-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praktické
Top novinky
ReklamaAnimal Models of Inflammatory Bowel Disease at the Dawn of the New Genetics Era
article has not abstract
Published in the journal: . PLoS Med 5(9): e198. doi:10.1371/journal.pmed.0050198
Category: Research in Translation
doi: https://doi.org/10.1371/journal.pmed.0050198Summary
article has not abstract
Crohn disease (CD) and ulcerative colitis (UC) are chronic inflammatory intestinal diseases with multifactorial etiologies. CD and UC are distinguished both by the location and by the nature of the inflammation. CD displays a transmural discontinuous inflammation, often with granulomas, in any part of the digestive system (most often ileum and/or colon). UC is almost exclusively restricted to the colon with a continuous superficial mucosal and submucosal inflammation. Both CD and UC can be further subphenotyped, suggesting that there is heterogeneity within each disorder. Despite many clinical and pathological features that distinguish CD and UC, the collective term inflammatory bowel disease (IBD) is often used for the two diseases. Clustering of IBD in families without specificity for a given form of IBD supports the notion of common genetic factors in the etiologies of the two conditions. In addition, higher concordance rates in monozygotic twins than dizygotic twins, particularly in CD, points to the importance of genes [1]. Recent advances in genetics have proven that both CD and UC are truly polygenic, but there are also strong environmental influences on IBD. This notion is first and foremost supported by the rapid increase in incidence of IBD during the past 50 years.
Understanding the complex interplay between genetic and environmental factors that lead to IBD is a formidable challenge. Animal models have been very helpful in this respect, providing fundamental insight into the importance of immunologic dysregulation and intestinal microbiota [2,3]. Animal models are continuing to give new insights into the pathogenesis of IBD, as shown by two recent papers in PLoS Medicine [4,5]. Simultaneously, there have been remarkable advances in the understanding of the genetics of human IBD, and the mapping of IBD genes is continuously progressing [6–8]. The new genetic findings bear promise for new and better therapies. This translation of basic science into the clinic will be much facilitated by relevant and good animal models. We can thus expect that the new genetic findings will set the stage for future development of animal models in IBD.
Animal Models of IBD
