysine and arginine, whereas the side chains of other residues, including histidine, glutamine, and asparagine represent minor targets for methylation [4]–[6]. Dicarboxylic amino acids (glutamate, aspartate) and cysteine are also known to be respectively O - and S-methylated on occasion [7], [8]. In addition, some proteins were shown to be methylated on their N-terminal and C-terminal ends [9]–[11]. The vast majority of methyltransferases are grouped into three large families based on their structure, namely seven-β-strand, SET and SPOUT domain-containing methyltransferases [2]. All protein R methyltransferases (PRMT) are part of the seven-β-strand superfamily, while protein K methyltransferases (PKMT) fall almost exclusively within the SET domain-containing group. Until recently, the only known seven-β-strand PKMT was Dot1 [12].
Efforts to characterize substrates of PKMT have mostly focused on nucleosome components. Methylation of histone H3 residues K4, K36, and K79 are associated with transcriptionally active euchromatin, whereas methylation of H3K9, H3K27 and H4K20 coincides with heterochromatin and transcriptional repression [13], [14]. Recent reports have furthermore shown that the type of lysine methylation (i.e., mono-, di - or trimethylation) should also be taken into consideration when assessing chromatin state [15]–[17]. Epigenetics has been paramount in demonstrating that a modification as seemingly insignificant as the addition of a methyl group can have a considerable impact on a biological process as crucial as gene expression. Evidence of lysine methylation-driven regulation has been documented for an ever-increasing number of non-histone proteins, including calmodulin, cytochrome C, Rubisco, ribosomal proteins, p53, and NF-κB [18]–[27].
As part of an effort to systematically map protein-protein interactions, we came across a previously uncharacterized protein sharing distant homology with PRMTs nestled within a network of molecular chaperones involved in protein complex assembly. Subsequent local alignement searches using that protein as seed uncovered a group of 10 distantly related putative methyltransferases. Characterization of the interaction network of this novel subgroup of methyltransferases was undertaken by Affinity Purification coupled to Mass Spectrometry (AP-MS) and then computationally assessed. Our results revealed that enzymes of this subgroup preferentially interact with molecular chaperones. Validation experiments using three of the identified interactors, Kin17, Hsc70, and VCP/p97, indicated that they represent bona fide substrates. In each case, trimethylated lysine residues were identified in vivo and confirmed in vitro using recombinant methyltransferase-substrate pairs. In addition, we have shown that methylation of one of these substrates, VCP/p97, by METTL21D can be modulated by ASPSCR1/UBXD9 and that this modification regulates ATPase activity of the VCP chaperone. The results presented here bring to light an entirely new cast of PKMTs of the seven-β-strand variety and expands our knowledge of non-histones substrates, most notably molecular chaperones. This finding points to a new role for protein methylation in regulating protein folding, quality control, and turnover.
Results
Characterization of a newly discovered family of distantly related putative methyltransferases
The study of this group of previously uncharacterized methyltransferases was initiated when METTL22 was identified in the soluble fraction of a protein affinity purification that targeted the DNA/RNA binding protein Kin17/KIN. Local alignment searches were performed to ascertain the function of this protein (data not shown). It was discovered that METTL22 was part of a larger group of 10 proteins (if the diversity of FAM86 closely-related isoforms are considered as a single member) that shared distant homology with PRMTs. Phylogenetic analysis of the most conserved region of these two protein groups (Figure S1) confirmed this observation, suggesting that this family of uncharacterized methyltransferases is related to, but distinct from, PRMTs (Figure 1A; Figure S2). Computational structure prediction further demonstrated the similarity between the members of this family of putative methyltransferases and PRMTs (Figure 1B, 1C). The subsequent publication of the human “methyltransferasome” by Clarke and colleagues confirmed that these proteins are putative methyltransferases and that they form a distinct family [2]. Indeed, most of the methyltransferases described here fall within the so-called “Group J.”
Fig. 1. Computational analysis defines a novel family of putative protein methyltransferases. 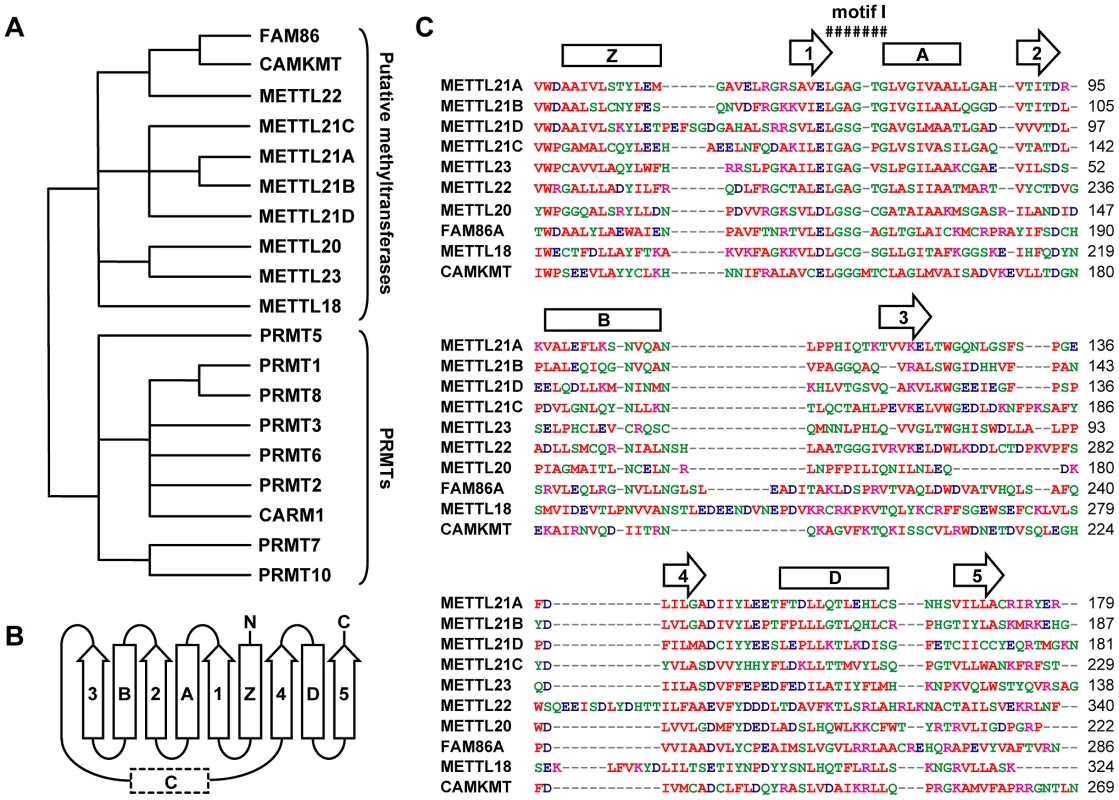
(A) Unrooted phylogenetic tree of a family of human putative methyltransferases distantly related to PRMTs. FAM86 represents a number of genetic variants (FAM86A, B1, B2, C, and D) whose duplication is observed only in primates. Branch lengths are not proportional to the actual evolutionary distances between the sequences. (B) Secondary structure organization of the Rossmann fold domain of PRMTs responsible for the methyltransferase activity. Arrows represent β strands and rectangles correspond to α helices, including typically ill-defined or inexistent α helix C. (C) Multiple sequence alignment of the Rossmann fold of all members within this family as generated by ClustalW2 (http://www.ebi.ac.uk/Tools/msa/clustalw2/). Red residues are small, hydrophobic, aromatic; blue are acidic; magenta are basic; and all other residues are green. Primary sequence alignment corresponds nicely with secondary structure prediction by Jpred3 (http://www.compbio.dundee.ac.uk/www-jpred/). Overhead β strands and α helices are shown as in (B). Conserved motif I, the site of S-adenosylmethionine binding, is also marked. Based on the observed homology with PRMTs, we hypothesized that these proteins were likely protein methyltransferases themselves. To identify possible substrates and cofactors, we elected to map the protein interaction network for each member of this novel family by AP-MS (main interactors are marked in Figure 2, additional targets are listed in Table S1) [28]–[35]. The main METTL22 interactor identified in the soluble fraction was KIN, which confirmed our initial observation and further strengthened the notion that these two factors interact. A common theme for most of these putative methyltransferases' interactors was chaperones, be they of the Hsp70 or Hsp90 variety (see METTL18, CAMKMT, METTL21C, METTL22, METTL23, METTL21A, and METTL21B), chaperonins HSPD1 and CCT (see METTL18, CAMKMT, METTL20, and METTL21B), and even the AAA ATPase VCP (see METTL21D) that is believed to act as a chaperone in various processes, most notably Endoplasmic Reticulum-Associated Protein Degradation (ERAD) [36]–. Recent publications have substantiated the accuracy of this interactome. Firstly, it was demonstrated in another report by Steven Clarke on the yeast homolog of METTL18, YIL110W, that this protein methylates the ribosomal protein RPL3 [6]. Our own purification of METTL18 identified RPL3 and its associated ribosome biogenesis factor GRWD1 [41] as the two main interactors. Secondly, CAMKMT has been shown to methylate calmodulin on a lysine residue [42]. We were likewise able to co-purify calmodulin protein CALM2 in our CAMKMT affinity purification. However, it should be noted that since our AP-MS protocol includes beads bearing the calmodulin-binding peptide (CBP), calmodulin often appears as a non-specific target, although usually with a weaker signal. Computational assessment showed that CALM2 was a high confidence interactor of CAMKMT (FDR<10%).
