-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praktické
Top novinky
ReklamaImprinted Genes … and the Number Is?
article has not abstract
Published in the journal: . PLoS Genet 8(3): e32767. doi:10.1371/journal.pgen.1002601
Category: Perspective
doi: https://doi.org/10.1371/journal.pgen.1002601Summary
article has not abstract
Genomic imprinting in mammals results in the expression of the alleles of a given gene being dependent on their parental origin. Although the existence of imprinted genes was postulated to explain aberrant development of uniparental embryos [1], it wasn't until 1991 that the first imprinted genes were identified by candidate approaches or fortuitously [2]. Given the serious developmental consequences of uniparental embryos, as well as some human syndromes associated with parental-specific deletion of particular chromosome regions, there has been great interest in discovering imprinted genes. As such, several unbiased approaches have been developed in the last 20 years with the goal of obtaining a complete list of imprinted genes. These approaches typically involved identifying genes that were present/absent in complete or partially uniparental embryos, although regions harbouring allele-specific DNA or chromatin modifications have also been used as an indicator of imprinted genes [3]. Earlier studies suggested that imprinted genes likely numbered in the low hundreds. Thus, it was startling to the imprinting community in 2010 when Gregg and colleagues reported >1,000 potential tissue-specific imprinted genes [4]. How could so many have been missed? In fact, others had previously used similar methodology but reported far fewer new imprinted genes [5], [6]. The answer, as discussed in a report from DeVeale and colleagues in this issue of PLoS Genetics [7], may not be that so many imprinted genes were missed, but that the limitations of the novel technology may not have been fully appreciated.
The experimental strategy that Gregg et al. and Babak and colleagues [4], [5] used to discover imprinted genes was to perform quantitative, whole-transcriptome sequencing (mRNA-seq) of samples from reciprocal hybrids (fetal or adult brain tissue from F1 hybrid mice, Figure 1) and to identify single nucleotide polymorphisms (SNPs) at which one parental allele is preferentially expressed. Comparison of reciprocal cross samples should rule out genetic effects and mitigate against some experimental noise. The approach is conceptually simple, but it requires robust statistical methods to account for false positives and it is probably fair to say that this remains an area of methodological development.
Fig. 1. Strategy for the generation and analysis of RNA from F1 hybrid mice. 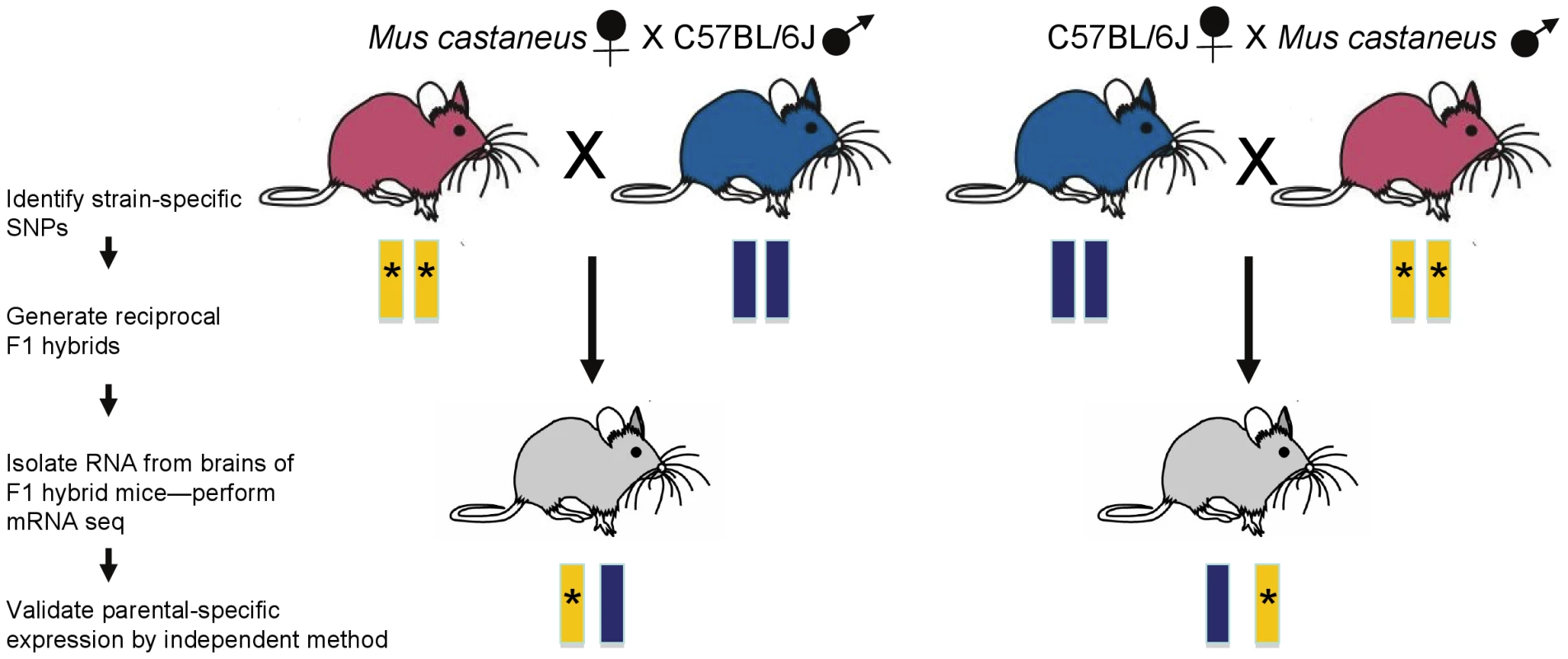
