-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaA Simple Auxin Transcriptional Response System Regulates Multiple Morphogenetic Processes in the Liverwort
In flowering plants it is hypothesized the complexity and the robustness of the auxin transcriptional response could be generated by a large number of paralogs encoding components of this signaling network. Yet, it is not known whether alternative embryophyte body plans can be patterned with a simpler auxin transcriptional machinery. We demonstrate that in the liverwort Marchantia polymorpha, the type rather than the number of components of the auxin transcriptional response pathway are sufficient to pattern a complex three-dimensional gametophyte with multiple tissue and cell types. In M. polymorpha, mutations in components of the auxin signaling pathway cause dramatic pleiotropic effects despite having a single class A activating AUXIN RESPONSE FACTOR (MpARF1). This supports the hypothesis that auxin is context dependent and that it facilitates rather than specifies particular developmental processes. We show that auxin signaling is not necessary for survival but it is required to pattern the transition from two-dimensional to three-dimensional growth. This suggests that the evolution of the auxin transcriptional response was critical for the evolution of developmental complexity in land plants.
Published in the journal: . PLoS Genet 11(5): e32767. doi:10.1371/journal.pgen.1005207
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1005207Summary
In flowering plants it is hypothesized the complexity and the robustness of the auxin transcriptional response could be generated by a large number of paralogs encoding components of this signaling network. Yet, it is not known whether alternative embryophyte body plans can be patterned with a simpler auxin transcriptional machinery. We demonstrate that in the liverwort Marchantia polymorpha, the type rather than the number of components of the auxin transcriptional response pathway are sufficient to pattern a complex three-dimensional gametophyte with multiple tissue and cell types. In M. polymorpha, mutations in components of the auxin signaling pathway cause dramatic pleiotropic effects despite having a single class A activating AUXIN RESPONSE FACTOR (MpARF1). This supports the hypothesis that auxin is context dependent and that it facilitates rather than specifies particular developmental processes. We show that auxin signaling is not necessary for survival but it is required to pattern the transition from two-dimensional to three-dimensional growth. This suggests that the evolution of the auxin transcriptional response was critical for the evolution of developmental complexity in land plants.
Introduction
Genetic and physiological studies have revealed that the phytohormone auxin, indole-3-acetic acid (IAA), controls central aspects of plant development in both haploid and diploid generations of plant species. Auxin triggers growth and tissue differentiation in places where it accumulates, perhaps acting as a morphogenetic trigger [1]. In the flowering plant Arabidopsis thaliana, a balance between auxin synthesis, transport, conjugation/de-conjugation and degradation regulates the spatial distribution of auxin, leading to activation of transcriptional responses that influence a variety of morphological and physiological processes [2,3,4,5,6,7,8].
The auxin transcriptional response in its simplest outline is as follows. In the absence of auxin, AUXIN/INDOLE-3-ACETIC ACID (AUX/IAA) repressors heterodimerize with AUXIN RESPONSE FACTOR (ARF) transcription factors (via two conserved domains, D34) and recruit TOPLESS (TPL) corepressors, thus preventing ARFs from regulating target genes [9,10,11,12,13]. In the presence of auxin, the AUX/IAA proteins are targeted for degradation by an SCF E3 ubiquitin ligase (TRANSPORT INHIBITOR RESPONSE1 [TIR1]/AUXIN SIGNALING F-BOX [AFB]), via auxin dependent interactions between AUX/IAA and TIR1 proteins [14,15,16,17]. Phylogenetic analyses of ARF evolution in land plants identifies three ARF classes: class A (orthologues of Arabidopsis ARF5-8), class B (orthologues of Arabidopsis ARF1-4, ARF9) and class C (orthologues of Arabidopsis ARF10, ARF16, ARF17) [18,19,20]. However, not all classes of ARFs are equal, with class A acting primarily as activators, and classes B and C as repressors [19,20]. Since auxin application predominantly activates gene expression, it has been argued that class B and C ARFs may act to repress common target genes activated by class A ARFs since all classes can potentially bind the same auxin responsive element (AuxRE) in promoters of auxin responsive genes [19,20,21]. Among the direct targets of ARFs are genes encoding AUX/IAA proteins, establishing a negative feedback loop [22,23]. Studies in Arabidopsis have suggested that activator ARFs interact with AUX/IAAs and that AUX/IAAs may also homodimerize, but that repressor ARFs do not interact efficiently with the AUX/IAAs, leaving most class B and C repressors outside this protein network [10]. These results imply that many ARF repressors regulate gene expression independently of auxin, as TIR1/AFB mediated degradation of AUX/IAAs would not affect their activity. However, all ARF proteins potentially compete for the same targets as they recognize similar AuxREs [11,21]. Thus, a current view is that transcriptional activation by class A ARFs is negatively regulated in an auxin dependent manner by AUX/IAAs and an auxin independent manner by class B and C ARFs.
That 23 ARF, 29 AUX/IAA, 6 TIR1/AFB, and 5 TPL paralogs are encoded in the Arabidopsis genome has prompted speculation that interactions differing in affinity among paralogs could influence auxin sensitivity in different cells [10,24]. Key features of the auxin signaling pathway are mechanistically similar in the moss Physcomitrella patens suggesting conservation of the auxin transcriptional response throughout land plants [25]. Given the complexity arising from multiple paralogs encoding each of the components of the auxin transcriptional response pathway in both flowering plants and P. patens [18], we sought to establish a model system where such complexity is reduced. It has been speculated that an increase in complexity of the auxin genetic toolkit (including genes for metabolism, synthesis, transport and signaling) is correlated with an increased complexity in land plant architecture, from bryophytes to flowering plants [26]. Although there is no universal agreement on the phylogenetic relationships of extant land plant clades, it is generally accepted that liverworts diverged earlier than other land plants based on morphology, fossil evidence, and some molecular analyses [27,28,29]. Liverworts in general have experienced a low rate of chromosomal evolution, with little evidence of widespread polyploidy, and molecular evolution within the Marchantiopsida is reported to be slow, compared with more derived liverwort lineages [30,31]. Given these evolutionary attributes, we reasoned that Marchantia polymorpha could possess a simplified version of the auxin response system. Furthermore, as M. polymorpha has been used as a model system for nearly two centuries to investigate both universal and specific biological processes, an extraordinary body of literature is available to inform our studies.
In M. polymorpha, the haploid spore produces a short protonemal filament only a few cells long [32,33,34]. The brief protonemal stage is followed by the formation of a globular sporeling that subsequently develops a marginal row of cells in its apical (distal) region, a central cell of which will form the apical cell of the gametophytic shoot meristem [33,34,35]. The marginal row of cells is conspicuous and forms the distal growing tip, and represents the transition from a sporeling to a 'prothallus'—a flattened unistratose heart shaped plant that grows by means of an apical cell with two alternate cutting faces [33,35]. Subsequently, a change in the wedge-shaped apical cell such that divisions occur along four planes, rather than two, transforms the prothallus into a thallus. Accompanying this transition is the establishment of dorsiventrality, with the production of specialized cell types both dorsally and ventrally [36]. M. polymorpha can also reproduce asexually via small, flattened lecticular gemmae [32,33,37,38]. Mature gemmae are dormant and lack dorsiventral polarity while inside dorsal gemmae cups, but acquire environmentally induced polarity when displaced from the gemmae cup. Individual gemmae are clonal and derived from a single cell of the parental thallus [32,33,37,39].
The effects of exogenous auxin on M. polymorpha gemmaling growth have been reported in previous studies. Some generalizations of the effects of exogenous auxin in young gemmalings can be made as follows: in low concentrations auxin increases the size of the thallus [40], breaks gemmae dormancy [41,42], produces formation of additional meristems in dormant gemmae [43], and confers apical dominance [44]. At higher concentrations auxin promotes the formation of numerous rhizoids on the dorsal side of the gemmae, a dorsiventral patterning defect, and further increases in concentration results in formation of a callus-like tissue with limited growth [40,41,42,43,44,45,46,47,48,49,50,51,52,53,54]. In transgenic M. polymorpha, an auxin-responsive regulatory sequence derived from soybean, the GRETCHEN HAGEN 3 (GH3) promoter, proGmGH3, responds to exogenous auxin in a dose dependent manner [40], and consistent with previous pharmacological experiments, sites of proGmGH3 expression in un-induced lines partially overlap with sites of auxin action, including incipient rhizoids and the bases of gemmae cups.
In the present work, we show that the liverwort M. polymorpha possesses the simplest auxin genetic toolkit so far found in any land plant. Dissection of the auxin response system of M. polymorpha, points to auxin being a facilitator rather than a determiner of cell fate, with auxin also being required for major developmental transitions during gametophyte development.
Results
Auxin influences most developmental stages of M. polymorpha gametophyte development
To address the role of auxin in the M. polymorpha gametophyte, young undifferentiated gemmae and young thalli with established dorsiventral polarity were grown in the presence of the auxin analogues naphthalene-1-acetic acid (NAA) and 2,4-dichlorophenoxyactetic acid (2,4-D).
Auxin application to gemmae and gemmalings grown up to four days old on unsupplemented media prior to auxin treatment triggered ectopic rhizoid formation on the dorsal surface of the thallus (Fig 1A and 1B). Normal dorsal structures including air chambers, air pores, and gemmae cups did not develop. However, plants were not entirely ventralized since scales, which are ventral structures, did not develop dorsally. Thalli also failed to develop a normal branching pattern and lacked lateral growth.
Fig. 1. Developmental effects of auxin in the M. polymorpha thallus. 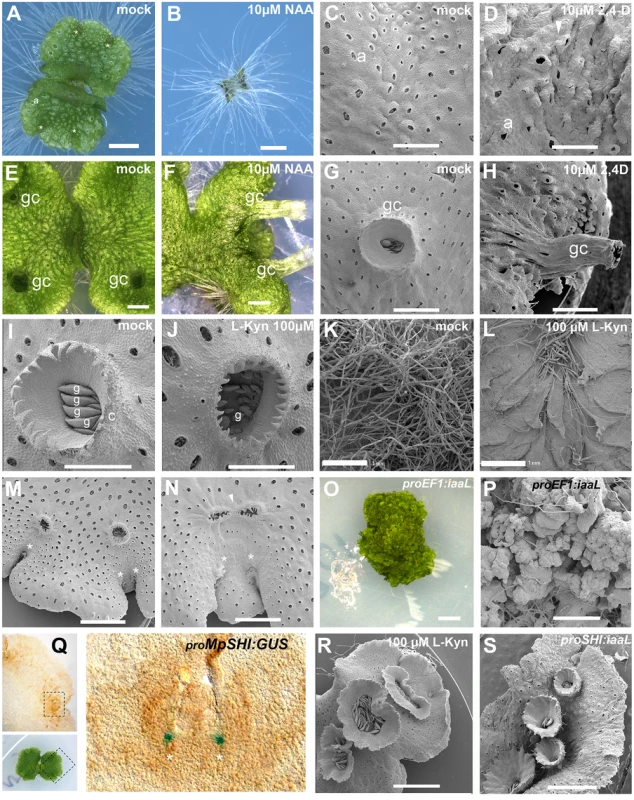
(A) 10-day-old wild-type gemmaling with four apical notches and well developed air pores. (B) 10-day-old wild-type gemmae grown on 10 μM NAA. (C) Dorsal epidermis of 15-day-old wild-type thallus. (D) Dorsal epidermis of 15-day-old wild-type thallus, transferred on day 6 to 10 μM 2,4-D with abnormal air pores. (E) 23-day-old wild-type thallus with three gemmae cups. (F) 23-day-old wild-type thallus transferred at day 7 to 10 μM NAA with elongated gemmae cup. (G) Wild-type mature gemmae cup. (H) Wild-type mature gemmae cup from 17-day-old plant, transferred at day 12 to 10 μM 2,4-D. (I) Wild-type mature gemmae cup with gemmae inside. (J) Wild-type mature gemmae cup from a plant grown on 100 μM L-Kyn. (K) Ventral side of wild-type thallus grown on mock media. (L) Ventral side of wild-type thallus grown on 100 μM L-Kyn; a decrease in rhizoid number exposes ventral scales. (M) Mature thallus showing a thallus lobe separating two recently bifurcated apical notches. (N) Mature thallus of plants transferred on day 7 to 100 μM L-Kyn, showing a smaller lobe spacing apical notches and fused gemmae cup primordia (arrowhead). (O) Strong proEF1:iaaL lines are composed of an undifferentiated mass of cells that fail to establish a clear dorsiventrality. (P) Close up view of proEF1:iaaL line shows a mass of undifferentiated cells although rhizoids are able to differentiate. (Q) Expression pattern of proMpSHI in mature thallus with staining in the vicinity of the apical cell. (R) Different degrees of gemmae cup fusion observed in 100 μM L-Kyn treated plants. (S) proMpSHI:iaaL plants show an irregular zig-zag arrangement of gemmae cups that are in close proximity to each other. Scale bars in A, B, E, F, G, H, J, K, L, O, R and S = 1 mm; C, D and P = 0.6 mm; I = 0.5 mm; M = 1.2 mm; N = 0.75 mm; Q = 0.2 mm. Asterisk, apical notch; a, air pores; gc, gemma cups; g, gemmae. Application of auxin to gemmalings with established dorsiventral polarity, those grown for six and sixteen days on unsupplemented media prior to auxin treatment, resulted in two classes of aberrant phenotype. In the first class, occurring primarily in plants with auxin treatment after one week, the thallus developed excessive rhizoids, although they were spatially restricted to the original central zones of the gemmalings and to the ventral side of the mature thallus (S1A and S1B Fig)—locations where rhizoids arise in untreated plants. Thus, by day six of gemmaling development dorsal tissues have lost the capacity to produce rhizoids in response to auxin (S1B Fig). However, typical dorsal structures of the thallus were affected, with air pores often either partially formed or unopened (Fig 1C and 1D, S1B and S1C Fig), air chambers structure asymmetric and disorganized, and an undulating dorsal epidermal surface with conspicuously protruding air pores (Fig 1D). Additionally, plants subject to this treatment were unable to form any gemmae cups (S1A–S1C Fig).