Fig. 2. Tandem-affinity purification coupled to mass spectrometry reveals protein interaction network of putative methyltransferases. 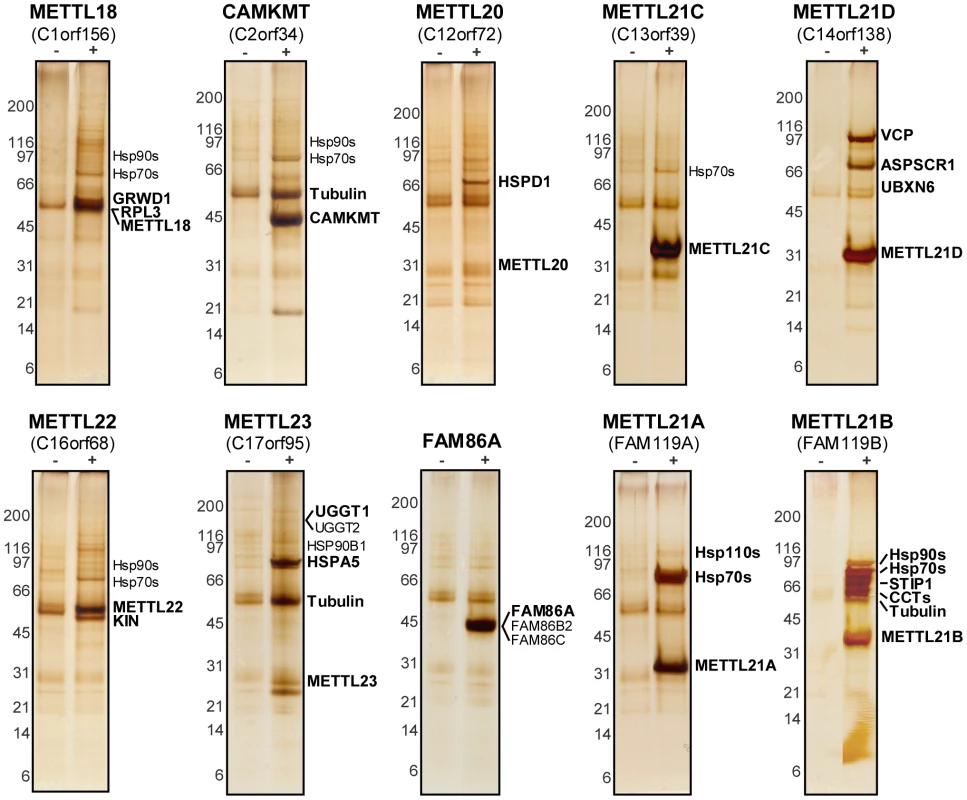
Purification of 10 TAP-tagged putative methyltransferases from ponasterone-inducible strains of HEK 293 cells. Eluted proteins were separated by SDS-PAGE. Gels were silver stained and cut in slices that were then trypsin digested before protein identification by LC-MS/MS. Tagged baits and major interactors are marked. Protein database searches were repeated allowing for mono-, di-, and trimethylation of lysine residues (as a variable modification). Of note, lysine trimethylation and acetylation are sometimes mis-annotated due to the closeness in mass of these modifications (+42.0468 Dalton and +42.0105 Dalton, respectively) [43]. Fortunately, the high mass accuracy obtained with the LTQ Orbitrap mass spectrometer was sufficient to distinguish between these PTMs. The most promising hits were a trimethylated lysine on KIN at position 135 in the METTL22 purification, another trimethylated lysine at position 315 on VCP in the METTL21D purification, and a number of trimethylated lysines on multiple Hsp70 isoforms, which correspond to a homologous site, in the METTL21A purification (Figure 3A; see corresponding mass spectra in Figure S3). These methylation sites were highly conserved through evolution (Figure 3B). Conservation of the VCP methylation site K315 is not surprising considering the relative immutability of the overall protein (roughly 70% sequence identity from Homo sapiens to Saccharomyces cerevisiae). The target lysine 561 in HSPA8/Hsc70 was likewise conserved through evolution and orthologs are found in species as distant as S. cerevisiae. Moreover, and as mentioned previously, this site is also retained in a number human Hsp70 paralogs (HSPA1, HSPA1L, HSPA2, HSPA5, and HSPA6). In fact the only conserved residue in this region of loose homology is the target lysine, pointing to a possible important regulatory role for this modification. The target lysine in KIN, K135, is present in a number of species, including Arabidopsis thaliana and Drosophila melanogaster, but is absent in Saccharomyces cerevisiae and Plasmodium falciparum. Interestingly, METTL22 orthologs are concurrently absent in species where the corresponding lysine in KIN is not conserved, which further suggests a strong link between the two.
Fig. 3. KIN, VCP, and a number of hsp70 isoforms are each trimethylated on lysine residues by specific methyltransferases within this family. 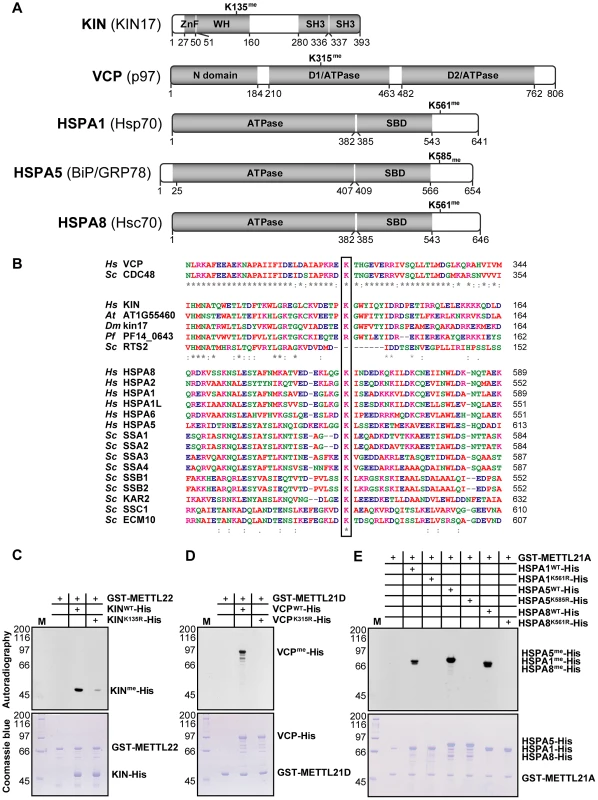
(A) Linear representation of all identified substrates with domain architecture. Residues delineating each domain are marked below. ZnF, Zinc Finger; WH, Winged Helix; SH3, Src Homology 3; SBD, substrate binding domain. Position of the methylated lysines is shown above. (B) Multiple sequence alignment of the region surrounding trimethylated lysines (boxed) in human VCP, KIN, and Hsp70 isoforms compared to paralogous genes in various organisms. Hs, Homo sapiens; Sc, Saccharomyces cerevisiae; Dm, Drosophila melanogaster; At, Arabidopsis thaliana; Pf, Plasmodium falciparum. Color code is as in Figure 1. Strong and weak residue similarity are represented by a colon (:) and period (.), respectively, and asterisk (*) denotes identity. (C–E) In vitro methylation assays with tritium-labeled S-adenosylmethionine of KIN-His with GST-METTL22 (C), VCP-His with GST-METTL21D (D), and three His-tagged hsp70 isoforms (HSPA1, HSPA5, and HSPA8) with GST-METTL21A (E). In each case, substitution of the methylated lysine by an arginine leads to loss of methylation signal as detected by autoradiography. Coomassie staining of the gel shows total proteins loaded onto the gel and serves as control. We then proceeded with in vitro methylation assays to confirm the identity of these methylation targets (Figure 3C–3E). A positive signal was observed for each reaction, confirming that these are in fact protein methyltransferases. Moreover, substitution of each identified lysine to an arginine, a relatively conserved substitution, led to the abrogation of the methylation signal. In the case of KIN, the K135R substitution greatly diminished the methylation signal, but did not completely abolish it as with other mutants tested. This could mean that there might be a second methylation site on KIN, but given that no other methylated peptide was ever observed by mass spectrometry, either in the original purification of METTL22 or in the in vitro methylation reaction itself (see corresponding mass spectra in Figure S4), we believe that the residual methylation is more likely to occur on a cryptic site, i.e., one that is not normally methylated in wild-type KIN. Of note, methylation by METTL21A was assayed on three Hsp70 homologs (HSPA1, HSPA5, and HSPA8), but it stands to reason that the modification would also apply to other isoforms where the lysine residue is conserved.
To further characterize the function of the methyltransferases, intracellular localization was determined by immunofluorescence. To this end, recombinant FLAG-tagged proteins were expressed in HeLa cells (Figure 4). For most methyltransferases, a marked preference for the cytoplasm was observed, although this trend is reversed in METTL21C and METT22, where the localization is predominantly nuclear. This nuclear distribution could hint at a nucleosomal methylation activity, since it is frequent with most other protein methyltransferases, but none of the four major histones appear to be methylated in vitro by the members of this family (see Figure S5). Localization of METTL18 was never determined, since no significant expression of the recombinant methyltransferase has ever been observed. It is tempting to speculate that this effect could be the result of impaired translation, since METTL18 interacts with, and probably methylates, ribosomal subunit RPL3. Whereas most methyltransferases display a diffuse distribution, METTL20 is concentrated in cytoplasmic granular foci and METTL23 displays internal membrane-like structures.