F1 hybrid progeny were generated from reciprocal matings between C57BL/6J and Mus musculus castaneus mice. Tissues were isolated from fetal and adult brain of the F1 hybrid mice and mRNA-seq was performed. Using SNPs (asterisks) that were identified in the parental DNA, biases in transcription of parental alleles can be assessed. By reanalysing mRNA-seq datasets from embryonic day 15 (e15) brain published by Gregg et al. [4] and e17.5 brain (their own, [5]), and using the same statistical approach, DeVeale et al. detect similar numbers of known imprinted genes. However, there was far less overlap in the new imprinted genes predicted from the two experiments: each predicted 400–500 candidates, but only about 50 were in common. Although these studies assayed fetal brain from different times, DeVeale and colleagues suspected that the discrepancy was more likely caused by technical issues in generation, mapping, or analysis of the mRNA-seq data.
A prerequisite in analysing large sequence datasets is to know how many candidates could appear “by chance” and to set thresholds to account for this. Although a false discovery rate (FDR) for a dataset can be predicted, there may be sources of experimental noise in the data that are not fully taken into account. Alternatively, it may be possible to determine an FDR empirically. DeVeale and colleagues did so by assuming that SNPs in the same coding exon of an imprinted transcript, but sufficiently distant to be sampled independently, should show the same parental allele expression bias; SNPs discordant in their direction of bias are more likely the consequence of sampling effects at the two positions. Of 1,388 SNP pairs, ∼20% disagreed on direction of bias, suggesting that as many as 40% of the predicted imprinted genes could be false positives. In a second approach, the authors analysed the number of candidates predicted in a “mock reciprocal” cross. This involves taking one F1 mRNA-seq dataset and comparing it with a second F1 dataset as if they were from reciprocal crosses. Worryingly, nearly as many candidate genes emerged from the mock reciprocal as a true reciprocal cross once known imprinted genes had been taken into account.
Using the FDRs determined from mock reciprocal crosses to set a threshold of significance, the authors then reanalysed reciprocal cross mRNA-seq datasets from four tissues: e15 and e17.5 whole brain, adult prefrontal cortex, and preoptic area [4], [5]. They detected 53 putative novel imprinted genes, including three that had already been validated by Gregg et al. Discounting 11 that were associated with known imprinted clusters, 42 candidates remained. They went on to verify a number of transcripts using an independent RT-PCR-based assay, including 17 candidates predicted by Gregg et al. (albeit of the “complex category”, in which there was discordance between parental allele ratios at different SNPs in the same transcript). Six of their 11 candidates validated with parental origin-specific allelic expression bias, but none of the “Gregg candidates” did. Not surprisingly, validation was best in genes with the highest “imprinting score” (a combination of allelic bias and read depth), including genes with biased parental allele expression in multiple samples and concordant at multiple SNPs. These criteria make sense, but such reasoning does not exclude the possibility that there may be additional imprinted genes among the longer candidate lists that exhibit spatiotemporally restricted imprinted expression.