In the second, milder class, primarily occurring in plants with auxin treatment after two weeks, plants exhibited abnormal tubular elongation of gemmae cups that had been initiated prior to auxin treatment (Fig 1E–1H and S1D Fig). The gemmae cups exhibited a positive gravitropic response in contrast to their normal negative graviotropism [55]. While previously formed gemmae cups continued to develop, no new cups were formed. Air chambers were asymmetric, spatially disorganized, and protruding upwards as in the first class (Fig 1F–1H). The growing thallus margins were bent downwards (Fig 1F and S1D Fig), an epinastic growth pattern reminiscent of epinastic angiosperm leaf growth in response to high levels of auxin [56].
Lack of auxin affects gemmae cup formation and thallus branching
As a complementary approach we examined the phenotypic consequences of reducing endogenous auxin levels. We first tested the effects of the auxin biosynthesis inhibitor L-Kynurenine (L-Kyn) on gemmaling development [57]. In Arabidopsis L-Kyn acts as an alternative substrate and competitive inhibitor for the TRYPTOPHAN AMINOTRANSFERASE OF ARABIDOPSIS1 (TAA1) and TRYPTOPHAN AMINOTRANSFERASE RELATED1, 2 (TAR1, 2) enzymes, inhibiting the conversion of tryptophan (trp) to indole-3-pyruvic acid (IPA), a key step in auxin biosynthesis [4,57,58]. Arabidopsis plants grown in the presence of 100 μM L-Kyn phenocopy wei8 tar2 double mutants (the weak ethylene insensitive 8 mutant is a null loss-of-function allele of TAA1) [57,59].
Wild-type gemmae grown on 100 to 250 μM L-Kyn were unable to produce normal gemmae cups, with gemmae cup rims failing to fully develop (Fig 1I and 1J). Growth of gemmae was slower as compared to wild type (S1E and S1F Fig), with smaller propagules forming inside the cup (Fig 1I and 1J). Decreased rhizoid production was also observed but was not measured quantitatively (Fig 1K and 1L).
During vegetative growth, a gemmae cup is usually initiated following each branching event [36] and only a single cup is formed per branching event (S2 Fig). Immediately after branching a growth phase separates the two apical notches creating a mid-lobe prior to a new round of branching and gemmae cup formation. Application of L-Kyn to established seven-day-old thalli resulted in gemmae cups derived from successive branching events fusing into a single structure (Fig 1M and 1N). The extent of fusion varied, with the most extreme fusion resulting in single oval gemmae cups (e.g. Fig 1N and 1R). Therefore, inhibiting auxin synthesis reduces the rate of branching and the extent of growth separating branches.
In order to further deplete free auxin levels in the thallus, we created transgenic plants overexpressing a heterologous bacterial auxin conjugation enzyme iaaL [60,61,62] under the control of the EF1-α promoter (proEF1), which drives high levels of expression in the meristematic regions and throughout the thallus [63]. proEF1:iaaL plants exhibited disruptions to patterning and growth, with thallus growth severely stunted, despite evidence of continuing cell division (Fig 1O and 1P). Except for a few cells developing as rhizoids, most cells in proEF1:iaaL plants do not acquire a particular fate, but instead resemble cells present in early stages of sporeling development.
To manipulate free auxin levels specifically in the meristematic region we utilized a 3.2kb promoter fragment of the single M. polymorpha orthologue of the SHORT INTERNODES/STYLISH (SHI/STY) gene family [64,65,66]. Under normal growth conditions, proMpSHI:GUS activity was detected mainly in the apical notch (meristematic region) during all stages of development (Fig 1Q). Most proMpSHI:iaaL transformants exhibited weaker phenotypes than proEF1:iaaL and were able to form a narrow thallus with compromised lamina expansion and a reduced rate of branching (Fig 1S, S1G and S1H Fig). Gemmae cups sometimes developed to the side of the midline, and/or close together, suggesting an irregular zig-zag formation and a degree of fusion (Fig 1S), resembling L-Kyn treatments (Fig 1R). We thus conclude that auxin influences the growth and differentiation of multiple tissues and processes in different developmental stages of the M. polymorpha gametophyte.
Auxin transcriptional response genes are conserved in M. polymorpha
Using PCR based methods and genomic sequences provided by the Joint Genome Institute, we identified single TPL (MpTPL, genbank accession number KP877967), AUX/IAA (MpIAA, KP877968), and TIR1/AFB (MpTIR) orthologues in M. polymorpha, as well as three ARF paralogs (MpARF1, KP877969; MpARF2, KP877970; MpARF3, KP877971; S13 Fig).
Analysis of the MpTPL sequence revealed highly conserved LisH, CTLH and WD40 repeat domains, characteristic of this gene family (amino acid sequence identity between MpTPL and Arabidopsis TPL is 68%). MpIAA harbors a conserved EAR domain (D1), as well as a proteolysis domain (D2); D1 allows AUX/IAA proteins to interact with the TPL CTLH domain, and D2 domain has been shown to facilitate interactions between AUX/IAA proteins and TIR1/AFB [15]. Both MpIAA and all three MpARFs possess conserved protein-protein interaction domains (D34; S3 Fig and S1 Table). In an accompanying paper, Kato et al [67] describe the extent of MpARF-MpIAA interactions in M. polymorpha using yeast two-hybrid (Y2H) and biomolecular fluorescence complementation (BiFC).
Bayesian phylogenetic analyses were performed to elucidate the evolution of the TPL, AUX/IAA, and ARF gene families using sequences from M. polymorpha, P. patens, Selaginella moellendorffii, Pseudotsuga menziesii, Amborella tricopoda, Solanum lycopersicum and Arabidopsis thaliana. The TPL phylogeny suggests that the common ancestor of land plants had a single TPL gene, with the M. polymorpha genome retaining this ancestral feature. Our analysis suggests a gene duplication in the lineage leading to the origin of tracheophytes (maximum support), creating two classes of TPL genes (Class A and B). This topology implies a loss of class A genes in S. moellendorffii (S4 Fig) and further sequence sampling may clarify if this is a trend in other lycophytes.
Phylogenetic analysis of the AUX/IAA family also suggests a single gene in the embryophyte ancestor, rendering M. polymorpha ancestral for AUX/IAA gene copy number. Our phylogenetic tree indicates that angiosperms and gymnosperms also have two types of AUX/IAAs, class A (similar to the Arabidopsis IAA1-7, IAA9, IAA14-17, IAA19 and IAA27) and class B AUX/IAAs (including IAA10-13, IAA18, IAA26, IAA28 and IAA29), suggesting a gene duplication in the common ancestor of seed plants (S5 Fig).
We performed Bayesian analysis of land plant ARF genes, and as noted in previous analyses, our phylogenetic analysis results in a tree with three distinct and well-supported ARF clades (A, B, C) (S6 Fig). M. polymorpha possesses a single homolog of each of these classes, MpARF1 (A), MpARF2 (B), and MpARF3 (C).
Loss of MpIAA results in auxin hypersensitivity
To test whether the components identified by homology act in an auxin signaling pathway, we created loss - and gain-of-function alleles of the genes. Two independent artificial microRNAs (amiRs) were designed to target MpIAA (amiRMpIAA7 and amiRMpIAA9). For amiR design in M. polymorpha, we used the backbone of the endogenous MpMIR160 precursor (S7 Fig), replacing the miR and miR* sequences with sequences to target the gene of interest, and cleavage was corroborated using RLM-RACE (Fig 2A; [68]). Both amiRs were constitutively expressed under the EF1-α promoter (proEF1) and the reduction of target MpIAA transcripts was corroborated (S8 Fig). While most proEF1:amiRMpIAA7 and proEF1:amiRMpIAA9 lines had nearly wild-type phenotypes, some lines exhibited a curly and convoluted thallus with long rhizoids, and miRMpIAA transcript levels are largely, albeit not perfectly, correlated with the severity of the phenotype (S8 Fig). Consistent with MpIAA having a role in auxin signaling, a majority of amiRMpIAA7/9 lines showed severe growth defects in exogenous auxin compared to wild-type controls (Fig 2). When grown in 2.5 and 5μM 2,4-D auxin media from the sporeling stage, proEF1:amiRMpIAA7/9 plants were dramatically smaller compared to wild-type controls, suggesting that exogenous auxin causes lethality when transcript levels of MpIAA are reduced (Fig 2B and S8D Fig). When mature proEF1:amiRMpIAA9 thalli were grown in mock media for 15 days and then transferred to 7.5μM 2,4-D plates for 20 days, loss of MpIAA resulted in profuse rhizoid production and a decrease in growth compared to controls (Fig 2B). In summary, knock-down MpIAA lines were auxin hypersensitive, indicating that MpIAA represses the auxin signaling pathway in M. polymorpha.