Fig. 4. Intracellular distribution of putative methyltransferases and colocalization with identified substrates as shown by immunofluorescence. 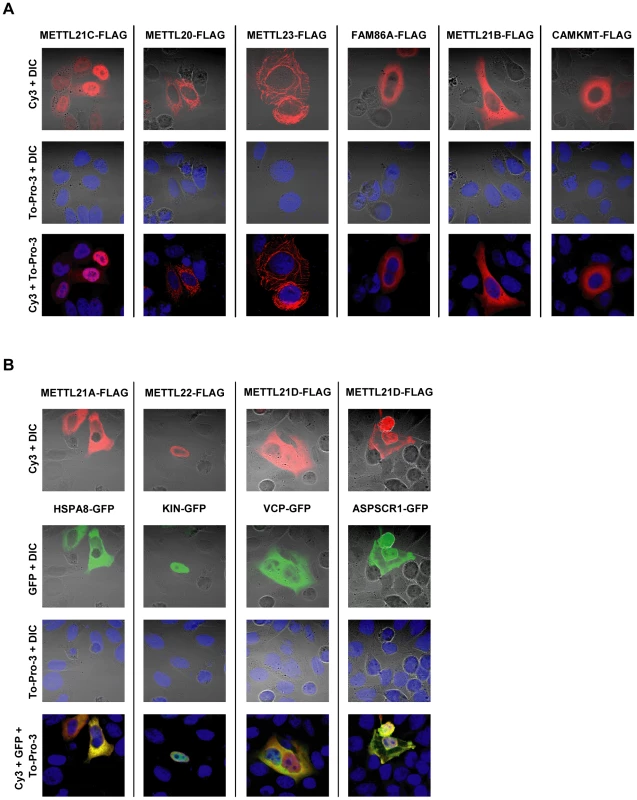
(A) HeLa cells were transiently transfected with vectors encoding C-terminally FLAG-tagged putative methyltransferases. Localization of the recombinant proteins is revealed by immunofluorescence with a Cy3-labelled antibody. DNA is stained with To-Pro-3 to determine the position of the nucleus, and overall morphology of the cell is shown by Differential Interference Contrast microscopy (DIC). (B) Comparative localization of methyltransferase with associated proteins is shown by transient cotransfection of vectors for the expression of FLAG-tagged methyltransferase and C-terminally GFP-tagged interactors. In the three instances where a methylation target has been identified, the substrate proteins (or associated protein ASPSCR1, in the case of METTL21D) bearing a GFP marker were co-expressed with the corresponding methyltransferase (Figure 4B). All methyltransferases and methyl acceptors more or less colocalize within the same cell compartments. In the case of METTL21A with HSPA8 and METTL21D with VCP or ASPSCR1, the colocalization is nearly perfect. METTL22 and KIN are both present in the nucleus, although METTL22 is clearly more concentrated in the periphery than KIN.
ASPSCR1/UBXD9 interacts with METTL22 to stimulate methylation of VCP
Methyltransferases often require cofactors to aid in the modification of their substrates. A good example of this is methylation of spliceosomal Sm proteins by the PRMT5/WD45 complex with the help of pICIn [44]. In the purification of METTL21D, we were able to identify two poorly documented VCP binding proteins, UBXN6/UBXD1 and ASPSCR1/UBXD9. This came as a surprise since VCP was shown to interact with an impressive number of cofactors including the entirety of the UBX (ubiquitin regulatory X) family [45]. Given this apparent specificity, we tested whether these proteins could act as cofactors in the methylation of VCP. As shown in Figure 5A, methylation experiments revealed that neither ASPSRC1 nor UBXN6 could be methylated directly by METTL21D but that only the addition of ASPSCR1, not UBXN6, could enhance methylation of VCP. N - and C-terminal fragments of ASPSCR1 were generated in an effort to determine which domain of the cofactor is responsible for this effect (Figure 5B). To our surprise, we observed that only the C-terminal fragment (residues 280–553), which was previously shown to interact weakly with VCP [46], could enhance VCP methylation in a similar manner as full-length ASPSCR1. An in vitro GST pull-down experiment (Figure 5C) confirms direct binding of the methyltransferase METTL21D to its substrate VCP, but also shows interaction with ASPSCR1, more specifically, to its C-terminal fragment. Furthermore, addition of VCP and ASPSCR1 or VCP and the C-terminal fragment of ASPSCR1 together appear to have a synergetic effect on binding to METTL21D, which could account for the concomitant increase in methylation signal.
Fig. 5. ASPSCR1 promotes methylation of VCP by METTL21D. 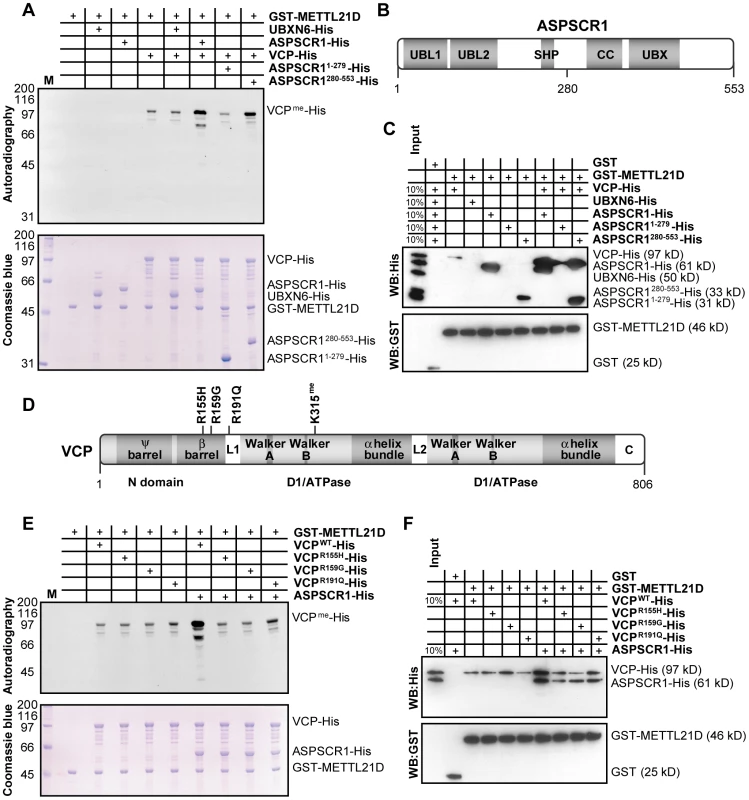
(A) In vitro methylation assays of VCP-His, UBXN6-His, ASPSCR1-His and fragments of ASPSCR1 with GST-METTL21D. Various combinations of the UBX proteins were added to reactions containing VCP. Coomassie staining of the gel is shown. (B) Linear representation of ASPSCR1 showing domain architecture of the protein (UBL, UBiquitin-Like domain; SHP, SHP box; UBX, UBiquitin regulatory X domain; CC, Coiled-Coil domain) and localization of residue 280 which marks the boundary between the N- and C- terminal fragments used in these experiments. (C) In vitro GST pull-down assays of VCP-His, UBXN6-His, ASPSCR1-His and fragments of ASPSCR1 with GST-METTL21D. Combinations of full-length ASPSCR1 and its fragments were once again employed with VCP. (D) Linear representation of VCP showing domain architecture of the protein (including double Ψ barrel superfold and 4-stranded β barrel of the N-terminal domain, Walker A and B motifs, as well as 4 α helices bundle of ATPase domains D1 and D2 as well as linker regions L1 and L2 and C-terminal domain) and localization of the mutants used in this study as well as the site of methylation. (E) In vitro methylation assays of wild-type VCP-His and IBMPFD and ALS-causing mutations R155H, R159G and R191Q in presence or absence of ASPSCR1-His. (F) In vitro GST pull-down assays of the same combination of proteins as in (E). Numerous mutations in the VCP gene have been linked with genetic disorders such as Inclusion Body Myopathy with Paget's disease of bone and Fronto-temporal Dementia (IBMPFD) and familial Amyotrophic Lateral Sclerosis (ALS) [47], [48]. Substitutions R155H and R191Q have been implicated in both IBMPFD and ALS. Furthermore, R159G was observed in patient with ALS, although other substitutions targeting arginine 159 were found in patients with IBMPFD (Figure 5D). In vitro methylation assay using recombinant VCP bearing these substitutions was done in order to test whether disease-causing mutations can also impact VCP methylation (Figure 5E). Although all mutant proteins appears to be methylated to a similar degree as wild-type VCP in vitro, the addition of the UBX protein no longer seems to enhance the methylation signal. These results can be explained by the notion the mutants used in this assay, as with most described VCP mutations, reside within the N-terminal domain believed to be involved in cofactor association [49]–[51]. In vitro GST pull-down experiment (Figure 5F) confirms that mutation of VCP has no impact on affinity of METTL21D for its substrate. However, when ASPSCR1 is added to the mix, the synergetic increase in binding is only observed with wild-type VCP.