Many strains of mice with genetic defects in immune regulation develop colitis (Figure 1). Such genetic defects include targeted deletion of receptors for the regulatory cytokines transforming growth factor (TGF)-ß or interleukin (IL)-10, and other mutations causing a disruption of signaling from these cytokines [9]. Specific T cell involvement in dysregulation can be shown by transfer of a subpopulation of T cells (CD4+CD45RB+) to lymphopenic hosts, which leads to colitis [10]. It was early established that cotransfer of CD4+CD25+ T cells could suppress colitis development, and the concept that FoxP3+ regulatory T (Treg) cells play an important general anti-inflammatory role is now also established in human studies [11].
Fig. 1. Mucosal Homeostasis in the Gut 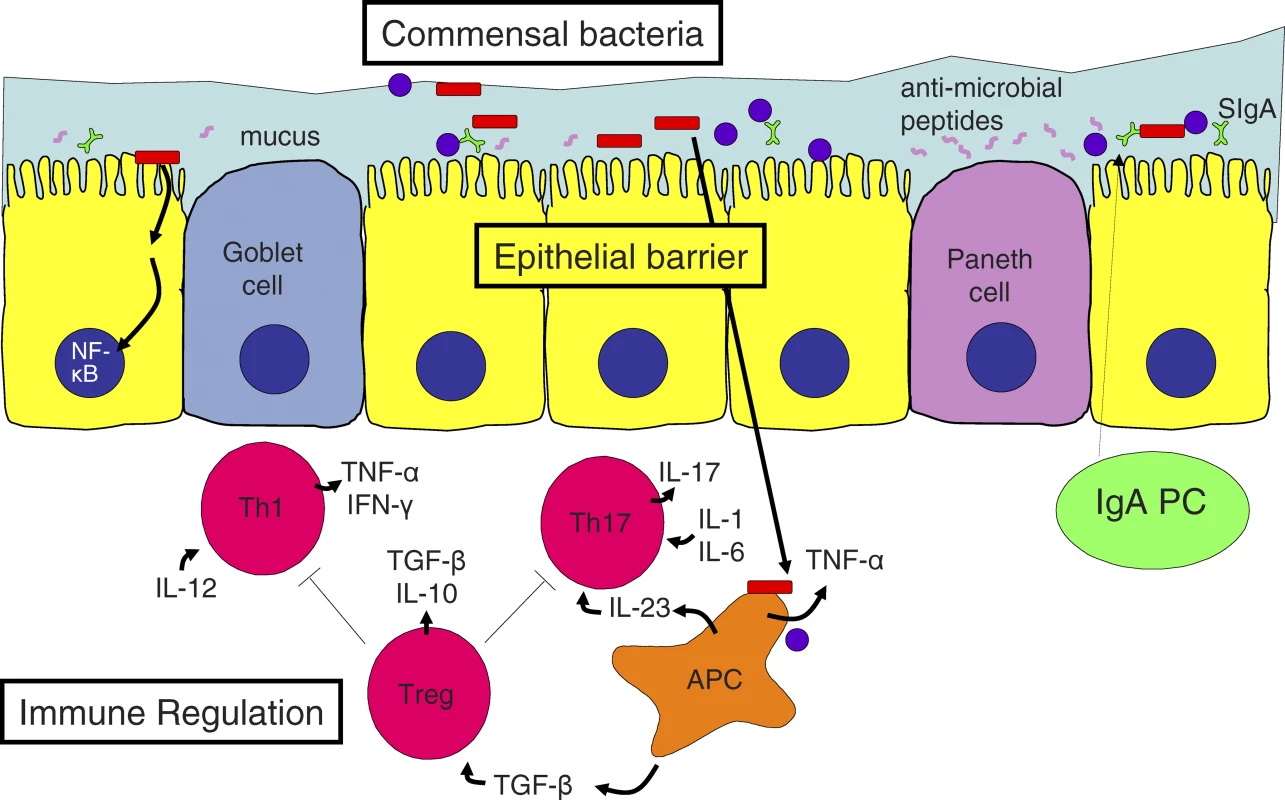
Goblet cell-secreted mucus, Paneth cell-secreted antimicrobial peptides, and secretory immunoglobulin A (SIgA), generated from epithelial transport of dimeric IgA produced by lamina propria plasma cells (PC), cooperate to reinforce epithelial barrier function. Breach of this barrier causes excessive stimulation of the mucosal immune system by commensal bacteria and leads to pro-inflammatory T cell–mediated immune responses. Gene deficiencies in immune regulatory factors may also lead to intestinal inflammation. Commensal flora and bacterial products trigger antigen presenting cells (APC) to produce several cytokines including TNF-α, IL-23, and TGF-β, depending on context. Epithelial activation of the transcription factor NF-κB by bacterial products may enhance host resistance by reinforcing the epithelial barrier. A mixed Th1/Th17-driven inflammation is controlled by regulatory T cells (Treg) through suppressive cytokines (TGF-ß and IL-10) and other mechanisms. Some key cytokines important for T helper cell differentiation or for their function are shown. Appropriate barrier function of the epithelium is critical to gut homeostasis (Figure 1) [12]. An early demonstration of this concept was the spontaneous intestinal inflammation occurring in mice with mucosal leakiness caused by dominant negative epithelial N-cadherin [13]. Although the picture in these mice is complicated by epithelial proliferation, apoptosis, and neoplasms, further support of the barrier concept comes from the fact that mice lacking the major component of intestinal mucus, Muc2−/− mice, develop spontaneous colitis [14]. Furthermore, mice with enterocyte-specific ablation of nuclear factor kappa B (NF-κB) activation fail to control the luminal microbial flora, and inflammation is triggered by an exaggerated immune response to invading bacteria [15,16]. Other defects in innate or adaptive defense, or chemical irritation of the colonic mucosa, may have similar outcomes. For instance, dextran sulfate sodium added to drinking water causes colitis by this mechanism and has been used to identify a number of genes required for appropriate protection and reinforcement of the epithelial barrier.