Fig. 2. MpIAA represses auxin signaling in M. polymorpha. 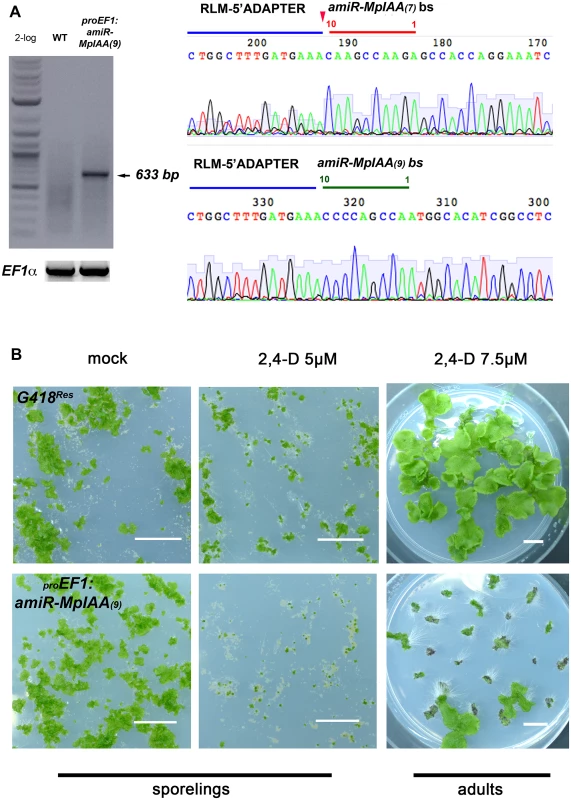
(A) RLM RACE of amiRMpIAA expressing lines. Left panel shows gel electrophoresis of RLM-RACE products. Right panel shows direct sequencing of the 633bp RLM-RACE product from amiRMpIAA9. Sequencing detected the expected cleavage of MpIAA between bases 10/11 of both miR sequences. (B) Auxin hypersensitivity in knock-down MpIAA lines at both the sporeling and adult stages. At the sporeling stage, proEF1:amiRMpIAA9 sporelings grown on 5μM 2,4-D from germination have severely affected area compared to controls. In the adult stage plants were grown for 16 days on B5 media prior to treatment with 2,4-D. proEF1:amiRMpIAA9 treated with 7.5μM 2,4-D for ten days form ectopic rhizoids compared to controls. In all panels, each of the plants is an independent transformant. Scale Bars = 1 cm. MpARF1 is a modulator of the auxin signaling pathway
To elucidate whether auxin hypersensitivity observed in MpIAA knockdown lines was due to misregulation of ARF genes, we explored the functional role of MpARF1. We focused on MpARF1 since MpARF1 transcripts are readily detectable in the thallus, whereas we did not detect transcripts of MpARF2 in the thallus (S8B Fig). Evidence that MpARF1 elicits a transcriptional output of the auxin signaling pathway comes from mutant alleles obtained by CRISPR-CAS (Sugano et al 2014) and artificial miRNAs [69], where plants deficient in MpARF1 activity are unable to form ectopic rhizoids in the presence of exogenous auxin. Our phylogenetic analysis places MpARF1 as a member of class A ARF activators, loss-of-function alleles of which cause auxin insensitivity in Arabidopsis. Thus, gain-of-function MpARF1 alleles should recapitulate aspects of loss-of-function MpIAA alleles. We cloned a wild-type allele of MpARF1 as well as a truncated dominant gain-of-function mutant of MpARF1 lacking D34 (MpARF1ΔD34; S9 Fig) that should produce a protein with limited interaction with MpIAA (Krogan et al 2012). When MpARF1 and MpARF1ΔD34 genes were driven by proEF1-α, plants displayed auxin hypersensitivity, exemplified by formation of ectopic rhizoids in adults exposed to 5μM 2,4-D (Fig 3A). Even in the absence of auxin, proEF1:MpARF1ΔD34 plants exhibited excessive rhizoid production with rhizoids growing to greater lengths compared to a HygRes control, visible when plants are grown on vertically oriented plates such that gravity influenced rhizoid growth is evident (Fig 3B). proEF1:MpARF1 and proEF1:MpARF1ΔD34 plants exhibited higher rates of branching, seen as a significant increase in apical notches compared to controls (Fig 3A and 3C; S3 Table), and proEF1:MpARF1ΔD34 lines had increased gemma cup production compared to proEF1:MpARF1 lines (Fig 3C). The extent of phenotypic aberration was correlated with MpARF1 transgene expression (Fig 3D). Additionally, the levels of MpIAA expression appear to be inversely correlated with levels of MpARF1 expression, supporting the idea that MpARF1 inhibits transcription of MpIAA
Fig. 3. Gain-of-function MpARF1 lines are auxin hypersensitive.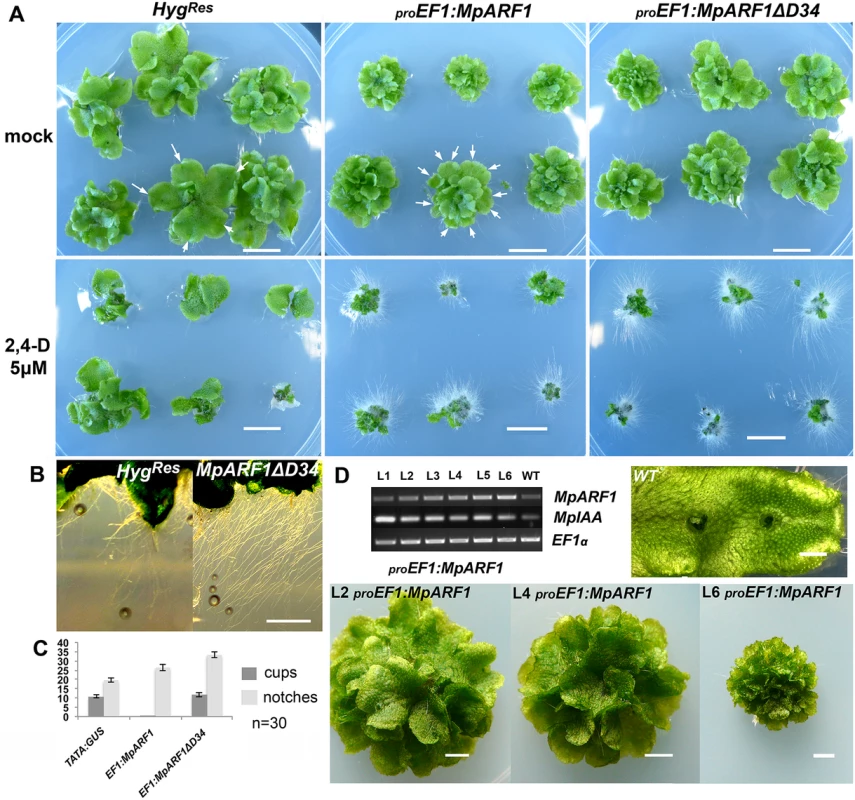 
(A) Each panel show six independent lines, transformed with constructs as indicated above the panels. Plants are treated as indicated to the left. Arrows indicate apical notches. (B) Left panel: Rhizoids from control HygR lines. Right panel: proEF1:MpARF1ΔD34 lines develop elongated and profuse rhizoids. (C) Apical notch and gemmae cup measurements for control, proEF1:MpARF1 and proEF1:MpARF1ΔD34 primary transformants after 50 days of growth (30 independent lines were scored per genotype). MpARF1 overexpression results in increased branching rates. Deletion of D34MpARF1 recovers the gemmae cup production lost by over-expressing full length MpARF1 transcripts. (D) Semi-quantitative RT-PCR showing six independent proEF1:MpARF1 lines exhibiting an inverse correlation between MpARF1 and MpIAA transcript levels. Three of the six lines are shown in the panel, with line L6 having more extensive branching and a smaller size compared to line L2. The constitutive translation ELONGATION FACTOR 1-alpha (EF1) was used as a control. All scale bars = 1 cm, except D = 1 mm. MpTPL is involved in patterning multiple tissues in M. polymorpha development
The lack of strong phenotypic effects in knock-down MpIAA lines prompted us to examine the single member of the TOPLESS corepressor family in M. polymorpha. Expression patterns of a proMpTPL:3xVENUS fusion, including 3.6kb upstream of MpTPL transcript up to the nearest upstream transcript, exhibited intense fluorescent signal in the meristematic regions during most developmental stages, including mature gemmae (Fig 4A), five-day-old gemmalings (Fig 4B), and mature 25-day-old thalli (Fig 4C) compared to autofluroesence in the wild type (S10A Fig). Expression was also detected in gemmae cups (Fig 4D). Transfer of mature proMpTPL:3xVENUS plants to media containing 10 μM 2,4-D resulted in a decreased level of expression of the reporter gene, suggesting regulatory feedback between auxin and MpTPL (Fig 4E). Expression of proMpTPL:3xVENUS was observed throughout the thallus, although the signal was weaker compared to proEF1:3xVENUS lines (Fig 4F).
Fig. 4. Developmental role of MpTPL in M. polymorpha. 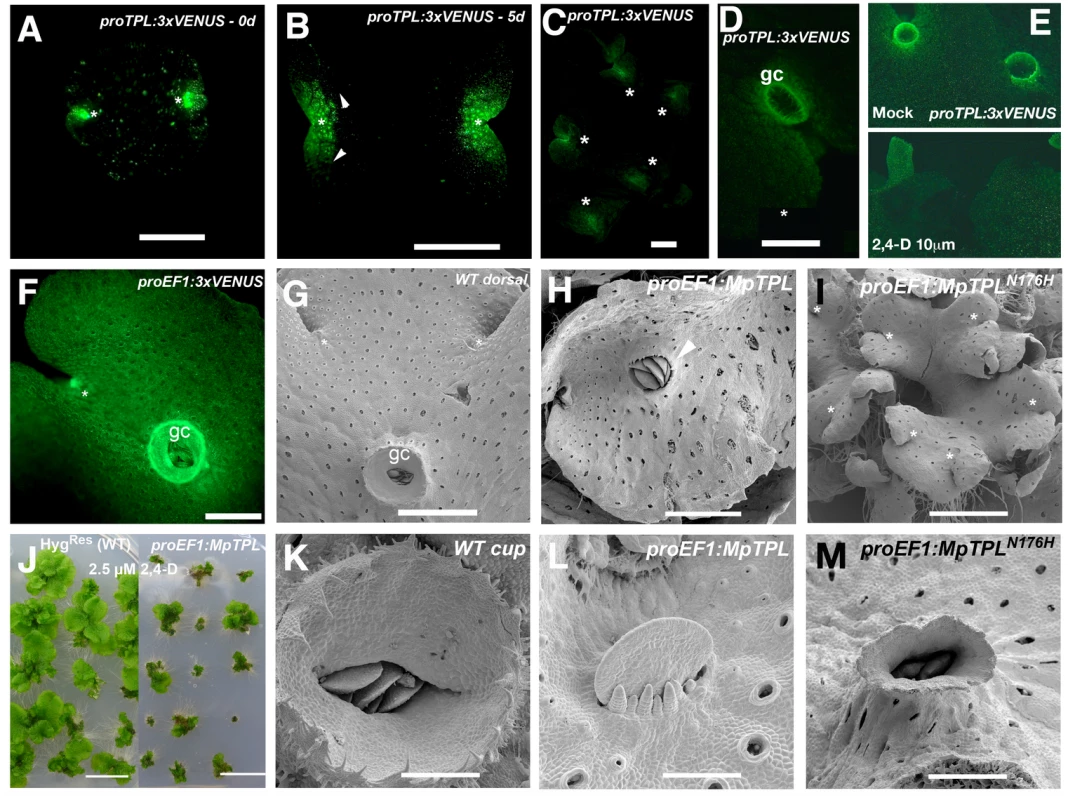
(A) Mature dormant gemma of proMpTPL:3xVENUSN7 with expression in two apical notches (asterisks). (B) 5-day-old gemmae of proMpTPL:3xVENUSN7. Expression has expanded from apical notches (asterisk) to zones of newly formed air chambers (arrowheads). Inset illustrates background fluorescence in the wild type with identical settings. (C) Mature 25-day-old proMpTPL:3xVENUSN7 line showing expression in meristematic regions (asterisks). (D) Young gemmae cup (gc) primordia in mature proMpTPL:3xVENUSN7 line. (E) Expression pattern of mature proMpTPL:3xVENUSN7 thallus on mock and 10μM 2,4-D media. Plants were transferred to exogenous auxin after 16 days on B5 media. (F) proEF1:VENUS indicating expression in apical notches (asterisk) and gemmae cups (gc). (G) Dorsal side of wild-type thallus covered with photosynthetic air chambers and two apical notches from which gemmae cups (gc) develop. (H) proEF1:MpTPL plants show dorsiventrality but are affected in cup development (arrowhead). (I) Representative proEF1:MpTPLN176H line with a compact and multi-bifurcated thalli, with at least 7 apical notches (asterisks) produced in a similar area compared to (G). (J) proEF1:MpTPL plants have diminished area in low auxin concentrations compared to controls. Each plant is an independent primary transformant grown for a month on 2.5μM 2,4-D. (K) Wild-type mature gemmae cup harboring dormant gemmae. (L) proEF1:MpTPL plants have no gemmae cups, instead naked gemmae and gemmae primordia with filaments growing by transversal cell divisions can be seen. (M) proEF1:MpTPLN176H gemmae cups are elongated and narrow, similar to developmental effects produced by exogenous auxin application in the wild type. Scale bars in A and B = 0.5mm; C, D, E, F, H, M = 1mm; G and I = 1.5mm; J = 1 cm; K = 300μm; L = 75μm; M = 0.6mm. The wild-type dorsiventral M. polymorpha thallus is several cell layers thick. The single apical cell in the shoot mersitem has four cleavage planes: one dorsal, one ventral, and two lateral [70,71]. Dorsal and lateral derivatives contribute to the formation of dorsal photosynthetic air chambers and gemmae cups (Fig 4G). Notably both dorsal gemmae cups and ventral rhizoids are produced along a midline (Fig 4F and 4G, and S2 Fig). Wild-type gemmae cups produce a characteristic cup rim that is several cell layers thick at its base and unistratose towards its apex and acts to store and protect dormant gemma (Fig 4K).
Ectopic expression of wild-type MpTPL using proEF1-α resulted in slightly smaller and darker green thalli able to form a bifurcating thallus with normal dorsiventral polarity (Fig 4H and S10B Fig). However, gemmae cup formation was disrupted in 80% of proEF1:MpTPL lines (N = 15), with gemmae cups entirely absent in some cases (Fig 4H and 4L). The loss of gemmae cups did not impair the formation of gemmae, as naked gemma and gemma primordia developed in the positions where gemmae cups would normally develop along the midline (Fig 4K and 4L), reminiscent of pharmacological effects of L-Kyn on the thallus. Intriguingly, proEF1:MpTPL lines were hypersensitive to 2.5 μM exogenous 2,4-D, similar to knock-down MpIAA lines (Fig 4J).