Methylation of VCP negatively impacts on its ATPase activity
Given VCP's involvement in disease, we decided to further scrutinize the functional implications of its methylation. This member of the AAA (ATPases Associated with various cellular Activities) family of ATPases contains dual ATPase domains. The methylation site falls in close proximity to the Walker B motif of VCP's first ATPase domain (Figure 6A). Knowing that Walker B motifs are usually involved in ATP hydrolysis, we hypothesized that trimethylation of lysine 315 might affect the ATPase activity of this domain. To test this idea, in vitro ATPase assays were performed with a fragment of VCP spanning its N-terminal and first ATPase domain. The reasoning behind this was that since most of VCP's ATPase activity stems from its second ATPase domain [52], if methylation of K315 only affects the activity of the first ATPase domain, we might not have detected a change in the overall activity of the full-length protein. Before going forward with in vitro ATPase assays, we first verified that the fragment could still be methylated and that methylation could be inhibited by S-adenosylhomocysteine (AdoHcy), a byproduct of methylation that also acts as a methylation inhibitor for most methyltransferases. A catalytically inactive mutated form of the methyltransferase was also created that targets the conserved acidic residue in METTL21D's AdoMet-binding motif (E73Q, see Figure 1C). The results show that the fragment is methylated as efficiently as full-length VCP (Figure 6B). Furthermore, substitution of the VCP fragment by an unmethylatable mutant (K315R), substitution of the methyltransferase by the catalytically inactive mutant (E73Q), or even addition of AdoHcy all resulted in nearly complete inhibition of methylation.
Fig. 6. Methylation of VCP decreases the activity of its N-terminal ATPase domain. 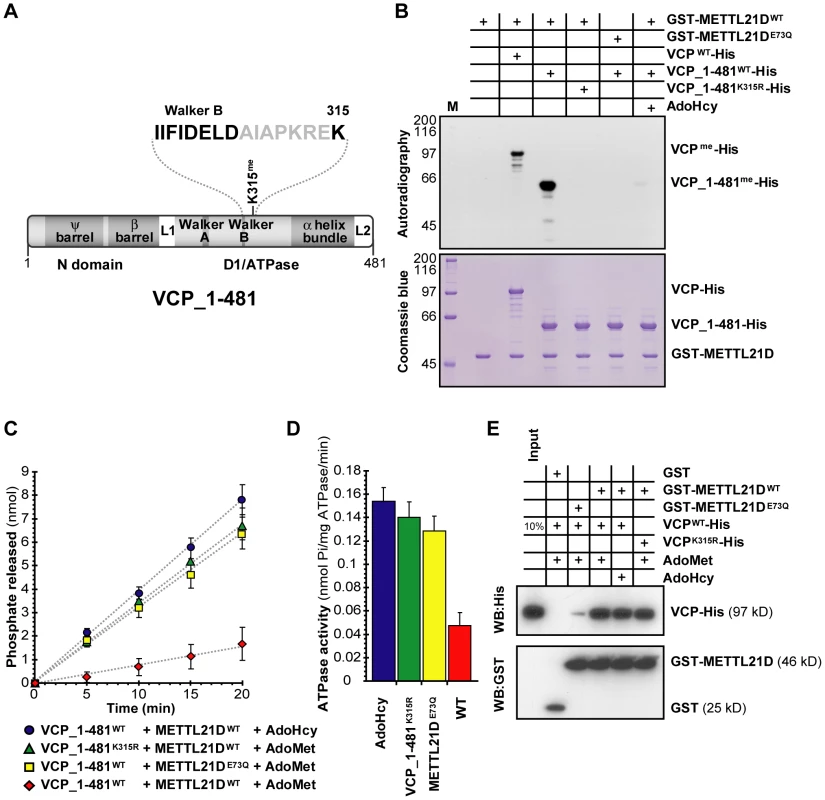
(A) Linear representation of a fragment of VCP encompassing its N-terminal and first ATPase domain employed in the ATPase assay. Proximity of the methylated lysine to the Walker B motif is highlighted above. (B) In vitro methylation assays of 1–481_VCP-His fragment by GST-METTL21D as compared to full length VCP. (C, D) Colorimetric assays to measure released phosphate (C) and relative ATPase activity (D) by the 1–481 fragment of VCP. The experiment was done in triplicate. Data from the last 3 time points (9 measurements in total for each condition) was compiled to generate the graph shown in (D). (E) In vitro GST pull-down assay of VCP-His with GST-METTL21D. In all experiments the effect of un methylatable VCP mutant K135R, catalytically inactive METTL21D mutant E73Q, and methylation inhibitor S-adenosylhomocysteine is shown. We then performed the in vitro ATPase assay and found that when a wild-type VCP fragment is pre-incubated with wild type METTL21D and the methyl donor, AdoMet, a significant decrease in ATPase activity was detected as compared to three separate control reactions where either the methyltransferase is replaced by its E73Q mutant; VCP is replaced with its unmethylatable mutant (K315R); or the methyl donor is replaced with AdoHcy (Figure 6C and 6D).
A possible interpretation of this finding is that methylation of VCP does not inhibit the ATPase activity per se, but that binding of the methyltransferase itself hinders the ATPase function. To eliminate this possibility, in vitro GST pulldowns were carried out (Figure 6E). Although we did observe a decreased binding between methyltransferase and substrate when METTL21D is replaced by the E73Q mutant, addition of AdoHcy and mutation of VCP lysine 315 do not appear to affect the interaction when compared to wild-type METTL21D and wild-type VCP in presence of AdoMet. This result confirms the conclusion that methylation of VCP directly modulates the ATPase activity of its first ATPase domain. Additionally, these experiments were repeated with a full-length form of VCP whose second ATPase domain has been inactivated by a mutation targeting a critical residue within the Walker B motif (E578Q; Figure S6) [53]. Again, a decrease in ATPase activity is observed when VCP_E578Q is preincubated with METTL21D and AdoMet.
Discussion
The data presented here bring an entirely new group of protein methyltransferases into light and suggest a role for this post-translational modification in modulating chaperone function. Hsp70 isoforms have been known to be methylated both on arginine and lysine residues for quite some time [54], [55], but up until now the exact sites of these modifications and the enzymes responsible for them had remained elusive. The role of these modifications is also uncertain, but we speculate that they may help direct specificity of these chaperones towards substrates and cofactors. Evidence for this could be derived from the AP-MS data presented here. Indeed, METTL21A, the only known Hsp70 methyltransferase identified so far, copurified with a number of Hsp70 isoforms but few cofactors aside from Hsp110s. The closely related METTL21B also copurified with significant amounts of Hsp70 but this time appeared to be complexed with STIP1/Hsp90 or CCT chaperonin. That differential methylation by these enzymes drives Hsp70 specificity is a hypothesis that remains to be tested.
What is certain based on the results presented in this article is that the ATPase activity of another seemingly unrelated chaperone, VCP, can be modulated by METTL21D-dependent lysine trimethylation. As with Hsp70s, VCP has also been shown to be extensively modified, mostly by phosphorylation and acetylation [56]–[60]. In this report, we demonstrate that methylation of the VCP requires a novel, specific methyltransferase, which in turn seems to be highly conserved throughout evolution. Indeed, tandem-affinity purification of a yeast homolog of METTL21D, Nnt1p, led to the identification of the yeast homolog of VCP, Cdc48p (Figure S7 and Table S2), hinting at the importance of this interaction.
Strickingly, methylation of VCP is further enhanced by direct interaction of the methyltransferase with ASPSCR1, a poorly characterized VCP cofactor, and this effect appears to require the C-terminal half of ASPSCR1. Mutations in the VCP gene have been linked to autosomal dominant disorders Inclusion Body Myopathy with Paget's disease of bone and Fronto-temporal Dementia (IBMPFD) and familial Amyotrophic Lateral Sclerosis (ALS) [47], [48]. Most VCP mutations reside within the N-terminal domain, which has been proposed to be involved in cofactor association [49]–[51].
Substitutions R155H, R159G and R191Q have no impact on the in vitro methylation of the protein. However, addition of ASPSCR1 no longer appears to increase the methylation of mutant VCP as compared to the wild-type protein. This observation opens up a whole new area of investigation in understanding the molecular physiopathology of IBMPFD and familial ALS. It may therefore be of interest to assess the relative methylation of VCP in affected patients as compared to healthy individuals. Many studies were performed to define how these disease mutations affect the function of VCP. From a biochemical point of view, the most promising alteration concerned the increased ATPase activity that may reflect structural changes induced by ATP binding [61], [62]. Methylation of VCP by METTL21D is shown here to significantly decrease activity of the first ATPase domain of this chaperone. This modification could eventually help modulate enzymatic activity of VCP that has gone haywire due to mutation.
Our work on the KIN protein, which eventually led to the discovery of its associated methyltransferase METTL22, began when it was detected in the interactome of a number of prefoldins (see supporting data in [32]). Thus, even though KIN is not known to have chaperone activity, it still appears to interact with chaperones and potentially affect their activity. The exact function of KIN is still a matter of debate. This DNA and RNA binding protein has been assigned a role in DNA repair and/or replication [63]–[66] and possibly mRNA processing as suggested by its identification in a number of proteomic analyses of the spliceosome [67], [68]. The role of the herein identified methylation will likely go hand in hand with the function of the winged helix domain in which it resides. Interestingly, yet another winged helix-containing protein was detected in the METTL22 purification, FOXK1. In this case, the function of the winged helix is known since it is required for DNA binding of this transcription factor. If METTL22 is shown to methylate FOXK1 as it did with KIN, this may in turn point to a more complex regulation of winged helix factors.