Our intimate and complex relationship with the 1013–1014 bacteria that reside in our colons has been explored in animal models. Activation of Toll-like receptor (TLR) signaling by luminal commensals has been shown to provide cytoprotective signals to the host [17]. However, commensals can also trigger disease development, because in germ-free conditions mice with defective immune regulation do not develop colitis [18,19]. Interestingly, a recent paper showed that a colitogenic microbial community spontaneously developed in susceptible Rag2/T-Bet double knockout mice [20]. This microbial community could transmit colitis to wild-type mice, and colitis was effectively cured by broad-spectrum oral antibiotics. Importantly, the animal models and analysis of immune responses in patients suggest that the immune response in IBD is largely directed against the luminal microbiota, rather than being a classical autoimmune disease. The presence of autoantibodies and extraintestinal symptoms in some patients are thus probably secondary to the intestinal inflammation.
A hallmark of IBD is an increase in the level of pro-inflammatory cytokines. The best studied of these are tumor necrosis factor (TNF)-α and interferon (IFN)-γ, but animal experiments also identified IL-23 as a contributor to intestinal inflammation [2]. This cytokine is a heterodimer of p40 (encoded by IL-12B) and a unique p19 subunit and functions by driving and/or sustaining a Th17 response. Recently, Th17 cells have been documented in human IBD [21], and the central involvement of Th17 cells is further supported by recent genetic findings (see below).
Two recent publications in PLoS Medicine shed new light on colitis development. Heazlewood et al. [4], in a model representative of reduced epithelial barrier, studied two independent noncomplementing strains of mutant mice that developed spontaneous colitis. Both strains had a missense mutation in the Muc2 gene. Mutant Muc2 oligomerized inappropriately, causing an endoplasmic reticulum (ER) stress response in goblet cells with subsequent reduced mucus secretion and concomitant increase in lamina propria proinflammatory cytokines. Importantly, the authors found MUC2 precursor accumulation and ER stress in human UC, even in noninflamed areas, suggesting that UC patients (at least some) have genetic alterations leading to ER stress and aberrant mucin assembly.
Kang et al. [5], in a model representative of immune dysregulation, reasoned that genetic predisposition to IBD is caused by interaction of several genes, while gene-targeted mouse models rely on the absence of a single gene product. Thus, they generated a mouse line with defective IL-10 signaling as well as defective TGF-ß signaling in T cells. Mice with defective signaling from both of these regulatory cytokines developed fulminant UC-like disease that was ameliorated by anti-cytokine therapy: anti-IFN - particularly in combination with anti-TNF-α alleviated most aspects of pathology. Furthermore, colitis could be completely prevented by oral broad-spectrum antibiotics, confirming an involvement of enteric bacteria in colitis development.
Genetics of Human IBD
The understanding of the genetics of human IBD is progressing with tremendous speed. NOD2 (CARD15) was identified as a susceptibility gene in 2001 [22,23]. In 2007, genome-wide association (GWA) studies were introduced [6]. This has led to the identification of a large number of new IBD genes, and additional genes are continuously being identified (see references within [7,8]). A recent meta-analysis reported on 32 CD genes and indicated that the actual number of CD genes is substantially higher [24]. With the exception of NOD2 (CARD15) and IL-23R, which each explain 1%–2% of the total genetic variance, the odd ratios of the other loci are low (generally <1.3), and their individual contribution to the genetic variance is meager. The 32 loci were altogether estimated to account for about 10% of the overall variance in disease risk, which may be as much as a fifth of the genetic risk [6,25]. Although no extensive GWA study has been performed on UC yet, it has become clear that some of the identified genetic factors are shared between CD and UC, but also that some of the genetic factors are specific for either CD or UC (Table 1) [25,26]. In particular, it is striking that NOD2 (CARD15), which is an intracellular sensor of bacterial peptidoglycan, and ATG16L1 and IRGM, which are involved in autophagy, are genetic factors for CD, but not for UC. This indicates that there are distinct pathogenic mechanisms related to microbial processing in CD and UC. The identification of IL-23R and IL-12B as risk factors in both CD and UC points to common inflammatory pathways of the two disorders, and underscores the observations of mouse models that inflammation involving Th17 cells is important.