We created a putative MpTPL loss-of-function allele analogous to the dominant negative Arabidopsis TPL(N176H) and drove its expression with proEF1 [9]. The most conspicuous phenotype of proEF1:MpTPLN176H plants was an increased frequency of thallus branching compared to wild type and proEF1:MpTPL plants (Fig 4I; S2 Table). In this regard proEF1:MpTPLN176H plants resemble MpARF1 overexpression lines (Fig 4M). Thalli of proEF1:MpTPLN176H plants produced nearly three apical notches in the time in which wild-type plants produce one apical notch (Fig 4G and 4I). proEF1:MpTPLN176H plants had diminished production of gemmae cups, and when produced, gemmae cups were aberrant, being narrow and elongate, reminiscent of gemmae cups that develop in the presence of exogenous auxin (Fig 4M). When reproductive development was induced under far-red light proEF1:MpTPLN176H gametophore stalks showed decreased elongation compared to wild type, resulting in gametophores of short stature and with dorsal protrusions reminiscent of rhizoids (S10C Fig).
Auxin insensitive M. polymorpha plants fail to establish a dorsiventral thallus
In Arabidopsis translational fusions of TPL and D34 of AUX/IAA proteins, such as IAA12/BODENLOS (BDL; TPL-D34BDL), constitutively bind to ARF5/MONOPTEROS (MP) via interactions with D34, keeping MP in a repressive complex even in the presence of auxin [9]. Given the putative role of MpIAA and MpTPL proteins in auxin signaling in M. polymorpha, chimeric translational fusions of MpTPL to D34 of all M. polymorpha loci bearing putative D34 of protein-protein interaction were generated and all fusions were regulated by proEF1, making sure non of the fusions carried an endogenous miR binding site (S11 Fig; [40]).
proEF1:MpTPL-D34MpIAA plants exhibited severe dwarfism (Fig 5A) and failed to establish a dorsiventral thallus. Individuals were composed of a mass of undifferentiated cells and incompletely developed organs with random orientations (Fig 5B). Occasional rhizoids, scales and air chambers were observed, but these structures were not organized relative to one another as in wild-type plants (Fig 5B). Similar phenotypes were observed for proEF1:MpTPL-D34MpARF1 (Fig 5C and 5D), proEF1:MpTPL-D34MpARF2 (Fig 5E). A majority of these lines only achieved the level of differentiation of the sporeling or prothallus stages of wild-type development (Fig 5F), with individuals not transitioning further than single cell layer tissues (Fig 5D). Fusions of TPL with D34 of BDL (proEF1:MpTPL-D34BDL) resulted in similar phenotypes, suggesting a high degree of functional conservation between liverwort and angiosperm D34IAA protein domains (Fig 5G). Weaker proEF1:MpTPL-D34BDL lines had a narrow thallus with fused gemmae cups, as in L-Kyn treatments and in lines expressing the IAA conjugating enzyme iaaL (S12A and S12B Fig).
Fig. 5. Developmental effects of auxin insensitivity. 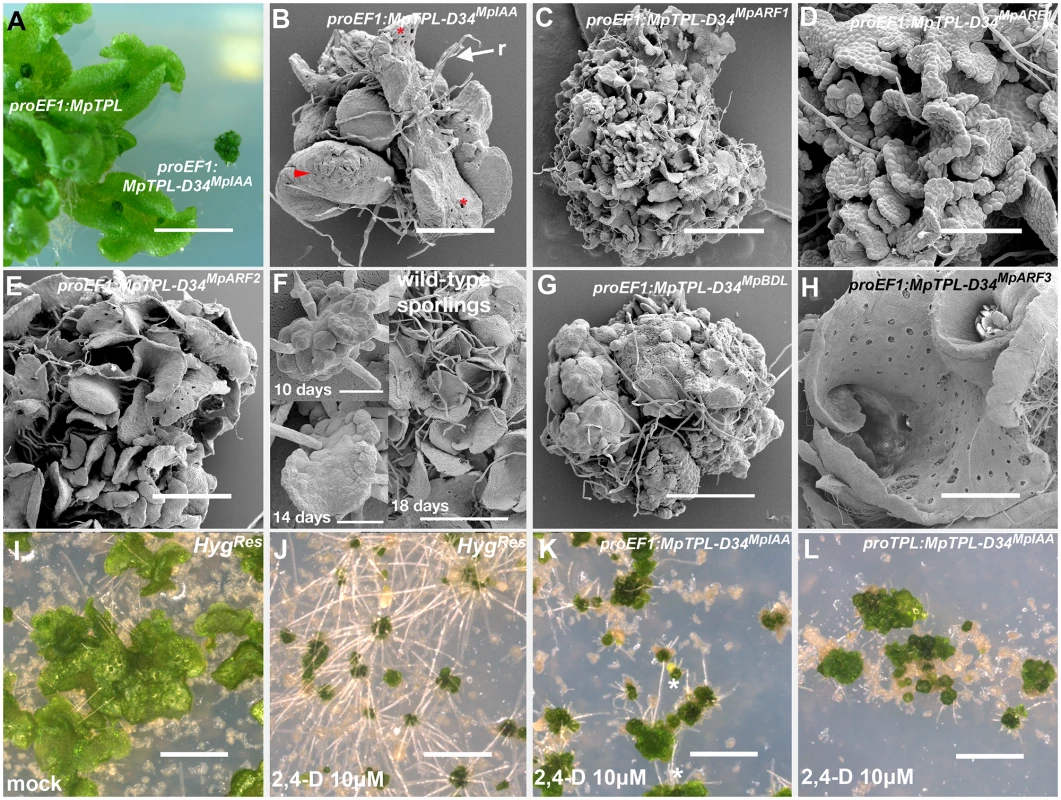
(A) proEF1:MpTPL-D34IAA lines suffer from severe dwarfism compared to proEF1:MpTPL plants. (B) Strong proEF1:MpTPL-D34IAA line showing a lack of organized patterning, dorsiventral polarity and a limited degree of differentiation. Scales (arrowhead), air pores (asterisk) and rhizoids (r) are visible in random places and growth is severely affected. (C) proEF1:MpTPL-D34MpARF1 representative line showing a degree of differentiation similar to the prothallus stage of wild-type development. (D) proEF1:MpTPL-D34MpARF1 showing single cell layer prothalli-like tisssue. (E) proEF1:MpTPL-D34MpARF2 representative line. (F) Wild-type sporeling (10 days) transitioning into a single cell layered prothallus (14 days) and fully established multiple prothalli 18 days after plating. (G) Strong proEF1:MpTPL-D34BDL line lacking growth, differentiation and dorsiventrality, but still capable of producing rhizoids. (H) proEF1:MpTPL-D34MpARF3 lines differentiate gemmae cups and a dorsiventral polarity with dorsal air chambers and ventral rhizoids. The thallus has an apparent hyponastic curvature. (I, J) Multiple independent HygRes sporelings (transformed with an empty binary vector) grown on media containing 10μg/ml Hyg and mock (I) or 10μM 2,4-D (J) for 14 days. (K, L) Multiple independent proEF1:MpTPL-D34MpIAA (K) or proMpTPL:MpTPL-D34MpIAA (L) lines grown on 10μM 2,4-D for 14 days. Scale bars in C, E, I, J, K and L = 1mm; A = 1cm; B = 430μm; D = 130μm; F = 30μm; G = 600μm; H = 860μm. In contrast to the severe developmental defects described above, proEF1:MpTPL-D34MpARF3 plants (Fig 5H) did not exhibit altered patterning in a manner comparable to other chimeric genes tested. proEF1:MpTPL-D34MpARF3 thalli had a convoluted dorsal surface (Fig 5H and S12C Fig), with thallus margins curled inwardly towards the midline, especially close to the apical notches, resulting in a more compact thallus than that of wild type (Fig 5H). proEF1:MpTPL-D34MpARF3 thalli produced ventral rhizoids and dorsal air chambers and gemmae cups. However, proEF1:MpTPL-D34MpARF3 plants grew bigger than other TPL-D34 fusions (S12C Fig).
As MpTPL acts as a repressor of auxin transcriptional response, we drove expression of the chimeric proteins with the MpTPL regulatory sequences. proMpTPL:MpTPL-D34MpIAA lines had a similar, but slightly milder, phenotype than proEF1:MpTPL-D34MpIAA lines: plants failed to pattern a flattened dorsiventral thallus and also formed a cluster of organs similar to wild-type prothalli (Fig 5F). Some proMpTPL:MpTPL-D34MpIAA lines however were able to form a single gemma cup able to produce gemmae (S12D and S12E Fig). The gemmae from these cups had distorted shapes with marginal extensions, and lacked the two apical notches conspicuous on wild-type gemmae (S12E Fig).
One prediction of constitutively expressing MpTPL-D34 chimeric proteins is a downregulation of activator ARF activity even if MpIAA proteins are degraded due to the presence of auxin, i.e. the transgenic plants should be auxin insensitive. When grown in the presence of exogenous auxin proEF1:MpTPL-D34MpIAA plants occasionally produce ectopic rhizoids compared to controls (Fig 5I–5K), while expression of MpTPL-D34MpIAA using proMpTPL produced auxin insensitive plants as assayed by the lack of ectopic rhizoid formation (Fig 5L). These results indicate that expression of the chimeric MpTPL-D34MpIAA protein is capable of creating auxin insensitive plants if it expressed in the appropriate cells at appropriate levels.
Discussion
A minimal auxin signaling network is sufficient to pattern a complex body plan
Phylogenetic analyses indicate the M. polymorpha genome possesses orthologues encoding components of the auxin transcriptional response machinery as it has been characterized in other land plants [72,73,74]. In contrast to other land plant lineages only single orthologs of each of the three classes of ARF and single orthologs of the AUX/IAA, TIR1/AFB, and TPL gene families are present in M. polymorpha. Phylogenetic analyses also suggest that the last common ancestor of land plants had single orthologs of these classes of genes, implying that M. polymorpha has retained the ancestral condition. Single orthologs of other regulatory gene families, e.g. the class III and IV HD-ZIP transcription factor families, in the last common ancestor of land plants have also been proposed and may be a common feature of the ancestral land plant [75,76]. As the genome of the filamentous charophycean alga Klebsormidium does not encode any of the canonical auxin transcriptional response machinery [77], its origin must lie either at the base of the land plant lineage or along the charophycean algal grade leading to land plants, as is the case for several other transcriptional regulators [78].
In flowering plants, the complexity and the robustness of the auxin transcriptional response has been hypothesized to be generated by a large number of signaling network paralogs encoding components of differing affinities [10,24]. However, the relative transcriptional simplicity of the auxin response system in M. polymorpha is sufficient to pattern its complex thallus with several tissue and cell types. While the common ancestor of mosses and flowering plants is also predicted to have possessed only single orthologs of the genes in question, relatively recent gene duplications in the lineage leading to the moss P. patens has resulted in this species possessing multiple closely related paralogs [74]. As both M. polymorpha and P. patens have arguably similar levels of complexity at the gametophytic level, our findings support the idea that the relationships between auxin signaling components—rather than the number of paralogs—are the key to establish a complex body plan. Regarding the sporophyte generation, it is still possible to speculate that increases in the numbers of paralogs of the auxin genetic toolkit could be correlated with increased morphological complexity but this requires further sampling of land plant genomes, in particular those of hornworts.
Conserved auxin transcriptional machinery in M. polymorpha
Several lines of evidence indicate a conserved role for MpIAA, MpARF1 and MpTPL in auxin response. First, these genes are orthologous to those that have been shown to act downstream of auxin in developmental and physiological processes in other land plants (S4–S6 Figs). Second, similar phenotypic defects are seen in M. polymorpha when expressing MpTPL-D34 translational fusions with D34 derived from either the Arabidopsis AUX/IAA protein BDL, or the M. polymorpha MpIAA, MpARF1 or MpARF2 proteins (Fig 3D–3F). Third, proMpTPL:MpTPL-D34IAA M. polymorpha plants are insensitive to exogenous auxin, similar to the corresponding plants in Arabidopsis [9]. Fourth, MpTPL-D34 alleles and loss - and gain-of-function alleles of MpIAA, MpARF1 and MpTPL recapitulate phenotypes observed when M. polymorpha is grown in the presence of either exogenous auxin or auxin biosynthesis inhibitors (Fig 1), as is discussed in detail below. Fifth, reduction of MpIAA transcripts or ectopic expression of MpARF1 and MpTPL results in auxin hypersensistivity (Figs 2, 3 and 4J). Sixth, loss-of-function alleles of MpARF1 result in auxin hyposensitivy [69,79]. Taken together, these data strongly suggest that the MpTPL, MpARF1, and MpIAA genes encode components of an auxin transcriptional response system in M. polymorpha.