Advances in proteomics have helped to catalog various post-translational modifications of the proteome, and it now seems evident that chaperones contain several occurrences of such modifications. Recent identification of Hsp90 methylation by lysine methyltransferase SMYD2 is further evidence of the significance of this modification in regulating chaperone function [69]. Just like post-translational modifications of histone tails were shown to modulate binding to multiple chromatin remodeling, transcription, and mRNA processing factors, we believe that chaperone modifications may compose a similar code to help define specificity of discrete subsets to their seemingly innumerable effectors. Further decrypting this “chaperone code” is now required to understand how the functional organization of the proteome is orchestrated.
Materials and Methods
Generation of cell lines for expressing TAP-tagged polypeptides
Coding sequences for methyltransferases discussed in this article were obtained from the I.M.A.G.E. consortium clone library (Thermo Scientific). The corresponding cDNAs were cloned into the mammalian expression vector pMZI [70] carrying a TAP tag at its C-terminus [71], [72]. Stable human embryonic kidney cell lines (EcR-293; derived from HEK293) carrying these constructs were produced as described previously [28], [33].
Tandem affinity purification
Induction for 48 hours with 3 µM ponasterone A (Life Technologies) was used to express the TAP-tagged proteins. Whole cell extracts prepared from induced and non-induced stable EcR-293 cell lines were subjected to purification by the TAP procedure as described previously [28], [33].
Protein identification by mass spectrometry
TAP eluates were desalted and concentrated on Amicon Ultra 10K centrifugal filter units (Millipore) and resolved on NuPAGE 4–12% Bis-Tris Gel (Life Technologies). The gels were silver-stained, and the entirety of the tracks where the eluates had migrated were cut in about 20 slices that were subsequently digested with trypsin as described previously [28], [33]. The resulting tryptic peptides were purified and identified by LC-tandem mass spectrometry (MS/MS) using a microcapillary reversed-phase high pressure liquid chromatography coupled to a LTQ Orbitrap (ThermoElectron) mass spectrometer with a nanospray interface. The peak list files were generated with extract_msn.exe (version February 15, 2005) using the following parameters: minimum mass set to 600 Da, maximum mass set to 6000 Da, no grouping of MS/MS spectra, precursor charge set to auto, and minimum number of fragment ions set to 10. Protein database searching was performed with Mascot 2.3 (Matrix Science) against the human NCBInr protein database (version April 2, 2009). There are 10,427,007 sequences in this database. The mass tolerances for precursor and fragment ions were set to 10 ppm and 0.6 Da, respectively. Trypsin was used as the enzyme allowing for up to two missed cleavages. Carbamidomethyl, oxidation of methionine, mono-, di-, and trimethylation of lysine were allowed as variable modifications. In cases where multiple gene products were identified from the same peptide set, all were unambiguously removed from the data set. In the case of multiple isoforms stemming from a unique gene, the isoform with the best sequence coverage was reported. Proteins identified on the basis of a single peptide were also discarded.
Reliability assessment of protein-protein interactions was performed by our previously published software Decontaminator [73]. A set of 17 pairs of matched non-induced bait expression (control) and induced bait expression vectors was used for the algorithm training. Those were chosen on the basis of the absence of leaky expression of the tagged protein in the non-induced experiments. Decontaminator builds Bayesian probabilistic models of the Mascot scores [74] for each protein observed in the training set. It then assigns a p-value to each bait-prey interaction by computing the significance of the observed prey Mascot score compared to its corresponding control Mascot score model. A False Discovery Rate (FDR) is then calculated for each protein-protein interactions in the dataset using a leave-one-out scheme. All interactions with a FDR below 10% were reported as bait-specific, resulting in a dataset of 234 interactions. In other words, less than 24 interactions are expected to be the consequence of contamination.
In vitro methylation assay
The protocol was slightly modified from Inamitsu et al. [75]. Coding sequences of methyltransferases were cloned into pGEX-4-T1 vector (GE Healthcare). Coding sequences of putative methylation substrates were cloned into pET-23a(+) vector (EMD Chemicals). All vectors were transformed in One Shot BL21 Star (DE3) (Life Technologies), and protein synthesis was induced with IPTG. Bacteria were harvested by centrifugation, and pellets were lysed with the use of an IEC French Press (Thermo Scientific). The resulting GST - and 6xHis-fusion proteins were purified using Glutathione Sepharose 4B (GE Healthcare) and Ni-NTA Agarose (QIAGEN) beads, respectively, according to the manufacturers' specifications. In each reaction described in this article, 1 µg of GST-tagged methyltransferase was incubated with 2.5 µg His-tagged substrate and 5 µCi of S-[methyl-3H]-Adenosyl-L-methionine (81.7 Ci/mmol; PerkinElmer) in 50 µl of PBS for 90 min at 37°C. The samples were resolved on 10% acrylamide gels that were stained with Coomassie to show total amounts of proteins. Gels were then treated with EN3HANCE (PerkinElmer) according to the manufacturer's specifications. Tritium-based methylation signals were detected by autoradiography with four hours of exposure on Amersham Hyperfilm MP (GE Healthcare) at −80°C. Alternatively, an assay was produced with unlabeled S-adenosyl-L-methionine that was resolved on NuPAGE 4–12% Bis-Tris Gel (Life Technologies) and Coomassie stained. The bands corresponding to the His-tagged substrates were excised, trypsin digested, and analyzed as described above.
Immunofluorescence
HeLa S3 (CCL-2.2 ATCC) cells were grown on Lab-Tek II chamber slides (Nalge Nunc) and co-transfected with FLAG and GFP expressing vectors (p3XFLAG-CMV-14 and pGFP2-N1; Sigma-Aldrich & PerkinElmer Life Sciences, respectively) using Lipofectamine 2000 according to the manufacturer's specifications (Life Technologies). Twenty-four hours following transfection, cells were fixed with 3.7% formaldehyde in Phosphate-Buffered Saline (PBS) and permeabilized with 0.3% Triton X-100 PBS. Slides were then fixed in 5% donkey serum PBS for 1 hour, incubated with 1∶200 monoclonal FLAG antibody (M2; Sigma-Aldrich) in 5% donkey serum PBS for 90 min, and then incubated for an additional hour with 1∶50 Cy3-conjugated donkey anti-mouse IgG secondary antibody (Jackson ImmunoResearch) in 5% donkey serum PBS. Slides were washed three times with PBS for 5 min after each step. DNA was stained with TO-PRO-3 (Molecular Probes). Slides were mounted using ProLong Gold antifade reagent (Life Technologies). Images were acquired with an LSM 700 confocal laser scanning microscope at 63× magnification and analyzed using ZEN 2010 software (Zeiss, Toronto, Canada).
In vitro GST pull-down assay
The assay was based on the protocol described in Zwijsen et al. [76]. Briefly, 500 ng of GST-tagged METTL21D was incubated for 1 hour at 4°C with 100 ng of His-tagged VCP, ASPSCR1 or UBXN6, and 25 µl Glutathione Sepharose 4B beads (GE Healthcare) in 1 ml of binding buffer (50 mM NaCl, 50 mM HEPES-KOH pH 7.6, 0.1% NP-40, 0.5% charcoal-stripped FBS) complemented with complete EDTA-free protease inhibitor cocktail (Roche). The beads were washed three times by centrifugation with the same buffer and the bound proteins were eluted by boiling for 5 min in sample buffer, and separated on NuPAGE 4–12% Bis-Tris Gel (Life Technologies). Binding of His-tagged proteins was detected by Western blot analysis using mouse monoclonal anti-6X His tag-antibody (abcam). A second blot was made with rabbit polyclonal anti-GST antibody (abcam) to ensure that comparable amounts of GST-tagged baits were purified by the pull-down.
ATPase assay
PiColorLock Gold Phosphate Detection System (Innova Biosciences) was used to quantify ATPase activity in vitro. Three micrograms of a His-tagged fragment corresponding to the first 481 residues of VCP were incubated beforehand with 2 µg of GST-METTL21D and 0.5 mM S-adenosyl-L-methionine for 30 min at 37°C in 100 µl of 0.1 M Tris pH 7.5. All assays were performed in triplicate. Independent experiments were carried out where the VCP fragment was replaced by an unmethylatable mutant (K315R); the methyltransferase was replaced by a catalytically inactive mutant (E73Q); or the methyl donor, S-adenosyl-L-methionine, was replaced by a methylation inhibitor, S-adenosyl-L-homocysteine. The rest of the protocol followed the guidelines provided by the manufacturer.
Phylogenetic analysis
All available ortholog sequences of PRMTs and of the family of 10 putative methyltransferases in the UniProt database [77] were aligned using the ClustalW2 multiple sequence alignment software (version 2.0.12) [78]. The most conserved region of the alignment was then selected to build an unrooted phylogenetic tree through the Jalview software [79] using the neighbor-joining algorithm [80] with the BLOSUM62 substitution matrix [81]. Orthologs forming monophyletic groups with their respective human sequences were collapsed to a single node in the phylogenetic tree shown in Figure 1A.
Supporting Information
Zdroje
1. MartinJL, McMillanFM (2002) SAM (dependent) I AM: the S-adenosylmethionine-dependent methyltransferase fold. Curr Opin Struct Biol 12 : 783–793.
2. PetrossianTC, ClarkeSG (2011) Uncovering the human methyltransferasome. Mol Cell Proteomics 10: M110 000976.
3. SchubertHL, BlumenthalRM, ChengX (2003) Many paths to methyltransfer: a chronicle of convergence. Trends Biochem Sci 28 : 329–335.
4. IshizawaT, NozakiY, UedaT, TakeuchiN (2008) The human mitochondrial translation release factor HMRF1L is methylated in the GGQ motif by the methyltransferase HMPrmC. Biochem Biophys Res Commun 373 : 99–103.