Tab. 1. Genetic Associations in Crohn Disease and Ulcerative Colitis 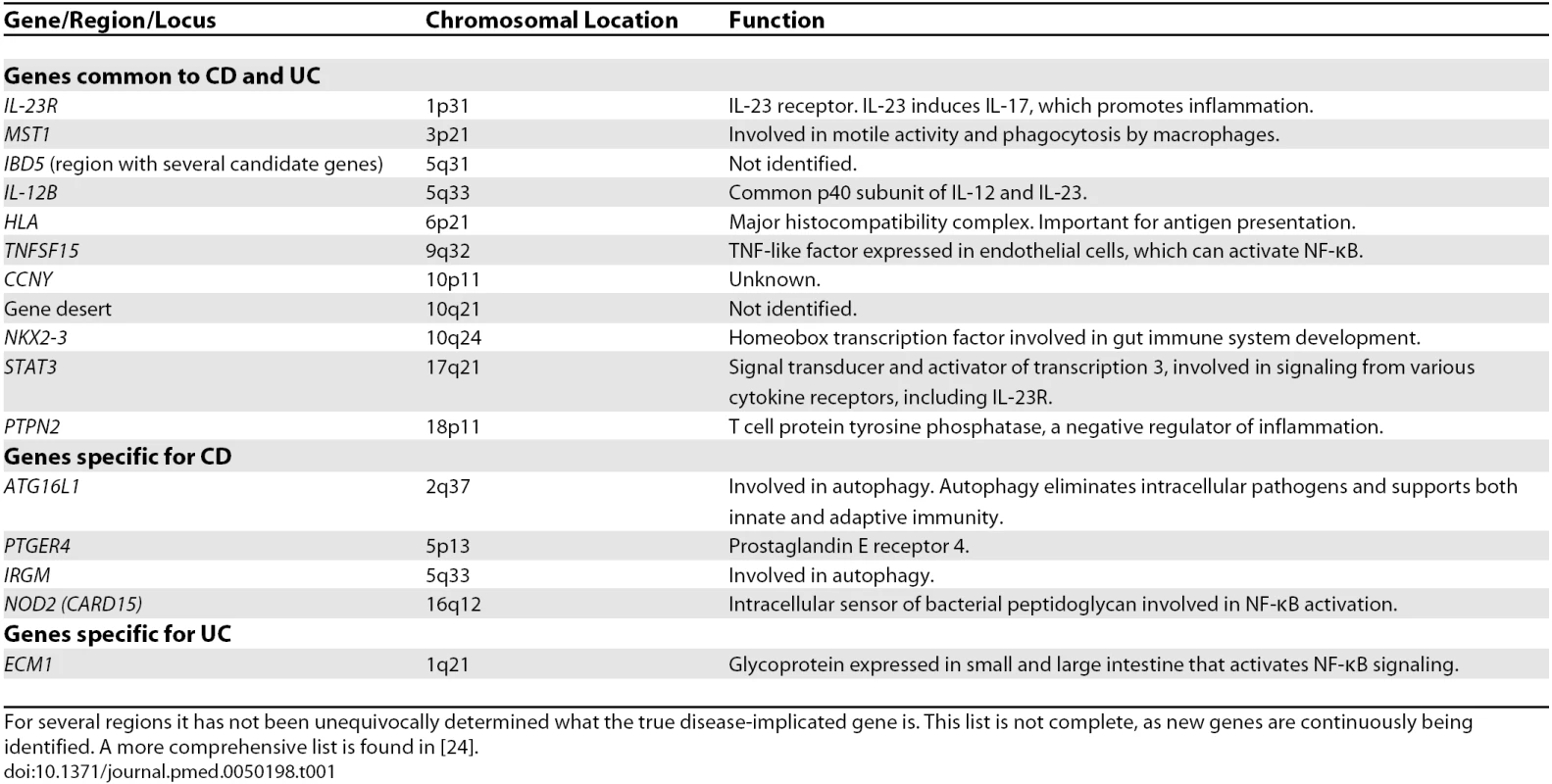
For several regions it has not been unequivocally determined what the true disease-implicated gene is. This list is not complete, as new genes are continuously being identified. A more comprehensive list is found in [24]. The new genetic findings have many implications. Most importantly, the identified susceptibility genes point at pathogenic pathways. These pathways also represent targets for therapy, which can either be the molecules encoded by the susceptibility genes themselves or other molecules involved in the same molecular pathways.