The auxin transcriptional response machinery in land plants
Our data in M. polymorpha support aspects of the model proposed for Arabidopsis explained in detail in the introduction. That indistinguishable phenotypes were observed when any of the chimeric proteins MpTPL-D34MpIAA, MpTPL-D34MpARF1, or MpTPL-D34MpARF2 were constitutively expressed suggests MpARF1 can homodimerize as well as heterodimerize with MpIAA and MpARF2 in planta. Y2H and in BiFC experiments on these proteins also suggest that they can all interact with one another, with the strongest interactions occurring between MpIAA and MpARF1 [67]. Furthermore, that proTPL:MpTPL-D34MpIAA plants are insensitive to auxin indicates that interactions between MpIAA, MpARF1, and MpARF2 mediate the activation of gene expression in response to auxin in M. polymorpha. In contrast, constitutive expression of MpTPL-D34MpARF3 chimeric proteins does not inhibit establishment of dorsiventrality or differentiation of tissues, suggesting MpARF3 does not interact effectively with MpARF1 and MpIAA proteins, pointing to a possible role of class C ARFs regulating gene expression in an auxin independent manner. Consistent with this hypothesis, is the observation that MpARF3 can’t homodimerize and its interaction with MpIAA is weaker than the interactions between MpIAA and either MpARF1 or MpARF2 [67].
Auxin patterns the development of many tissues in M. polymorpha
A large body of literature, primarily of pharmacological experiments whereby M. polymorpha has been grown in the presence of exongenous auxins, anti-auxins, and auxin transport inhibitors, has indicated auxin is involved in rhizoid formation, establishment of dorsiventral polarity in gemmalings, polar thallus regeneration, gemma and apical cell dormancy, gemmae cup development, air pore development, and spacing of organs during branching [40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,80]. As outlined below, our pharmacological and genetic studies confirm and extend the known roles of auxin in the development of multiple structures during several stages of M. polymorpha development. For the sake of clarity, the different experiments and their putative effects performed in this study are summarized in S4 Table.
Dorsiventral polarity
Auxin distribution influences establishment of the dorsiventral body plan. Consistent with earlier reports, growth of gemmae from germination on auxin containing media resulted in rhizoid formation from both dorsal and ventral surfaces. In addition, transfer of gemmae with established dorsiventrality to auxin plates caused several other developmental abnormalities, such as distorted growth of air chambers, air pores, and gemmae cups, but not changes in dorsiventrality (Fig 1A–1H). This is consistent with the observation that dorsiventrality is established within the first 24 hours of gemmaling growth and is irreversible after this time [32,45,81,82,83,84,85].
Rhizoids
Rhizoid formation is promoted by auxin in receptive tissues. When gemmae are germinated in the presence of exogenous auxin, cells on both the dorsal and ventral surfaces have the potential to differentiate into rhizoids, however, once dorsiventrality is established, dorsal cells lose the capacity to form rhizoids in response to high levels of auxin. However, in gemmalings with established polarity, auxin induces production of additional central and ventral rhizoids and they grow to greater lengths (S1B Fig). Conversely, when gemmae are grown in the presence of auxin biosynthesis inhibitors, fewer, and sometimes shorter, rhizoids develop from the central surface (Fig 1K and 1L). Finally, proMpTPL:TPL-D34IAA plants do not form rhizoids even in the presence of exogenous auxin.
Gemmae and gemmae cups
Gemmae cups are one of the most sensitive tissues to disruptions in auxin homeostasis. When grown in the presence of auxin biosynthesis inhibitors gemmae cup and gemmae formation is delayed (Fig 1I and 1J). Over-expression of MpTPL (as in proEF1:MpTPL lines), which presumably results in a dampening of auxin transcriptional response, also results in a reduction in gemmae cup growth (Fig 4H, 4K and 4L). In contrast, when thalli with established polarity are treated with exogenous auxin elongated tubular cups are formed (Fig 1E–1H). The tubular gemmae cups often show a gravitropic response opposite to the usual negative gravitropism previously described for gemma cup growth [55], and consistent with the notion that auxin transduces a gravity stimulus or inhibits a phototropic response [86,87]. Excess auxin may be transported into gemmae cups, or alternatively, auxin-conjugating enzymes (such as GH3) may metabolize auxin in most other tissues except the gemmae cup walls.
Branching
Auxin influences the rate of branching during thallus growth. Growth of the M. polymorpha thallus occurs via divisions of a single apical cell with four cutting faces [70,71]. Branching is dichotomous (S2 Fig), but the precise mechanism, e.g. via division of the apical cell into two apical cells or de novo formation of nearby apical cells, is not known. Application of the auxin biosynthesis inhibitor L-Kyn and expression of a bacterial auxin-conjugating enzyme (iaaL) appear to slow the rate of branching. L-Kyn in particular, causes a disruption of the growth phase that spatially separates apical notches (Fig 1M, 1N and 1R). Thalli often have gemmae cups produced by two adjacent apices fused into a single structure, suggesting fields of cells can act coordinately in the development of a single gemmae cup, indicating M. polymorpha exhibits a remarkable plasticity during organogenesis. In contrast, an increase in the rate of branching results from over-expression of MpARF1 and MpARF1ΔD34 (Fig 3A and 3C; S3 Table) as well as the putative reduction in MpTPL activity caused by the dominant negative construct proEF1:MpTPLN176H (Fig 4I; S2 Table). Because the overexpression of an MpIAA-insensitive MpARF1ΔD34 protein results in an enhanced phenotype compared to the one observed in MpARF1 overexpressing lines, it is unlikely that these lines have secondary loss-of-function effects as a result of increased MpIAA transcription [11,12,20,21,88]. Thus, MpARF1 facilitates activities as diverse as cell expansion (expansion of rhizoids) and pluoripotency reprograming (bifurcation).
Developmental stage transitions
Auxin is required for gametophyte developmental stage transitions. Several different MpTPL-D34 lines, especially in the cases of proEF1:MpTPL-D34MpARF1, and proEF1:MpTPL-D34MpARF2, phenotypically resemble wild type at the prothallus stage (Fig 5B–5F), with a multitude of undifferentiated unistratose lobes composed (Fig 5D). Additionally, the phenotypes of strong proEF1:iaaL lines are reminiscent of wild-type sporelings, prior to transitioning into a prothallus (Figs 1O, 1P and 5F). These results suggest that auxin is required for the morphological transitions between successive developmental stages during M. polymorpha gametophyte development—from sporeling to prothallus, which can be viewed as a transition from unorganized growth to 2-dimensional patterning, and from prothallus to thallus, which is a transition from 2-dimensional to 3-dimensional patterning. The requirement of auxin for developmental transitions has been observed in another bryophyte lineage, the mosses, where auxin is necessary to produce 3-dimensional gametophores from protonemal filaments [25,64]. The ubiquitous requirement for auxin for morphological transitions during development of these basal lineages of land plants suggests that the common ancestor of embryophytes likely utilized auxin as an internal cue to promote morphological transitions. Given the origin of the auxin transcription response machinery was at or near the origin of land plants and that a key innovation of land plants is development of 3-dimensional bodies from an apical cell of at least three cutting faces, it is plausible that the incorporation of auxin response into developmental programs facilitated the evolution of complex land plant body plans.
Organized tissue patterning
In many proEF1:iaaL lines there is little, if any, organized tissue growth even though the plants continue to grow for months, suggesting that a reduction in auxin levels, that while not inimical to life, results in an almost complete loss of tissue patterning in M. polymorpha (Fig 1). Auxin insensitive proMpTPL:MpTPL-D34IAA and proEF1:MpTPL-D34IAA plants can produce many of the tissues found in a wild-type thallus, including air chambers, air pores, gemmae cups and gemmae. However, the organization of the tissues relative to one another and of the tissues themselves is severely disrupted (Fig 5B–5G). Such distortions of tissue patterning, (e.g. shapes of air chambers, air pores, gemmae cups), are also observed when thalli are grown in the presence of exogenous auxin or auxin biosynthesis inhibitors. Taken together, these data suggest that auxin, or processes downstream of auxin are required for not only developmental transitions, but also for the patterning of tissues relative to each other and development most tissue types in the M. polymorpha gametophyte.
In a classic experiment nearly a century ago, geotropic sensitivity to decapitated roots was restored by placing either another root tip or, alternatively, a shoot tip on the decapitated stump, thus demonstrating that auxins produced in the shoot and root tips do not have specificity of action, but rather act in a context dependent manner [89]. Given that auxin acts in the patterning of most tissue and cell types of the M. polymorpha gametophyte and the lack of redundancy in its components of the auxin transcriptional response machinery, there appears to be no tissue or cell type specificity of action with respect to auxin signaling components. Rather, it seems that since the origin of land plants, auxin acted as a facilitator rather than a determiner of cell fate, with the specific output of ARF action depending upon the milieu of other transcription factors present in a particular cell [90,91].
Materials and Methods
Plant growth conditions
Sporangia from an Australian population of Marchantia polymorpha (from now on referred to as wild type) were collected from a southeastern Melbourne field location (37°57'48.36"S, 145° 6'20.41"E). Sporangia were sterilized in 1x sterilization solution (2% sodium hypochlorite, 0.1% TritonX-100) for 1 minute and subsequently washed with water. Sporangia were then dried and stored at 4°C.
Wild-type spores were suspended in water and plated on petri dishes with Gamborg’s B5 basal medium (B5; Austratech,). Dormant gemmae from adult plants were subcultured for pharmacological assays. Spores were also grown in liquid 0M51C media [92] for transformation (see transformation section below).
Pharmacological assays
Dormant gemmae, or gemmalings grown on B5 media for 2, 4, 6, 8, 10, 12, 14 and 16 days, were plated on B5 media with 10 μM 2,4-D or NAA (Sigma-Aldrich). L-Kyneurenine (Sigma-Aldrich) was used in a similar fashion as exogenous auxin.
Phylogenetic analyses
Nucleotide sequence alignments for seven species with complete genomes (Physcomitrella patens, Selaginella moellendorffii, Picea abies, Pseudotsuga menziesii (used only for TPL), Amborella trichopoda, Solanum lycopersicum and Arabidopsis thaliana) plus the isolated M. polymorpha genes (see below) were created manually using Se-Alv2.0a11 [93]. Ambiguous alignment regions were trimmed for phylogenetic inferences. Bayesian phylogenies were performed using Mr. Bayes 3.2.1 [94]. A nucleotide TPL phylogeny with 3111 characters and 23 taxa was performed assuming variable rates between first, second and third positions (prset ratepr = variable). The analysis was run for 500,000 generations, sufficient for two simultaneous runs to converge with statistical significance (P <0.01). A nucleotide AUX/IAA phylogeny with 309 characters and 49 taxa was performed assuming variable rates between first, second and third positions (prset ratepr = variable). The analysis was run for 5,000,000 generations, sufficient for two simultaneous runs to converge with statistical significance (P <0.01). For the ARF phylogeny, a nucleotide matrix with embryophyte sequences (44 taxa, 996 characters) were run in Mr. Bayes for 750,000 generations assuming variable rates between first, second and third positions (prset ratepr = variable). Each analysis was run until the standard deviation of split frequencies dropped below 0.01. For all phylogenies, 25% of trees were discarded in the burn-in phase.
Cloning and plasmids
All primer sequences are available in S5 Table. The MpEF1 promoter was amplified with primers MpEF1-NdeI-F and MpEF1-SalI-R, cloned into pCRII-TOPO (Invitrogen) and sequenced. proEF1 was subcloned into the shuttle vector BJ36 using NdeI and SalI. The resulting plasmid was called proEF1 BJ36 and was used to create constitutive expression constructs described below.
Cloning of MpSHI promoter and proMpSHI:GUS
A 3981bp promoter/UTR fragment of the single SHI/STY ortholog in M. polymorpha, MpSHI, was amplified using primers ME1 and ME2. The fragment was cloned in pCRII-TOPO, creating pME68, and subsequently transferred to pRITA, which has MCS upstream of a GUS reporter gene and transcriptonal terminator sequence [95] using NdeI and KpnI, creating pME82. A proMpSHI:GUS NotI fragment from pME82 was later subcloned to pSKF HART. pSKF HART, created by Dr. Sandra Floyd, is a binary vector based on pBART (used for Arabidopsis transformation), but in contrast to the BASTA/Glufosinate resistance gene in pBART, HART has two CaMV35S promoters in tandem driving an hptII gene (Hygromicin resistance). An NdeI/KpnI proMpSHI fragment from pME68 was subcloned into proEF1 BJ36, replacing the EF1 promoter, resulting in plasmid pME159.
EF1:iaaL and MpSHI:iaaL constructs
A bacterial iaaL CDS was amplified from plasmid iaaL-BJ36 with primers ME83 and ME84. The resulting fragment was cloned in pCRII-TOPO and sequenced. iaaL was subcloned into proEF1 BJ36 and pME159 using KpnI and HindIII. The resulting proEF1:iaaL and proMpSHI:iaaL fragments were then subcloned into pSKF HART.