5. KlotzAV, LearyJA, GlazerAN (1986) Post-translational methylation of asparaginyl residues. Identification of beta-71 gamma-N-methylasparagine in allophycocyanin. J Biol Chem 261 : 15891–15894.
6. WebbKJ, Zurita-LopezCI, Al-HadidQ, LaganowskyA, YoungBD, et al. (2010) A novel 3-methylhistidine modification of yeast ribosomal protein Rpl3 is dependent upon the YIL110W methyltransferase. J Biol Chem 285 : 37598–37606.
7. PeggAE (1990) Mammalian O6-alkylguanine-DNA alkyltransferase: regulation and importance in response to alkylating carcinogenic and therapeutic agents. Cancer Res 50 : 6119–6129.
8. SprungR, ChenY, ZhangK, ChengD, ZhangT, et al. (2008) Identification and validation of eukaryotic aspartate and glutamate methylation in proteins. J Proteome Res 7 : 1001–1006.
9. ClarkeS (1992) Protein isoprenylation and methylation at carboxyl-terminal cysteine residues. Annu Rev Biochem 61 : 355–386.
10. TooleyCE, PetkowskiJJ, Muratore-SchroederTL, BalsbaughJL, ShabanowitzJ, et al. (2010) NRMT is an alpha-N-methyltransferase that methylates RCC1 and retinoblastoma protein. Nature 466 : 1125–1128.
11. WuJ, TolstykhT, LeeJ, BoydK, StockJB, et al. (2000) Carboxyl methylation of the phosphoprotein phosphatase 2A catalytic subunit promotes its functional association with regulatory subunits in vivo. Embo J 19 : 5672–5681.
12. FengQ, WangH, NgHH, Erdjument-BromageH, TempstP, et al. (2002) Methylation of H3-lysine 79 is mediated by a new family of HMTases without a SET domain. Curr Biol 12 : 1052–1058.
13. MartinC, ZhangY (2005) The diverse functions of histone lysine methylation. Nat Rev Mol Cell Biol 6 : 838–849.
14. VolkelP, AngrandPO (2007) The control of histone lysine methylation in epigenetic regulation. Biochimie 89 : 1–20.
15. BarskiA, CuddapahS, CuiK, RohTY, SchonesDE, et al. (2007) High-resolution profiling of histone methylations in the human genome. Cell 129 : 823–837.
16. KochCM, AndrewsRM, FlicekP, DillonSC, KaraozU, et al. (2007) The landscape of histone modifications across 1% of the human genome in five human cell lines. Genome Res 17 : 691–707.
17. RosenfeldJA, WangZ, SchonesDE, ZhaoK, DeSalleR, et al. (2009) Determination of enriched histone modifications in non-genic portions of the human genome. BMC Genomics 10 : 143.
18. ChuikovS, KurashJK, WilsonJR, XiaoB, JustinN, et al. (2004) Regulation of p53 activity through lysine methylation. Nature 432 : 353–360.
19. HoutzRL, StultsJT, MulliganRM, TolbertNE (1989) Post-translational modifications in the large subunit of ribulose bisphosphate carboxylase/oxygenase. Proc Natl Acad Sci U S A 86 : 1855–1859.
20. HuangJ, Perez-BurgosL, PlacekBJ, SenguptaR, RichterM, et al. (2006) Repression of p53 activity by Smyd2-mediated methylation. Nature 444 : 629–632.
21. KruiswijkT, KunstA, PlantaRJ, MagerWH (1978) Modification of yeast ribosomal proteins. Methylation. Biochem J 175 : 221–225.
22. LeeSW, BergerSJ, MartinovicS, Pasa-TolicL, AndersonGA, et al. (2002) Direct mass spectrometric analysis of intact proteins of the yeast large ribosomal subunit using capillary LC/FTICR. Proc Natl Acad Sci U S A 99 : 5942–5947.
23. LevyD, KuoAJ, ChangY, SchaeferU, KitsonC, et al. (2011) Lysine methylation of the NF-kappaB subunit RelA by SETD6 couples activity of the histone methyltransferase GLP at chromatin to tonic repression of NF-kappaB signaling. Nat Immunol 12 : 29–36.
24. LhoestJ, LobetY, CostersE, ColsonC (1984) Methylated proteins and amino acids in the ribosomes of Saccharomyces cerevisiae. Eur J Biochem 141 : 585–590.
25. MorinoH, KawamotoT, MiyakeM, KakimotoY (1987) Purification and properties of calmodulin-lysine N-methyltransferase from rat brain cytosol. J Neurochem 48 : 1201–1208.
26. PolevodaB, MartzenMR, DasB, PhizickyEM, ShermanF (2000) Cytochrome c methyltransferase, Ctm1p, of yeast. J Biol Chem 275 : 20508–20513.
27. YangXD, HuangB, LiM, LambA, KelleherNL, et al. (2009) Negative regulation of NF-kappaB action by Set9-mediated lysine methylation of the RelA subunit. Embo J 28 : 1055–1066.
28. CloutierP, Al-KhouryR, Lavallee-AdamM, FaubertD, JiangH, et al. (2009) High-resolution mapping of the protein interaction network for the human transcription machinery and affinity purification of RNA polymerase II-associated complexes. Methods 48 : 381–386.
29. CloutierP, CoulombeB (2010) New insights into the biogenesis of nuclear RNA polymerases? Biochem Cell Biol 88 : 211–221.
30. CojocaruM, BouchardA, CloutierP, CooperJJ, VarzavandK, et al. (2011) Transcription factor IIS cooperates with the E3 ligase UBR5 to ubiquitinate the CDK9 subunit of the positive transcription elongation factor B. J Biol Chem 286 : 5012–5022.
31. CoulombeB, BlanchetteM, JeronimoC (2008) Steps towards a repertoire of comprehensive maps of human protein interaction networks: the Human Proteotheque Initiative (HuPI). Biochem Cell Biol 86 : 149–156.
32. ForgetD, LacombeAA, CloutierP, Al-KhouryR, BouchardA, et al. (2010) The protein interaction network of the human transcription machinery reveals a role for the conserved GTPase RPAP4/GPN1 and microtubule assembly in nuclear import and biogenesis of RNA polymerase II. Mol Cell Proteomics 9 : 2827–2839.
33. JeronimoC, ForgetD, BouchardA, LiQ, ChuaG, et al. (2007) Systematic analysis of the protein interaction network for the human transcription machinery reveals the identity of the 7SK capping enzyme. Mol Cell 27 : 262–274.
34. JeronimoC, LangelierMF, ZeghoufM, CojocaruM, BergeronD, et al. (2004) RPAP1, a novel human RNA polymerase II-associated protein affinity purified with recombinant wild-type and mutated polymerase subunits. Mol Cell Biol 24 : 7043–7058.
35. KruegerBJ, JeronimoC, RoyBB, BouchardA, BarrandonC, et al. (2008) LARP7 is a stable component of the 7SK snRNP while P-TEFb, HEXIM1 and hnRNP A1 are reversibly associated. Nucleic Acids Res 36 : 2219–2229.
36. BaysNW, WilhovskySK, GoradiaA, Hodgkiss-HarlowK, HamptonRY (2001) HRD4/NPL4 is required for the proteasomal processing of ubiquitinated ER proteins. Mol Biol Cell 12 : 4114–4128.
37. BraunS, MatuschewskiK, RapeM, ThomsS, JentschS (2002) Role of the ubiquitin-selective CDC48(UFD1/NPL4)chaperone (segregase) in ERAD of OLE1 and other substrates. Embo J 21 : 615–621.
38. JaroschE, TaxisC, VolkweinC, BordalloJ, FinleyD, et al. (2002) Protein dislocation from the ER requires polyubiquitination and the AAA-ATPase Cdc48. Nat Cell Biol 4 : 134–139.
39. RabinovichE, KeremA, FrohlichKU, DiamantN, Bar-NunS (2002) AAA-ATPase p97/Cdc48p, a cytosolic chaperone required for endoplasmic reticulum-associated protein degradation. Mol Cell Biol 22 : 626–634.
40. YeY, MeyerHH, RapoportTA (2001) The AAA ATPase Cdc48/p97 and its partners transport proteins from the ER into the cytosol. Nature 414 : 652–656.
41. IoukTL, AitchisonJD, MaguireS, WozniakRW (2001) Rrb1p, a yeast nuclear WD-repeat protein involved in the regulation of ribosome biosynthesis. Mol Cell Biol 21 : 1260–1271.
42. MagnaniR, DirkLM, TrievelRC, HoutzRL (2010) Calmodulin methyltransferase is an evolutionarily conserved enzyme that trimethylates Lys-115 in calmodulin. Nat Commun 1 : 43.
43. ZhangK, YauPM, ChandrasekharB, NewR, KondratR, et al. (2004) Differentiation between peptides containing acetylated or tri-methylated lysines by mass spectrometry: an application for determining lysine 9 acetylation and methylation of histone H3. Proteomics 4 : 1–10.
44. MeisterG, EggertC, BuhlerD, BrahmsH, KambachC, et al. (2001) Methylation of Sm proteins by a complex containing PRMT5 and the putative U snRNP assembly factor pICln. Curr Biol 11 : 1990–1994.
45. AlexandruG, GraumannJ, SmithGT, KolawaNJ, FangR, et al. (2008) UBXD7 binds multiple ubiquitin ligases and implicates p97 in HIF1alpha turnover. Cell 134 : 804–816.