Despite the recent success, the genetic studies of humans have several shortcomings. Novel and strong genetic factors can still be discovered, as GWA studies do not effectively address rare polymorphisms that may have high penetrance or copy number variation. As for the regions that have been discovered, it is often difficult to pinpoint the responsible gene and even more difficult to pinpoint the causative mutation, due to linkage disequilibrium. Animal models have the potential to assist in solving these problems, even though the background genes will be different and there are likely differences between the animal and human gut physiology. The function of the different genes suggested by the genetic studies can be tested in in vivo settings, and the function of particular polymorphisms can be tested by gene knock-in or other types of genetic manipulation. Such models have been developed in mouse for studies of the NOD2 gene.
Animal Models in the New Genetics Era: Lessons from NOD2
Individuals with mutations in the NOD2 gene leading to altered NOD2 protein have increased risk for ileal-only and ileocolonic CD, but not colonic-only CD nor UC. This risk is particularly high for homozygous or compound-heterozygous individuals. NOD2 activates an NF-κB signaling pathway upon binding the peptidoglycan component muramyl dipeptide.
Two models of genetically manipulated mice have been developed to study the function of NOD2 in IBD [27,28]. In one model, NOD2 has been knocked out. In the absence of NOD2, TLR2 activation by peptidoglycan (or Pam3Cys) leads to excessive IL-12 production by macrophages, and thus a Th1 polarizing environment, suggesting that the human NOD2 polymorphisms associated with CD are loss-of-function mutations. This result also suggests that there is inhibitory cross talk between different bacterial sensors. Furthermore, NOD2 transfection into epithelial cells enhanced their resistance to bacterial infection, and its expression in Paneth cells in vivo is required for expression of a subset of defensins, suggesting that loss-of-function mutations reduce innate epithelial defense and therefore upset mucosal homeostasis.
An alternative explanation for NOD2 involvement in CD has come from knock-in animals with a mutation equivalent to the most common human CD-associated allele, 3020insC. Macrophages from these mice showed increased production of mature IL-1ß upon muramyl dipeptide stimulation, suggested to be caused by a gain-of-function mutation in NOD2 that activated caspase-1, an enzyme required for the activation step of IL-1ß. At the same time, macrophages from 3020insC knock-in mice retained normal TLR signaling. Although work with engineered mice strains have provided mechanistic understanding of how NOD2 might work, these studies have not settled the debate on the mechanism by which NOD2 variants in human IBD enhance susceptibility to CD. Neither mouse model developed spontaneous IBD, so chemical irritation or antigen-driven UC-like models have been used, even though NOD2 mutations are linked to CD and not UC in humans.
The Way Forward for Animal Models
The list of human IBD genes is continuously expanding. Some of the identified risk genes fit with pathways suggested by animal models, like the central role of Th17 cells. It will be interesting to see how many other genes will map to pathways that have been identified by animal models. Most of our current animal models have been developed without human genetics as a roadmap, but we can expect that this will change drastically. New models will be developed to understand the function of the novel IBD genes. The experience from studies of NOD2 in mice suggests that the development of adequate models will not be straightforward. Most of the novel gene polymorphisms are low-penetrance genes, and the development of suitable animal models for such genes can be particularly demanding. The intestinal physiology of the mouse and human gut are different, and the function of many genes can have species differences. Moreover, if there are gene–gene and gene–environment interactions for the risk genes, reestablishing the context into which a gene is predisposing to IBD will be difficult. A major environmental factor is the composition of the luminal bacterial community. Although there are differences between human and test animal microbiota, and the host–bug interactions likely will be different, the Human Microbiome Project (US), MetaHIT (Europe), and other big science projects aimed at determining composition of gut microflora in humans and test animals will surely be useful to IBD researchers. A further challenge will be to develop models that are representative for CD or UC or the major phenotypes within. Despite these challenges, good animal models are badly needed to bring new therapies to the clinic. Animal models of IBD should hence receive a continued interest, not least from clinicians, in the future.