Cloning and mutagenesis of MpTPL
A 1kb MpTPL fragment was cloned from wild-type M. polymorpha thallus cDNA using a nested PCR approach with degenerate primers TPL-F1, TPL-F2, TPL-R1 and TPL-R2, flanking a conserved region in TPL transcripts. The obtained fragment was cloned in pCRII-TOPO and sequenced. For 5’RLM-RACE outer primers (UPM and TPLR4) and inner primers (NUP and TPLR3) were used. For 3’RACE TPLF3, TPLF4 and a GAGA primer were used. All fragments were cloned in pCRII-TOPO and sequenced. A full-length transcript was amplified using primers MpTPL-SalI-F and MpTPL-EagI-R and cloned and sequenced in pCRII-TOPO. This clone was named 1416G. PCR based mutagenesis was used to create MpTPLN176H. An initial set of two PCR fragments were amplified using primers M13F and MpTPL(N176H)-R (fragment 1), and primers MpTPL(N176H)-F and M13R (fragment 2), using 1416G as template. Both fragments 1 and 2 were used as template in a subsequent PCR reaction and amplified with primers M13F and M13R. The PCR product (named 1626) was cloned in pCRII-TOPO and sequenced.
MpTPL functional constructs
MpTPL was cut from 1416G with EcoRI and inserted in EF1proBJ36V2.0 resulting in proEF1:MpTPL BJ36. proEF1:MpTPL was subcloned into pSKF HART using NotI. MpTPLN176H was cloned into entry Gateway vector pE2B using EcoRI (replacing the ccdB gene), and subsequently recombined into PKIGWB2 (Gateway binary vector provided by Dr. Kimizune Ishizaki and Prof. Takayuki Kohchi).
To construct MpTPL-D34 fusions, an MpTPL transcript without stop codon was amplified using primers MluI-MpTPL-ATG-F and EcoRI-MpTPL-NoSTOP-R and cloned into EF1-BJ36V2.0 using MluI and EcoRI. D34 of MpIAA (EcoRI/HindIII), MpARF1 (EcoRI/EcoRI), MpARF2 (EcoRI/HindIII), MpARF3 (EcoRI/HindIII) were amplified from wild-type thallus cDNA with primers adding restriction enzyme cleavage sites (indicated above) and cloned and sequenced in pCRII-TOPO. Subsequently, each D34 was subcloned into proEF1:MpTPLNoSTOP BJ36. All resulting constructs were then subcloned into pSKF HART using NotI.
Fusions using the endogenous MpTPL promoter were made by excising the MpTPL promoter with NdeI and MluI and subcloning it into proEF1:MpTPL-D34MpIAA BJ36 and subsequently subcloning into pSKF HART using NotI.
MpTPL reporters
Promoter sequences were obtained by assembling partial 454 reads from the first M. polymorpha genome drafts provided by JGI. Once a putative assembly was made, experimental corroboration of promoters were performed using PCR. A 3.6Kb fragment of the MpTPL promoter was amplified using primer MpTPLproF5-NdeI and MpTPLproR1-SalI, and cloned and sequenced in pCRII-TOPO (named 3685). The MpTPL promoter was subcloned into 3xVENUSN7BJ36 using NdeI and SalI. 3xVENUSN7 has 3 copies of YFP Venus with a C-terminal Nuclear localized sequence [96]. proMpTPL:3xVENUSN7 BJ36 was subcloned into pSKF HART using NotI.
Cloning of MpARF1
MpARF1 was amplified using RACE with nested PCRs. For 5’RACE, primers UPM, MpARF(miR167)R4-out, NUP and MpARF(miR167)R3-in were used. For 3’RACE, GAGA and ARFmiR167F were used. Full length transcripts were amplified with MpARF1(BamHI)-F1 and MpARF1(EcoRV)-R1 and cloned and sequenced in pCRII-TOPO (Invitrogen). The full-length transcript was subcloned into pE2B using BamHI and EcoRV. The resulting plasmid was subcloned into a modified version of pSKF HART with attR1 and attR2 sites that allow a Gateway LR recombination (Invitrogen). The dominant MpARF1ΔD34 allele was created by assembly PCR, erasing the internal NotI site with a silent mutation using Universal M13 Primers and primers MpARF1-mNotI-F2 and MpARF1-mNotI-R2. A second round of PCR using primers MpARF1-ATG-F-KpnI and MpARF1-DeltaD34-R-HindIII removed domains D34 of MpARF1. The resulting fragment was sequenced and subcloned into EF1:BJ36 using KpnI/HindIII. All constructs were subcloned into HART binary vector using NotI.
Cloning of amiRs
amiRs 268 bp in length were designed in silico with flanking KpnI and HindIII sites and synthesized at GeneScript (USA). amiRs were subcloned into EF1:BJ36 using KpnI and HindIII and later into HART or KART as described in previous sections.
RLM-RACE
RNA was extracted using QIAGEN Plant RNAeasy Kit. Total RNA was ligated to an RNA oligo (First Choice RLM-RACE) using T4 RNA ligase at 37° for one hour. Samples were precipitated using Phenol:Chlroform and eluates were used as a template for cDNA synthesis using Bioscript Reverse Transcriptase (Bioline), according to manufacturers instructions. PCRs were performed using Ex-Taq (TAKARA) using nested PCR using 5’RACE-Out-F and 5’RACE-in-F (First Choice RLM-RACE) as forward primers. To detect cleaved MpIAA transcripts, RLM-MpIAA-R3 and RLM-MpIAA-R1 were used as outer and inner reverse primers, respectively.
Branching rates measurements
Primary sporeling-transformants expressing either proEF1:MpTPLN176H or a HygRes control T-DNA were simultaneously plated and notches were counted by eye after 25 days of growth in B5 media (N = 17).
Plant transformation and culture
M. polymorpha transformation was performed essentially as described previously [92]. M. polymorpha spores were grown in 25ml liquid 0M51C media for 9 to 11 days prior to Agrobacterium co-cultivation. Agrobacterium strain GV3001 was transformed with binary vectors and cultured for 2 days at 28°C in LB media whereafter the bacteria was pelleted and cultured for an additional 4 hours in 10ml 0M51C media with 100μM acetorsyringone (Austratech). 500ul bacterial suspension and acetosyringone (100μM final concentration) was added to 25ml spore suspension. Spores and bacteria were co-cultivated for 48–60 hours. Primary transformants (T1 generation) were selected on Gamborgs B5 plates with 10μg/ml Hygromicin (Sigma-Aldrich) and/or G418 (Sigma-Aldrich) and 200μM Timentin (Austratech). After 10–15 days, survivors were re-plated to a second round of selection. After approximately 1 week, survivors were transferred to unsupplemented B5 media or stored at 4°C for later use. Gemmae from the T1 generation (G1 generation) or gemmae from the G1 generation (G2) were used for all experiments, except in cases where no viable gemmae were formed in T1 plants.
Microscopy
Plants were observed in a Lumar.V12 dissecting microscope (Zeiss) and photographed with an AxioCam HRc (Zeiss). Fixation for SEM was performed in FAA (ethanol 50%, acetic acid 5%, Formaldehyde 10%) overnight at 4°C. Samples were dehydrated in ethanol before critical point drying in a CPD 030 (Baltec). Samples were sputter coated with gold using a SCD 005 (Baltec). Scanning Electron Microscopy was performed with microscope S-570 (Hitachi) at 10kV and photos were digitalized using SPECTRUM Software (Dindima).
Supporting Information
Zdroje
1. Benkova E, Ivanchenko MG, Friml J, Shishkova S, Dubrovsky JG (2009) A morphogenetic trigger: is there an emerging concept in plant developmental biology? Trends in Plant Science 14 : 189–193. doi: 10.1016/j.tplants.2009.01.006 19285906
2. Zhao Y, Christensen SK, Fankhauser C, Cashman JR, Cohen JD, et al. (2001) A role for flavin monooxygenase-like enzymes in auxin biosynthesis. Science 291 : 306–309. 11209081
3. Zhao Y, Hull AK, Gupta NR, Goss KA, Alonso J, et al. (2002) Trp-dependent auxin biosynthesis in Arabidopsis: involvement of cytochrome P450s CYP79B2 and CYP79B3. Genes & Development 16 : 3100–3112.
4. Stepanova AN, Robertson-Hoyt J, Yun J, Benavente LM, Xie DY, et al. (2008) TAA1-mediated auxin biosynthesis is essential for hormone crosstalk and plant development. Cell 133 : 177–191. doi: 10.1016/j.cell.2008.01.047 18394997
5. Benkova E, Michniewicz M, Sauer M, Teichmann T, Seifertova D, et al. (2003) Local, efflux-dependent auxin gradients as a common module for plant organ formation. Cell 115 : 591–602. 14651850
6. Friml J, Vieten A, Sauer M, Weijers D, Schwarz H, et al. (2003) Efflux-dependent auxin gradients establish the apical-basal axis of Arabidopsis. Nature 426 : 147–153. 14614497
7. Petrasek J, Friml J (2009) Auxin transport routes in plant development. Development 136 : 2675–2688. doi: 10.1242/dev.030353 19633168
8. Staswick PE, Serban B, Rowe M, Tiryaki I, Maldonado MT, et al. (2005) Characterization of an Arabidopsis enzyme family that conjugates amino acids to indole-3-acetic acid. The Plant Cell 17 : 616–627. 15659623
9. Szemenyei H, Hannon M, Long JA (2008) TOPLESS mediates auxin-dependent transcriptional repression during Arabidopsis embryogenesis. Science 319 : 1384–1386. doi: 10.1126/science.1151461 18258861
10. Vernoux T, Brunoud G, Farcot E, Morin V, Van den Daele H, et al. (2011) The auxin signalling network translates dynamic input into robust patterning at the shoot apex. Molecular Systems Biology 7: doi: 10.1038/Msb.2011.39
11. Ulmasov T, Hagen G, Guilfoyle TJ (1997) ARF1, a transcription factor that binds to auxin response elements. Science 276 : 1865–1868. 9188533
12. Ulmasov T, Murfett J, Hagen G, Guilfoyle TJ (1997) Aux/IAA proteins repress expression of reporter genes containing natural and highly active synthetic auxin response elements. The Plant Cell 9 : 1963–1971. 9401121
13. Kim J, Harter K, Theologis A (1997) Protein-protein interactions among the Aux/IAA proteins. Proceedings of the National Academy of Sciences of the United States of America 94 : 11786–11791. 9342315
14. Kepinski S, Leyser O (2005) The Arabidopsis F-box protein TIR1 is an auxin receptor. Nature 435 : 446–451. 15917798
15. Gray WM, Kepinski S, Rouse D, Leyser O, Estelle M (2001) Auxin regulates SCF(TIR1)-dependent degradation of AUX/IAA proteins. Nature 414 : 271–276. 11713520
16. Dharmasiri N, Dharmasiri S, Estelle M (2005) The F-box protein TIR1 is an auxin receptor. Nature 435 : 441–445. 15917797
17. Tiwari SB, Wang XJ, Hagen G, Guilfoyle TJ (2001) AUX/IAA proteins are active repressors, and their stability and activity are modulated by auxin. The Plant Cell 13 : 2809–2822. 11752389
18. Finet C, Berne-Dedieu A, Scutt CP, Marletaz F (2012) Evolution of the ARF Gene Family in Land Plants: Old Domains, New Tricks. Molecular Biology and Evolution 30 : 45–56. doi: 10.1093/molbev/mss220 22977118
19. Tiwari SB, Hagen G, Guilfoyle T (2003) The roles of auxin response factor domains in auxin-responsive transcription. The Plant Cell 15 : 533–543. 12566590
20. Ulmasov T, Hagen G, Guilfoyle TJ (1999) Activation and repression of transcription by auxin-response factors. Proceedings of the National Academy of Sciences of the United States of America 96 : 5844–5849. 10318972
21. Ulmasov T, Hagen G, Guilfoyle TJ (1999) Dimerization and DNA binding of auxin response factors. The Plant Journal 19 : 309–319. 10476078
22. Theologis A (1985) Rapid Gene-Regulation by Auxin in Pea Tissue. In Vitro Cellular & Developmental Biology 21: A39–A39.