46. OrmeCM, BoganJS (2012) The ubiquitin regulatory X (UBX) domain-containing protein TUG regulates the p97 ATPase and resides at the endoplasmic reticulum-golgi intermediate compartment. J Biol Chem 287 : 6679–6692.
47. JohnsonJO, MandrioliJ, BenatarM, AbramzonY, Van DeerlinVM, et al. (2010) Exome sequencing reveals VCP mutations as a cause of familial ALS. Neuron 68 : 857–864.
48. WattsGD, WymerJ, KovachMJ, MehtaSG, MummS, et al. (2004) Inclusion body myopathy associated with Paget disease of bone and frontotemporal dementia is caused by mutant valosin-containing protein. Nat Genet 36 : 377–381.
49. HanzelmannP, SchindelinH (2011) The structural and functional basis of the p97/valosin-containing protein (VCP)-interacting motif (VIM): mutually exclusive binding of cofactors to the N-terminal domain of p97. J Biol Chem 286 : 38679–38690.
50. SchuberthC, BuchbergerA (2008) UBX domain proteins: major regulators of the AAA ATPase Cdc48/p97. Cell Mol Life Sci 65 : 2360–2371.
51. YeungHO, KloppsteckP, NiwaH, IsaacsonRL, MatthewsS, et al. (2008) Insights into adaptor binding to the AAA protein p97. Biochem Soc Trans 36 : 62–67.
52. WangQ, SongC, YangX, LiCC (2003) D1 ring is stable and nucleotide-independent, whereas D2 ring undergoes major conformational changes during the ATPase cycle of p97-VCP. J Biol Chem 278 : 32784–32793.
53. DalalS, RosserMF, CyrDM, HansonPI (2004) Distinct roles for the AAA ATPases NSF and p97 in the secretory pathway. Mol Biol Cell 15 : 637–648.
54. WangC, LazaridesE (1984) Arsenite-induced changes in methylation of the 70,000 dalton heat shock proteins in chicken embryo fibroblasts. Biochem Biophys Res Commun 119 : 735–743.
55. WangC, LinJM, LazaridesE (1992) Methylations of 70,000-Da heat shock proteins in 3T3 cells: alterations by arsenite treatment, by different stages of growth and by virus transformation. Arch Biochem Biophys 297 : 169–175.
56. KleinJB, BaratiMT, WuR, GozalD, SachlebenLRJr, et al. (2005) Akt-mediated valosin-containing protein 97 phosphorylation regulates its association with ubiquitinated proteins. J Biol Chem 280 : 31870–31881.
57. KoikeM, FukushiJ, IchinoheY, HigashimaeN, FujishiroM, et al. (2010) Valosin-containing protein (VCP) in novel feedback machinery between abnormal protein accumulation and transcriptional suppression. J Biol Chem 285 : 21736–21749.
58. LiG, ZhaoG, SchindelinH, LennarzWJ (2008) Tyrosine phosphorylation of ATPase p97 regulates its activity during ERAD. Biochem Biophys Res Commun 375 : 247–251.
59. LivingstoneM, RuanH, WeinerJ, ClauserKR, StrackP, et al. (2005) Valosin-containing protein phosphorylation at Ser784 in response to DNA damage. Cancer Res 65 : 7533–7540.
60. Mori-KonyaC, KatoN, MaedaR, YasudaK, HigashimaeN, et al. (2009) p97/valosin-containing protein (VCP) is highly modulated by phosphorylation and acetylation. Genes Cells 14 : 483–497.
61. HalawaniD, LeBlancAC, RouillerI, MichnickSW, ServantMJ, et al. (2009) Hereditary inclusion body myopathy-linked p97/VCP mutations in the NH2 domain and the D1 ring modulate p97/VCP ATPase activity and D2 ring conformation. Mol Cell Biol 29 : 4484–4494.
62. MannoA, NoguchiM, FukushiJ, MotohashiY, KakizukaA (2010) Enhanced ATPase activities as a primary defect of mutant valosin-containing proteins that cause inclusion body myopathy associated with Paget disease of bone and frontotemporal dementia. Genes Cells 15 : 911–922.
63. AnguloJF, MoreauPL, MaunouryR, LaporteJ, HillAM, et al. (1989) KIN, a mammalian nuclear protein immunologically related to E. coli RecA protein. Mutat Res 217 : 123–134.
64. BiardDS, MiccoliL, DesprasE, HarperF, PichardE, et al. (2003) Participation of kin17 protein in replication factories and in other DNA transactions mediated by high molecular weight nuclear complexes. Mol Cancer Res 1 : 519–531.
65. KannoucheP, MauffreyP, Pinon-LatailladeG, MatteiMG, SarasinA, et al. (2000) Molecular cloning and characterization of the human KIN17 cDNA encoding a component of the UVC response that is conserved among metazoans. Carcinogenesis 21 : 1701–1710.
66. MiccoliL, FrouinI, NovacO, Di PaolaD, HarperF, et al. (2005) The human stress-activated protein kin17 belongs to the multiprotein DNA replication complex and associates in vivo with mammalian replication origins. Mol Cell Biol 25 : 3814–3830.
67. MakarovEM, MakarovaOV, UrlaubH, GentzelM, WillCL, et al. (2002) Small nuclear ribonucleoprotein remodeling during catalytic activation of the spliceosome. Science 298 : 2205–2208.
68. RappsilberJ, RyderU, LamondAI, MannM (2002) Large-scale proteomic analysis of the human spliceosome. Genome Res 12 : 1231–1245.
69. DonlinLT, AndresenC, JustS, RudenskyE, PappasCT, et al. (2012) Smyd2 controls cytoplasmic lysine methylation of Hsp90 and myofilament organization. Genes Dev 26 : 114–119.
70. ZeghoufM, LiJ, ButlandG, BorkowskaA, CanadienV, et al. (2004) Sequential Peptide Affinity (SPA) system for the identification of mammalian and bacterial protein complexes. J Proteome Res 3 : 463–468.
71. PuigO, CasparyF, RigautG, RutzB, BouveretE, et al. (2001) The tandem affinity purification (TAP) method: a general procedure of protein complex purification. Methods 24 : 218–229.
72. RigautG, ShevchenkoA, RutzB, WilmM, MannM, et al. (1999) A generic protein purification method for protein complex characterization and proteome exploration. Nat Biotechnol 17 : 1030–1032.
73. Lavallee-AdamM, CloutierP, CoulombeB, BlanchetteM (2010) Modeling contaminants in AP-MS/MS experiments. J Proteome Res 10 : 886–895.
74. PerkinsDN, PappinDJ, CreasyDM, CottrellJS (1999) Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20 : 3551–3567.
75. InamitsuM, ItohS, HellmanU, Ten DijkeP, KatoM (2006) Methylation of Smad6 by protein arginine N-methyltransferase 1. FEBS Lett 580 : 6603–6611.
76. ZwijsenRM, BuckleRS, HijmansEM, LoomansCJ, BernardsR (1998) Ligand-independent recruitment of steroid receptor coactivators to estrogen receptor by cyclin D1. Genes Dev 12 : 3488–3498.
77. ApweilerR, Jesus MartinM, O'onovanC, MagraneM, Alam-FaruqueY, et al. (2012) Reorganizing the protein space at the Universal Protein Resource (UniProt). Nucleic Acids Res 40: D71–75.
78. LarkinMA, BlackshieldsG, BrownNP, ChennaR, McGettiganPA, et al. (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23 : 2947–2948.
79. WaterhouseAM, ProcterJB, MartinDM, ClampM, BartonGJ (2009) Jalview Version 2–a multiple sequence alignment editor and analysis workbench. Bioinformatics 25 : 1189–1191.
80. SaitouN, NeiM (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4 : 406–425.
81. HenikoffS, HenikoffJG (1992) Amino acid substitution matrices from protein blocks. Proc Natl Acad Sci U S A 89 : 10915–10919.