Seven Key Papers in the Field
The Wellcome Trust Case Control Consortium, 2007 [6] A groundbreaking paper reporting GWA studies of seven diseases, including CD. Several new genes implicated in IBD were reported.
Rakoff-Nahoum et al., 2004 [17] This paper demonstrated recognition and activation of TLRs by commensal bacteria under normal steady-state conditions. Such activation had a crucial role in the protection against gut injury and associated mortality. Thus, commensal bacteria signal actively via TLRs to the host to promote mucosal homeostasis.
Hugot et al., 2001 [22] and Ogura et al., 2001 [23] These papers identified NOD2 (CARD15) as a susceptibility gene in CD. The papers lead to a paradigm shift directing the focus toward innate immunity and the role of dysregulated immune response to luminal bacteria in the pathogenesis of CD.
Hermiston and Gordon, 1995 [13] In this paper, a dominant negative N-cadherin expressed in the crypts of the small intestine caused a disruption of the epithelial barrier and led to ileal colitis resembling CD. Thus, a normal immune response to an abnormal stimulus could be an underlying cause of IBD.
Sadlack et al., 1993 [19] and Kuhn et al., 1993 [18] These papers knocked out the genes for IL-2 and IL-10, respectively, and showed that both strains of mice developed intestinal inflammation. Furthermore, both strains were cured of their IBD when rederived to germ-free conditions. Thus, an inappropriate immune response to a normal stimulus could be an underlying cause of IBD.
Zdroje
1. HalfvarsonJBodinLTyskCLindbergEJarnerotG
2003
Inflammatory bowel disease in a Swedish twin cohort: A long-term follow-up of concordance and clinical characteristics.
Gastroenterology
124
1767
1773
2. WirtzSNeurathMF
2007
Mouse models of inflammatory bowel disease.
Adv Drug Deliv Rev
59
1073
1083
3. SartorRB
2008
Microbial influences in inflammatory bowel diseases.
Gastroenterology
134
577
594
4. HeazlewoodCKCookMCEriRPriceGRTauroSB
2008
Aberrant mucin assembly in mice causes endoplasmic reticulum stress and spontaneous inflammation resembling ulcerative colitis.
PLoS Med
5
e54
doi:10.1371/journal.pmed.0050054
5. KangSSBloomSMNorianLAGeskeMJFlavellRA
2008
An antibiotic-responsive mouse model of fulminant ulcerative colitis.
PLoS Med
5
e41
doi:10.1371/journal.pmed.0050041
6. The Wellcome Trust Case Control Consortium
2007
Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls.
Nature
447
661
678
7. ChoJH
2008
The genetics and immunopathogenesis of inflammatory bowel disease.
Nat Rev Immunol
8
458
466
8. MathewCG
2008
New links to the pathogenesis of Crohn disease provided by genome-wide association scans.
Nat Rev Genet
9
9
14
9. LiMOFlavellRA
2008
Contextual regulation of inflammation: A duet by transforming growth factor-beta and interleukin-10.
Immunity
28
468
476
10. PowrieFLeachMWMauzeSMenonSCaddleLB
1994
Inhibition of Th1 responses prevents inflammatory bowel disease in scid mice reconstituted with CD45RBhi CD4+ T cells.
Immunity
1
553
562
11. SakaguchiSSakaguchiNAsanoMItohMTodaM
1995
Immunologic self-tolerance maintained by activated T cells expressing IL-2 receptor alpha-chains (CD25). Breakdown of a single mechanism of self-tolerance causes various autoimmune diseases.
J Immunol
155
1151
1164
12. McGuckinMAEriRSimmsLAFlorinTHRadford-SmithG
2008
Intestinal barrier dysfunction in inflammatory bowel diseases.
Inflamm Bowel Dis
E-pub 11 July 2008
13. HermistonMLGordonJI
1995
Inflammatory bowel disease and adenomas in mice expressing a dominant negative N-cadherin.
Science
270
1203
1207
14. Van der SluisMDe KoningBADe BruijnACVelcichAMeijerinkJP
2006
Muc2-deficient mice spontaneously develop colitis, indicating that MUC2 is critical for colonic protection.