23. Abel S, Nguyen MD, Theologis A (1995) The Ps-Iaa4/5-Like Family of Early Auxin-Inducible Messenger-Rnas in Arabidopsis-Thaliana. Journal of Molecular Biology 251 : 533–549. 7658471
24. Villalobos LIAC, Lee S, De Oliveira C, Ivetac A, Brandt W, et al. (2012) A combinatorial TIR1/AFB-Aux/IAA co-receptor system for differential sensing of auxin. Nature Chemical Biology 8 : 477–485. doi: 10.1038/nchembio.926 22466420
25. Prigge MJ, Lavy M, Ashton NW, Estelle M (2010) Physcomitrella patens Auxin-Resistant Mutants Affect Conserved Elements of an Auxin-Signaling Pathway. Current Biology 20 : 1907–1912. doi: 10.1016/j.cub.2010.08.050 20951049
26. Cooke TJ, Poli D, Sztein AE, Cohen JD (2002) Evolutionary patterns in auxin action. Plant molecular biology 49 : 319–338. 12036257
27. Kenrick P, Crane PR (1997) The origin and early evolution of land plants: a cladistic study. Washington, D.C.: Smithsonian Institution Press. 21238287
28. Qiu YL, Li LB, Wang B, Chen ZD, Knoop V, et al. (2006) The deepest divergences in land plants inferred from phylogenomic evidence. Proceedings of the National Academy of Sciences of the United States of America 103 : 15511–15516. 17030812
29. Mishler BD, Churchill SP (1984) A Cladistic Approach to the Phylogeny of the Bryophytes. Brittonia 36 : 406–424.
30. Forrest LL, Davis EC, Long DG, Crandall-Stotler BJ, Clark A, et al. (2006) Unraveling the evolutionary history of the liverworts (Marchantiophyta): multiple taxa, genomes and analyses. Bryologist 109 : 303–334.
31. Berrie GK (1963) Cytology and Phylogeny of Liverworts. Evolution 17 : 347–357.
32. Mirbel C-F (1835) Researches anatomiques et physiologiques sur le Marchantia polymorpha. Mém Acad Roy Sc Inst France 13 : 337–436.
33. Kny L (1890) Botanische wandtafeln mit erläuterndem text. Berlin: Wiegandt. Hempel & Parey.
34. O'Hanlon ME (1926) Germination of Spores and Early Stages in Development of Gametophyte of Marchantia polymorpha. Botanical Gazette 82 : 215–222.
35. Leitgeb H (1881) Untersuchungen über die Lebermoose. Heft VI. Die Marchantieen. Jena: Leuschner & Lubensky.
36. Douin R (1921) Reserches sur les Marchantiées. Revue générale de botanique 33 : 34–62; 99–145; 190–209. doi: 10.1007/s00221-011-2827-4 21842191
37. Barnes CR, Land WJG (1908) Bryological papers II. The origin of the cupule of marchantia—Contributions from the hull botanical laboratory 120. Botanical Gazette 46 : 401–409.
38. Hedwig J (1783) Theoria Generationis et Fructificationis Plantarum Cryptogamicarum Linnaei, mere propriis Observationibus et Experimentis Superstructa. Petropoli: Academiae Imper. Scientiarum.
39. Hofmeister WFB (1862) On the germination, development, and fructification of the higher Cryptogamia, and on the fructification of the Coniferae. Currey F, translator. London: Ray Society. 506 p.
40. Ishizaki K, Nonomura M, Kato H, Yamato KT, Kohchi T (2012) Visualization of auxin-mediated transcriptional activation using a common auxin-responsive reporter system in the liverwort Marchantia polymorpha. Journal of Plant Research 125 : 643–651. doi: 10.1007/s10265-012-0477-7 22311005
41. Tarén N (1958) Regulating the Initial Development of Gemmae in Marchantia polymorpha. The Bryologist 61 : 191–204.
42. Halbsguth W, Kohlenbach H-W (1953) Einige versuche über die wirkung von heteroauxin auf die symmetrieentwicklung der brutkörperkeimlinge von Marchanta polymorpha L. Planta 42 : 349–366.
43. Kaul KN, Mitra GC, Tripathi BK (1962) Responses of Marchantia in Aseptic Culture to Well-known Auxins and Antiauxins. Annals of Botany 26 : 447–466.
44. Davidonis GH, Munroe MH (1972) Apical Dominance in Marchantia—Correlative Inhibition of Neighbor Lobe Growth. Botanical Gazette 133 : 177–184.
45. Fitting H (1939) Untersuchungen über den Einfluss von Licht und Dunkelheit auf die Entwicklung von Moosen. I. Die Brutkörper der Marchantieen. Jahrbücher für wissenschaftliche Botanik 88 : 633–722
46. Rousseau J (1950) Action de l'acide indol beta-acétique sur les propagules de marchantia polymorpha et Lunularia cruciata. Comptes rendus hebdomadaires des séances de l'Académie des sciences 230 : 675–676. 17597171
47. Rousseau J (1951) Action de l'acide x napthaléne acétique sur les corbeilles á propagules de Marchantia polymorpha L. et de Lunularia cruciata Adans. Comptes rendus hebdomadaires des séances de l'Académie des sciences 232 : 107–108. 17597171
48. Rousseau J (1951) Action des acides 2.4-dichloro-phénoxy-acétique et 2.5-dichloro-thio-acétique sur les propagules de Marchantia polymorpha L. Comptes rendus hebdomadaires des séances de l'Académie des sciences 232 : 749–751. 17597171
49. Rousseau J (1952) Influence des hétéroauxines sur la croissance des corbeilles a propagules de Marchantia polymorpha L. et de Lunularia cruciata Adans. Revue Bryologique et Lichénologique 75 : 239–241.
50. Rousseau J (1953) Action de hétéroauxines sur les thalles de Lunularia cruciata Adans. et de Marchantia polymorpha L. Revue Bryologique et Lichénologique 76 : 22–25.
51. Rousseau J (1953) Action des hetero-auxines sur les chapeaux du Marchantia polymorpha L. Bulletin de la Société Botanique de France 100 : 179–180.
52. Kohlenbach HW (1957) Die Bedeutung des heretoauxins für die entwicklung der Dorsiventralität der Brutkörperkeimlinge von Marchantia polymorpha L. Biologishes Zentralblatt 76 : 70–125.
53. Maravolo NC, Voth PD (1966) Morphogenic Effects of 3 Growth Substances on Marchantia Gemmalings. Botanical Gazette 127 : 79–86.
54. Otto K-R (1976) Der Einfluss von ausseren Faktoren auf die Bildung von Primärrhizoiden bei Brutkörpern von Marchantia polymorpha 1. Z Pflanzenphysiol 80 : 189–196.
55. Miller MW, Voth PD (1962) Geotropic Responses of Marchantia. The Bryologist 65 : 146–154.
56. Keller CP, Van Volkenburgh E (1997) Auxin-Induced Epinasty of Tobacco Leaf Tissues (A Nonethylene-Mediated Response). Plant physiology 113 : 603–610. 12223629
57. He WR, Brumos J, Li HJ, Ji YS, Ke M, et al. (2011) A Small-Molecule Screen Identifies L-Kynurenine as a Competitive Inhibitor of TAA1/TAR Activity in Ethylene-Directed Auxin Biosynthesis and Root Growth in Arabidopsis. The Plant Cell 23 : 3944–3960. doi: 10.1105/tpc.111.089029 22108404
58. Tao Y, Ferrer JL, Ljung K, Pojer F, Hong FX, et al. (2008) Rapid synthesis of auxin via a new tryptophan-dependent pathway is required for shade avoidance in plants. Cell 133 : 164–176. doi: 10.1016/j.cell.2008.01.049 18394996
59. Nomura T, Itouga M, Kojima M, Kato Y, Sakakibara H, et al. (2014) Copper mediates auxin signalling to control cell differentiation in the copper moss Scopelophila cataractae. Journal of Experimental Botany 66 : 1205–1213. doi: 10.1093/jxb/eru470 25428998
60. Glass NL, Kosuge T (1986) Cloning of the Gene for Indoleacetic Acid-Lysine Synthetase from Pseudomonas-Syringae Subsp Savastanoi. Journal of Bacteriology 166 : 598–603. 3084452
61. Romano CP, Hein MB, Klee HJ (1991) Inactivation of Auxin in Tobacco Transformed with the Indoleacetic-Acid Lysine Synthetase Gene of Pseudomonas-Savastanoi. Genes & Development 5 : 438–446.
62. Landberg K, Pederson ERA, Viaene T, Bozorg B, Friml J, et al. (2013) The Moss Physcomitrella patens Reproductive Organ Development Is Highly Organized, Affected by the Two SHI/STY Genes and by the Level of Active Auxin in the SHI/STY Expression Domain. Plant physiology 162 : 1406–1419. 23669745
63. Althoff F, Kopischke S, Zobell O, Ide K, Ishizaki K, et al. (2014) Comparison of the MpEF1α and CaMV35 promoters for application in Marchantia polymorpha overexpression studies. Transgenic Research 23 : 234–244.
64. Eklund DM, Thelander M, Landberg K, Staldal V, Nilsson A, et al. (2010) Homologues of the Arabidopsis thaliana SHI/STY/LRP1 genes control auxin biosynthesis and affect growth and development in the moss Physcomitrella patens. Development 137 : 1275–1284. doi: 10.1242/dev.039594 20223761
65. Kuusk S, Sohlberg JJ, Magnus Eklund D, Sundberg E (2006) Functionally redundant SHI family genes regulate Arabidopsis gynoecium development in a dose-dependent manner. The Plant Journal 47 : 99–111. 16740146
66. Sohlberg JJ, Myrenas M, Kuusk S, Lagercrantz U, Kowalczyk M, et al. (2006) STY1 regulates auxin homeostasis and affects apical-basal patterning of the Arabidopsis gynoecium. The Plant Journal 47 : 112–123. 16740145
67. Kato H, Ishizaki K, Kouno M, Shirakawa M, Bowman JL, et al. (2015) Auxin-mediated transcriptional system with a minimal set of components is critical for morphogenesis through the life cycle in Marchantia polymorpha. PLOS Genetics 11: doi: 10.1371/journal.pgen.1005084
68. Alvarez JP, Pekker I, Goldshmidt A, Blum E, Amsellem Z, et al. (2006) Endogenous and synthetic microRNAs stimulate simultaneous, efficient, and localized regulation of multiple targets in diverse species. Plant Cell 18 : 1134–1151. 16603651
69. Flores-Sandoval E, Dierschke T, Fisher TJ, Bowman JL (2015) Efficient and inducible use of artificial microRNAs in Marchantia polymorpha. Plant and Cell Physiology 56: in press.
70. Douin MR (1923) Reserches sur le gamétophyte des Marchantiées. Revue générale de botanique 35 : 213–226. doi: 10.1007/s00221-011-2827-4 21842191
71. Apostolakos P, Galatis B, Mitrakos K (1982) Studies on the Development of the Air Pores and Air Chambers of Marchantia-Paleacea. I. Light-Microscopy. Annals of Botany 49 : 377–396.
72. De Smet I, Voss U, Lau S, Wilson M, Shao N, et al. (2011) Unraveling the Evolution of Auxin Signaling. Plant physiology 155 : 209–221. doi: 10.1104/pp.110.168161 21081694
73. Banks JA, Nishiyama T, Hasebe M, Bowman JL, Gribskov M, et al. (2011) The Selaginella Genome Identifies Genetic Changes Associated with the Evolution of Vascular Plants. Science 332 : 960–963. doi: 10.1126/science.1203810 21551031
74. Rensing SA, Lang D, Zimmer AD, Terry A, Salamov A, et al. (2008) The Physcomitrella genome reveals evolutionary insights into the conquest of land by plants. Science 319 : 64–69. 18079367
75. Floyd SK, Zalewski CS, Bowman JL (2006) Evolution of Class III Homeodomain leucine zipper genes in streptophytes. Genetics 173 : 373–388. 16489224
76. Zalewski CS, Floyd SK, Furumizu C, Sakakibara K, Stevenson DW, et al. (2013) Evolution of the Class IV HD-Zip Gene Family in Streptophytes. Molecular Biology and Evolution 30 : 2347–2365. doi: 10.1093/molbev/mst132 23894141
77. Hori K, Maruyama F, Fujisawa T, Togashi T, Yamamoto N, et al. (2014) Klebsormidium flaccidum genome reveals primary factors for plant terrestrial adaptation. Nature Communications 5; doi: 10.1038/ncomms4978
78. Bowman JL (2013) Walkabout on the long branches of plant evolution. Current Opinion in Plant Biology 16 : 70–77. doi: 10.1016/j.pbi.2012.10.001 23140608
79. Sugano SS, Shirakawa M, Takagi J, Matsuda Y, Shimada T, et al. (2014) CRISPR/Cas9-Mediated Targeted Mutagenesis in the Liverwort Marchantia polymorpha L. Plant and Cell Physiology 55 : 475–481. doi: 10.1093/pcp/pcu014 24443494
80. Binns AN, Maravolo NC (1972) Apical Dominance, Polarity, and Adventitious Growth in Marchantia Polymorpha. American Journal of Botany 59 : 691–696.