Štítky
Genetika Reprodukční medicína
Článek Comparative Genome Structure, Secondary Metabolite, and Effector Coding Capacity across PathogensČlánek TATES: Efficient Multivariate Genotype-Phenotype Analysis for Genome-Wide Association StudiesČlánek Secondary Metabolism and Development Is Mediated by LlmF Control of VeA Subcellular Localization inČlánek Human Disease-Associated Genetic Variation Impacts Large Intergenic Non-Coding RNA ExpressionČlánek The Roles of Whole-Genome and Small-Scale Duplications in the Functional Specialization of GenesČlánek The Role of Autophagy in Genome Stability through Suppression of Abnormal Mitosis under Starvation
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2013 Číslo 1
-
Všechny články tohoto čísla
- A Model of High Sugar Diet-Induced Cardiomyopathy
- Comparative Genome Structure, Secondary Metabolite, and Effector Coding Capacity across Pathogens
- Emerging Function of Fat Mass and Obesity-Associated Protein (Fto)
- Positional Cloning Reveals Strain-Dependent Expression of to Alter Susceptibility to Bleomycin-Induced Pulmonary Fibrosis in Mice
- Genetics of Ribosomal Proteins: “Curiouser and Curiouser”
- Transposable Elements Re-Wire and Fine-Tune the Transcriptome
- Function and Regulation of , a Gene Implicated in Autism and Human Evolution
- MAML1 Enhances the Transcriptional Activity of Runx2 and Plays a Role in Bone Development
- Predicting Mendelian Disease-Causing Non-Synonymous Single Nucleotide Variants in Exome Sequencing Studies
- A Systematic Mapping Approach of 16q12.2/ and BMI in More Than 20,000 African Americans Narrows in on the Underlying Functional Variation: Results from the Population Architecture using Genomics and Epidemiology (PAGE) Study
- Transcription of the Major microRNA–Like Small RNAs Relies on RNA Polymerase III
- Histone H3K56 Acetylation, Rad52, and Non-DNA Repair Factors Control Double-Strand Break Repair Choice with the Sister Chromatid
- Genome-Wide Association Study Identifies a Novel Susceptibility Locus at 12q23.1 for Lung Squamous Cell Carcinoma in Han Chinese
- Genetic Disruption of the Copulatory Plug in Mice Leads to Severely Reduced Fertility
- The [] Prion Exists as a Dynamic Cloud of Variants
- Adult Onset Global Loss of the Gene Alters Body Composition and Metabolism in the Mouse
- Fis Protein Insulates the Gene from Uncontrolled Transcription
- The Meiotic Nuclear Lamina Regulates Chromosome Dynamics and Promotes Efficient Homologous Recombination in the Mouse
- Genome-Wide Haplotype Analysis of Expression Quantitative Trait Loci in Monocytes
- TATES: Efficient Multivariate Genotype-Phenotype Analysis for Genome-Wide Association Studies
- Structural Basis of a Histone H3 Lysine 4 Demethylase Required for Stem Elongation in Rice
- The Ecm11-Gmc2 Complex Promotes Synaptonemal Complex Formation through Assembly of Transverse Filaments in Budding Yeast
- MCM8 Is Required for a Pathway of Meiotic Double-Strand Break Repair Independent of DMC1 in
- Comparative Genomic Analysis of the Endosymbionts of Herbivorous Insects Reveals Eco-Environmental Adaptations: Biotechnology Applications
- Integration of Nodal and BMP Signals in the Heart Requires FoxH1 to Create Left–Right Differences in Cell Migration Rates That Direct Cardiac Asymmetry
- Pharmacodynamics, Population Dynamics, and the Evolution of Persistence in
- A Hybrid Likelihood Model for Sequence-Based Disease Association Studies
- Aberration in DNA Methylation in B-Cell Lymphomas Has a Complex Origin and Increases with Disease Severity
- Multiple Opposing Constraints Govern Chromosome Interactions during Meiosis
- Transcriptional Dynamics Elicited by a Short Pulse of Notch Activation Involves Feed-Forward Regulation by Genes
- Dynamic Large-Scale Chromosomal Rearrangements Fuel Rapid Adaptation in Yeast Populations
- Heterologous Gln/Asn-Rich Proteins Impede the Propagation of Yeast Prions by Altering Chaperone Availability
- Gene Copy-Number Polymorphism Caused by Retrotransposition in Humans
- An Incompatibility between a Mitochondrial tRNA and Its Nuclear-Encoded tRNA Synthetase Compromises Development and Fitness in
- Secondary Metabolism and Development Is Mediated by LlmF Control of VeA Subcellular Localization in
- Single-Stranded Annealing Induced by Re-Initiation of Replication Origins Provides a Novel and Efficient Mechanism for Generating Copy Number Expansion via Non-Allelic Homologous Recombination
- Tbx2 Controls Lung Growth by Direct Repression of the Cell Cycle Inhibitor Genes and
- Suv4-20h Histone Methyltransferases Promote Neuroectodermal Differentiation by Silencing the Pluripotency-Associated Oct-25 Gene
- A Conserved Helicase Processivity Factor Is Needed for Conjugation and Replication of an Integrative and Conjugative Element
- Telomerase-Null Survivor Screening Identifies Novel Telomere Recombination Regulators
- Genome-Wide Analysis Reveals Selection for Important Traits in Domestic Horse Breeds
- Coordinated Degradation of Replisome Components Ensures Genome Stability upon Replication Stress in the Absence of the Replication Fork Protection Complex
- Nkx6.1 Controls a Gene Regulatory Network Required for Establishing and Maintaining Pancreatic Beta Cell Identity
- HIF- and Non-HIF-Regulated Hypoxic Responses Require the Estrogen-Related Receptor in
- Delineating a Conserved Genetic Cassette Promoting Outgrowth of Body Appendages
- The Telomere Capping Complex CST Has an Unusual Stoichiometry, Makes Multipartite Interaction with G-Tails, and Unfolds Higher-Order G-Tail Structures
- Comprehensive Methylome Characterization of and at Single-Base Resolution
- Loci Associated with -Glycosylation of Human Immunoglobulin G Show Pleiotropy with Autoimmune Diseases and Haematological Cancers
- Switchgrass Genomic Diversity, Ploidy, and Evolution: Novel Insights from a Network-Based SNP Discovery Protocol
- Centromere-Like Regions in the Budding Yeast Genome
- Sequencing of Loci from the Elephant Shark Reveals a Family of Genes in Vertebrate Genomes, Forged by Ancient Duplications and Divergences
- Mendelian and Non-Mendelian Regulation of Gene Expression in Maize
- Mutational Spectrum Drives the Rise of Mutator Bacteria
- Human Disease-Associated Genetic Variation Impacts Large Intergenic Non-Coding RNA Expression
- The Roles of Whole-Genome and Small-Scale Duplications in the Functional Specialization of Genes
- Sex-Specific Signaling in the Blood–Brain Barrier Is Required for Male Courtship in
- A Newly Uncovered Group of Distantly Related Lysine Methyltransferases Preferentially Interact with Molecular Chaperones to Regulate Their Activity
- Is Required for Leptin-Mediated Depolarization of POMC Neurons in the Hypothalamic Arcuate Nucleus in Mice
- Unlocking the Bottleneck in Forward Genetics Using Whole-Genome Sequencing and Identity by Descent to Isolate Causative Mutations
- The Role of Autophagy in Genome Stability through Suppression of Abnormal Mitosis under Starvation
- MTERF3 Regulates Mitochondrial Ribosome Biogenesis in Invertebrates and Mammals
- Downregulation and Altered Splicing by in a Mouse Model of Facioscapulohumeral Muscular Dystrophy (FSHD)
- NBR1-Mediated Selective Autophagy Targets Insoluble Ubiquitinated Protein Aggregates in Plant Stress Responses
- Retroactive Maintains Cuticle Integrity by Promoting the Trafficking of Knickkopf into the Procuticle of
- Phenome-Wide Association Study (PheWAS) for Detection of Pleiotropy within the Population Architecture using Genomics and Epidemiology (PAGE) Network
- Genetic and Functional Modularity of Activities in the Specification of Limb-Innervating Motor Neurons
- A Population Genetic Model for the Maintenance of R2 Retrotransposons in rRNA Gene Loci
- A Quartet of PIF bHLH Factors Provides a Transcriptionally Centered Signaling Hub That Regulates Seedling Morphogenesis through Differential Expression-Patterning of Shared Target Genes in
- A Genome-Wide Integrative Genomic Study Localizes Genetic Factors Influencing Antibodies against Epstein-Barr Virus Nuclear Antigen 1 (EBNA-1)
- Mutation of the Diamond-Blackfan Anemia Gene in Mouse Results in Morphological and Neuroanatomical Phenotypes
- Life, the Universe, and Everything: An Interview with David Haussler
- Alternative Oxidase Expression in the Mouse Enables Bypassing Cytochrome Oxidase Blockade and Limits Mitochondrial ROS Overproduction
- An Evolutionarily Conserved Synthetic Lethal Interaction Network Identifies FEN1 as a Broad-Spectrum Target for Anticancer Therapeutic Development
- The Flowering Repressor Underlies a Novel QTL Interacting with the Genetic Background
- Telomerase Is Required for Zebrafish Lifespan
- and Diversified Expression of the Gene Family Bolster the Floral Stem Cell Network
- Susceptibility Loci Associated with Specific and Shared Subtypes of Lymphoid Malignancies
- An Insertion in 5′ Flanking Region of Causes Blue Eggshell in the Chicken
- Increased Maternal Genome Dosage Bypasses the Requirement of the FIS Polycomb Repressive Complex 2 in Arabidopsis Seed Development
- WNK1/HSN2 Mutation in Human Peripheral Neuropathy Deregulates Expression and Posterior Lateral Line Development in Zebrafish ()
- Synergistic Interaction of Rnf8 and p53 in the Protection against Genomic Instability and Tumorigenesis
- Dot1-Dependent Histone H3K79 Methylation Promotes Activation of the Mek1 Meiotic Checkpoint Effector Kinase by Regulating the Hop1 Adaptor
- A Heterogeneous Mixture of F-Series Prostaglandins Promotes Sperm Guidance in the Reproductive Tract
- Starvation, Together with the SOS Response, Mediates High Biofilm-Specific Tolerance to the Fluoroquinolone Ofloxacin
- Directed Evolution of a Model Primordial Enzyme Provides Insights into the Development of the Genetic Code
- Genome-Wide Screens for Tinman Binding Sites Identify Cardiac Enhancers with Diverse Functional Architectures
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Function and Regulation of , a Gene Implicated in Autism and Human Evolution
- An Insertion in 5′ Flanking Region of Causes Blue Eggshell in the Chicken
- Comprehensive Methylome Characterization of and at Single-Base Resolution
- Susceptibility Loci Associated with Specific and Shared Subtypes of Lymphoid Malignancies
Přihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