Gastroenterology
131
117
129
15. NenciABeckerCWullaertAGareusRvan LooG
2007
Epithelial NEMO links innate immunity to chronic intestinal inflammation.
Nature
446
557
561
16. ZaphCTroyAETaylorBCBerman-BootyLDGuildKJ
2007
Epithelial-cell-intrinsic IKK-beta expression regulates intestinal immune homeostasis.
Nature
446
552
556
17. Rakoff-NahoumSPaglinoJEslami-VarzanehFEdbergSMedzhitovR
2004
Recognition of commensal microflora by toll-like receptors is required for intestinal homeostasis.
Cell
118
229
241
18. KuhnRLohlerJRennickDRajewskyKMullerW
1993
Interleukin-10-deficient mice develop chronic enterocolitis.
Cell
75
263
274
19. SadlackBMerzHSchorleHSchimplAFellerAC
1993
Ulcerative colitis-like disease in mice with a disrupted interleukin-2 gene.
Cell
75
253
261
20. GarrettWSLordGMPunitSLugo-VillarinoGMazmanianSK
2007
Communicable ulcerative colitis induced by T-bet deficiency in the innate immune system.
Cell
131
33
45
21. AnnunziatoFCosmiLSantarlasciVMaggiLLiottaF
2007
Phenotypic and functional features of human Th17 cells.
J Exp Med
204
1849
1861
22. HugotJPChamaillardMZoualiHLesageSCezardJP
2001
Association of NOD2 leucine-rich repeat variants with susceptibility to Crohn's disease.
Nature
411
599
603
23. OguraYBonenDKInoharaNNicolaeDLChenFF
2001
A frameshift mutation in NOD2 associated with susceptibility to Crohn's disease.
Nature
411
603
606
24. BarrettJCHansoulSNicolaeDLChoJHDuerrRH
2008
Genome-wide association defines more than 30 distinct susceptibility loci for Crohn's disease.
Nat Genet
40
955
962
25. FisherSATremellingMAndersonCAGwilliamRBumpsteadS
2008
Genetic determinants of ulcerative colitis include the ECM1 locus and five loci implicated in Crohn's disease.
Nat Genet
40
710
712
26. FrankeABalschunTKarlsenTHHedderichJMayS
2008
Replication of signals from recent studies of Crohn's disease identifies previously unknown disease loci for ulcerative colitis.
Nat Genet
40
713
715
27. EckmannLKarinM
2005
NOD2 and Crohn's disease: loss or gain of function.
Immunity
22
661
667
28. StroberWFussIMannonP
2007
The fundamental basis of inflammatory bowel disease.
J Clin Invest
117
514
521
Štítky
Interní lékařství
Článek vyšel v časopisePLOS Medicine
Nejčtenější tento týden
2008 Číslo 9- Alternativní léčebné možnosti u hypercholesterolemie při intoleranci statinů
- Vliv kombinace nutraceutik na remodelaci levé komory srdeční u osob s metabolickým syndromem
- Nutraceutika a jejich ovlivnění mírného kardiometabolického rizika
- Princip účinku medu v léčbě chronických i infikovaných ran
- Superoxidovaný roztok a jeho využití v léčbě ran
-
Všechny články tohoto čísla
- Towards a Data Sharing Culture: Recommendations for Leadership from Academic Health Centers
- A 50-Year-Old Woman with Recurrent Generalised Seizures
- Tobacco Control Yields Clear Dividends for Health and Wealth
- Ethical and Practical Issues Associated with Aggregating Databases
- Publication of Clinical Trials Supporting Successful New Drug Applications: A Literature Analysis
- Informed Consent in the Genomics Era
- Birth Size and the Pathogenesis of Breast Cancer
- Animal Models of Inflammatory Bowel Disease at the Dawn of the New Genetics Era
- Making Sense of Non-Financial Competing Interests
- PLOS Medicine
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Informed Consent in the Genomics Era
- Publication of Clinical Trials Supporting Successful New Drug Applications: A Literature Analysis
- A 50-Year-Old Woman with Recurrent Generalised Seizures
- Ethical and Practical Issues Associated with Aggregating Databases
Přihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