81. Pfeffer W (1871) Studien über Symmetric und spezifische Wachstumsursachen. Arbeiten Des Botanischen Instituts in Würzburg 1 : 77–98.
82. Fitting H (1936) Untersuchungen über die Induktion der Dorsiventralität bei den keimenden Brutkörpern von Marchantia und Lunularia. I. Die Induktoren und ihre Wirkungen. Jahrbücher für wissenschaftliche Botanik 82 : 333–376.
83. Fitting H (1936) Untersuchungen über die Induktion der Dorsiventralität bei den Marehantieen brutkörpern. II. Die Schwerkraft als Induktor der Dorsiventralität. Jahrbücher für wissenschaftliche Botanik 82 : 696–740.
84. Fitting H (1937) Untersuchungen über die Induktion der Dorsiventralität bei den Brutkörperkeimlingen der Marchantieen. III. Das Licht als Induktor der Dorsiventralität. Jahrbücher für wissenschaftliche Botanik 85 : 169–242.
85. Fitting H (1938) Untersuchungen über die Induktion der Dorsiventralität bei den Brutkörperkeimlingen der Marchantieen. V. Die Umkehrbarkeit der durch Aussenfaktoren induzierten Dorsiventralität. Jahrbücher für wissenschaftliche Botanik 86 : 107–227.
86. Muller A, Guan C, Galweiler L, Tanzler P, Huijser P, et al. (1998) AtPIN2 defines a locus of Arabidopsis for root gravitropism control. The EMBO Journal 17 : 6903–6911. 9843496
87. Esmon CA, Tinsley AG, Ljung K, Sandberg G, Hearne LB, et al. (2006) A gradient of auxin and auxin-dependent transcription precedes tropic growth responses. Proceedings of the National Academy of Sciences of the United States of America 103 : 236–241. 16371470
88. Hardtke CS, Ckurshumova W, Vidaurre DP, Singh SA, Stamatiou G, et al. (2004) Overlapping and non-redundant functions of the Arabidopsis auxin response factors MONOPTEROS and NONPHOTOTROPIC HYPOCOTYL 4. Development 131 : 1089–1100. 14973283
89. Cholodny N (1924) Über die hormonale Wirkung der Organspitze bei der geotropischen Krümmung. Berichte der Deutschen Botanischen Gesellschaft 42 : 356–362.
90. Stewart JL, Nemhauser JL (2010) Do trees grow on money? Auxin as the currency of the cellular economy Cold Spring Harb Perspect Biol 2: a001420. doi: 10.1101/cshperspect.a001420 20182619
91. Bennett T, Leyser O (2014) The Auxin Question: A Philosophical Overview. In: Zažímalová E, Petrášek J, Benková E, editors. Auxin and Its Role in Plant Development. Vienna: Springer. pp. 3–19.
92. Ishizaki K, Chiyoda S, Yamato KT, Kohchi T (2008) Agrobacterium-mediated transformation of the haploid liverwort Marchantia polymorpha L., an emerging model for plant biology. Plant and Cell Physiology 49 : 1084–1091. doi: 10.1093/pcp/pcn085 18535011
93. Rambaut A (1996) Se-Al: Sequence alignment editor. http://tree.bio.ed.ac.uk/software/seal/
94. Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17 : 754–755. 11524383
95. Eshed Y, Baum SF, Bowman JL (1999) Distinct mechanisms promote polarity establishment in carpels of Arabidopsis. Cell 99 : 199–209. 10535738
96. Heisler MG, Ohno C, Das P, Sieber P, Reddy GV, et al. (2005) Patterns of auxin transport and gene expression during primordium development revealed by live imaging of the Arabidopsis inflorescence meristem. Current Biology 15 : 1899–1911. 16271866
Štítky
Genetika Reprodukční medicína
Článek Minor Type IV Collagen α5 Chain Promotes Cancer Progression through Discoidin Domain Receptor-1Článek A Unique Virulence Gene Occupies a Principal Position in Immune Evasion by the Malaria ParasiteČlánek The Centrosomal Linker and Microtubules Provide Dual Levels of Spatial Coordination of CentrosomesČlánek Dissecting the Function and Assembly of Acentriolar Microtubule Organizing Centers in Cells In VivoČlánek MicroRNA-Dependent Transcriptional Silencing of Transposable Elements in Drosophila Follicle CellsČlánek PARP2 Is the Predominant Poly(ADP-Ribose) Polymerase in Arabidopsis DNA Damage and Immune ResponsesČlánek Cooperative Action of Cdk1/cyclin B and SIRT1 Is Required for Mitotic Repression of rRNA SynthesisČlánek The Developmental Intestinal Regulator ELT-2 Controls p38-Dependent Immune Responses in Adult .Článek Feeding and Fasting Signals Converge on the LKB1-SIK3 Pathway to Regulate Lipid Metabolism in
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2015 Číslo 5
-
Všechny články tohoto čísla
- Genomic Heritability: What Is It?
- Triglyceride-Increasing Alleles Associated with Protection against Type-2 Diabetes
- Epistasis Is a Major Determinant of the Additive Genetic Variance in
- Genetic Regulation of Bone Metabolism in the Chicken: Similarities and Differences to Mammalian Systems
- Minor Type IV Collagen α5 Chain Promotes Cancer Progression through Discoidin Domain Receptor-1
- The Philosophical Approach: An Interview with Ford Doolittle
- Downregulation of the Host Gene by miR-92 Is Essential for Neuroblast Self-Renewal in
- A Unique Virulence Gene Occupies a Principal Position in Immune Evasion by the Malaria Parasite
- Y Fuse? Sex Chromosome Fusions in Fishes and Reptiles
- Regulation of Active DNA Demethylation by a Methyl-CpG-Binding Domain Protein in
- Overlapping Patterns of Rapid Evolution in the Nucleic Acid Sensors cGAS and OAS1 Suggest a Common Mechanism of Pathogen Antagonism and Escape
- Autoselection of Cytoplasmic Yeast Virus Like Elements Encoding Toxin/Antitoxin Systems Involves a Nuclear Barrier for Immunity Gene Expression
- Genetic Architecture of Abdominal Pigmentation in
- Whole Genome DNA Binding Analysis of the Bacterial Replication Initiator and Transcription Factor DnaA
- The Centrosomal Linker and Microtubules Provide Dual Levels of Spatial Coordination of Centrosomes
- Parp3 Negatively Regulates Immunoglobulin Class Switch Recombination
- Burden Analysis of Rare Microdeletions Suggests a Strong Impact of Neurodevelopmental Genes in Genetic Generalised Epilepsies
- Cell Cycle Control by the Master Regulator CtrA in
- Myopathic Lamin Mutations Cause Reductive Stress and Activate the Nrf2/Keap-1 Pathway
- Monoallelic Loss of the Imprinted Gene Promotes Tumor Formation in Irradiated Mice
- Phylum-Level Conservation of Regulatory Information in Nematodes despite Extensive Non-coding Sequence Divergence
- Clustering and Negative Feedback by Endocytosis in Planar Cell Polarity Signaling Is Modulated by Ubiquitinylation of Prickle
- Dissecting the Function and Assembly of Acentriolar Microtubule Organizing Centers in Cells In Vivo
- MicroRNA-Dependent Transcriptional Silencing of Transposable Elements in Drosophila Follicle Cells
- β-Catenin Signaling Biases Multipotent Lingual Epithelial Progenitors to Differentiate and Acquire Specific Taste Cell Fates
- A Simple Auxin Transcriptional Response System Regulates Multiple Morphogenetic Processes in the Liverwort
- Parallel Gene Expression Differences between Low and High Latitude Populations of and .
- The Nutrient-Responsive Hormone CCHamide-2 Controls Growth by Regulating Insulin-like Peptides in the Brain of
- Characterization of TCF21 Downstream Target Regions Identifies a Transcriptional Network Linking Multiple Independent Coronary Artery Disease Loci
- PARP2 Is the Predominant Poly(ADP-Ribose) Polymerase in Arabidopsis DNA Damage and Immune Responses
- Drosophila Spaghetti and Doubletime Link the Circadian Clock and Light to Caspases, Apoptosis and Tauopathy
- Coronary Artery Disease Associated Transcription Factor TCF21 Regulates Smooth Muscle Precursor Cells That Contribute to the Fibrous Cap
- Rescue of DNA-PK Signaling and T-Cell Differentiation by Targeted Genome Editing in a Deficient iPSC Disease Model
- Disruption of Transcriptional Coactivator Sub1 Leads to Genome-Wide Re-distribution of Clustered Mutations Induced by APOBEC in Active Yeast Genes
- Yeast Killer Elements Hold Their Hosts Hostage
- Keeping in Shape the Dogma of Mitochondrial DNA Maternal Inheritance
- Extreme-Depth Re-sequencing of Mitochondrial DNA Finds No Evidence of Paternal Transmission in Humans
- Trading Places—Switching Frataxin Function by a Single Amino Acid Substitution within the [Fe-S] Cluster Assembly Scaffold
- Natural Variation Identifies , a Universal Gene Required for Cell Proliferation and Growth at High Temperatures in
- Mutations in Gene Are Associated with Predisposition to Breast Cancer
- The Whole of a Scientific Career: An Interview with Oliver Smithies
- Cell Specific eQTL Analysis without Sorting Cells
- Cooperative Action of Cdk1/cyclin B and SIRT1 Is Required for Mitotic Repression of rRNA Synthesis
- Systemic Regulation of RAS/MAPK Signaling by the Serotonin Metabolite 5-HIAA
- Reprogramming LCLs to iPSCs Results in Recovery of Donor-Specific Gene Expression Signature
- The Developmental Intestinal Regulator ELT-2 Controls p38-Dependent Immune Responses in Adult .
- Genetic Mechanism of Human Neutrophil Antigen 2 Deficiency and Expression Variations
- Feeding and Fasting Signals Converge on the LKB1-SIK3 Pathway to Regulate Lipid Metabolism in
- Early Lineage Priming by Trisomy of Leads to Myeloproliferation in a Down Syndrome Model
- Turning into a Frataxin-Dependent Organism
- Accounting for Experimental Noise Reveals That mRNA Levels, Amplified by Post-Transcriptional Processes, Largely Determine Steady-State Protein Levels in Yeast
- Biological Significance of Photoreceptor Photocycle Length: VIVID Photocycle Governs the Dynamic VIVID-White Collar Complex Pool Mediating Photo-adaptation and Response to Changes in Light Intensity
- CTXφ Replication Depends on the Histone-Like HU Protein and the UvrD Helicase
- Disruption of miR-29 Leads to Aberrant Differentiation of Smooth Muscle Cells Selectively Associated with Distal Lung Vasculature
- Notch Is Required in Adult Sensory Neurons for Morphological and Functional Plasticity of the Olfactory Circuit
- Post-transcriptional Regulation of Keratinocyte Progenitor Cell Expansion, Differentiation and Hair Follicle Regression by
- PERK Limits Lifespan by Promoting Intestinal Stem Cell Proliferation in Response to ER Stress
- Casein Kinase 1 and Phosphorylation of Cohesin Subunit Rec11 (SA3) Promote Meiotic Recombination through Linear Element Formation
- Fibroblast Growth Factor 9 Regulation by MicroRNAs Controls Lung Development and Links Loss to the Pathogenesis of Pleuropulmonary Blastoma
- Auxin-Mediated Transcriptional System with a Minimal Set of Components Is Critical for Morphogenesis through the Life Cycle in
- The Broad-Spectrum Antiviral Protein ZAP Restricts Human Retrotransposition
- The 4E-BP Caf20p Mediates Both eIF4E-Dependent and Independent Repression of Translation
- Turning into a Frataxin-Independent Organism
- Promotion of Bone Morphogenetic Protein Signaling by Tetraspanins and Glycosphingolipids
- Essential Role of the ESX-5 Secretion System in Outer Membrane Permeability of Pathogenic Mycobacteria
- The Zinc-Finger Antiviral Protein ZAP Inhibits LINE and Alu Retrotransposition
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Drosophila Spaghetti and Doubletime Link the Circadian Clock and Light to Caspases, Apoptosis and Tauopathy
- Autoselection of Cytoplasmic Yeast Virus Like Elements Encoding Toxin/Antitoxin Systems Involves a Nuclear Barrier for Immunity Gene Expression
- Parp3 Negatively Regulates Immunoglobulin Class Switch Recombination
- PERK Limits Lifespan by Promoting Intestinal Stem Cell Proliferation in Response to ER Stress
Přihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