To account for these discrepant findings, DeVeale and colleagues [7] argue that there are potentially multiple sources of systematic error in quantifying allele-specific expression by mRNA-seq, but whether these in aggregate could explain the substantially greater number of candidate imprinted transcripts reported by Gregg et al. is unclear. Nevertheless, the current study demonstrates the importance of appropriate empirically determined FDRs and extensive validation of new candidates by an independent method. Convergent evidence from other datasets, for example, parental-allele-specific DNA methylation or histone modifications, as they become available, will also be useful [8].
Transcriptome sequencing has also been applied to imprinted gene identification in the mouse placenta. The placenta is particularly significant in the physiology of imprinting, owing to its role in regulating fetal growth by controlling the supply of nutrients. In this case, an additional confounding factor is expression of genes in maternally derived cells within the placenta that can remain even after careful dissection [9]. Recently, Okae et al. elegantly demonstrated by embryo transfer that genes highly expressed in contaminating maternal decidual tissue, or other maternal cells infiltrating the placenta, can appear imprinted with maternal-allele-specific expression [10]. Using mRNA-seq of reciprocal hybrid placenta, they identified ∼1,000 genes expressed predominantly from the maternal allele. However, imprinted maternal allele expression was unequivocally demonstrated for only one of 269 genes they sought to verify (in some additional cases, genuine imprinted expression could have been obscured by extremely high expression in contaminating maternal tissue). The success rate for genes with paternal-allele-specific expression was much higher (1/6). This important study casts doubt on a number of apparent imprinted genes previously reported to exhibit maternal-allele-specific expression restricted to the placenta.
Where does this leave us? First, there are probably not more than the few hundred imprinted genes that were predicted many years ago. Moreover, with the rapid development of high-throughput transcriptome sequencing, we have an unprecedented opportunity to identify imprinted genes in any species with a sequenced genome. Thus, understanding the roles of imprinted genes in disease, development, and evolution will be within reach. Nevertheless, the studies by DeVeale and Okae suggest that results from high-throughput screens must be carefully interrogated, including substantial validation by alternative methods.
Zdroje
1. SolterD 1988 Differential imprinting and expression of maternal and paternal genomes. Annu Rev Genet 22 127 146
2. Ferguson-SmithAC 2011 Genomic imprinting: the emergence of an epigenetic paradigm. Nat Rev Genet 12 565 575
3. HenckelAArnaudP 2010 Genome-wide identification of new imprinted genes. Brief Funct Genomics 9 304 314
4. GreggCZhangJWeissbourdBLuoSSchrothGP 2010 High-resolution analysis of parent-of-origin allelic expression in the mouse brain. Science 329 643 648
5. BabakTDeVealeBArmourCRaymondCClearyMA 2008 Global survey of genomic imprinting by transcriptome sequencing. Curr Biol 18 1735 1741
6. WangXSunQMcGrathSDMardisERSolowayPD 2008 Transcriptome-wide identification of novel imprinted genes in neonatal mouse brain. PLoS ONE 3 e3839 doi:10.1371/journal.pone.0003839
7. DeVealeBvan der KooyDBabakT 2012 Critical evaluation of imprinted gene expression by RNA-Seq: a new perspective. PLoS Genet 8 e1002600 doi:10.1371/journal.pgen.1002600
8. XieWBarrCLKimAYueFLeeAYEubanksJ 2012 Base-resolution analyses of parent-of-origin and sequence dependent DNA methylation in the mouse genome. Cell 148 816 831
9. ProudhonCBourc'hisD 2010 Identification and resolution of artifacts in the interpretation of imprinted gene expression. Brief Funct Genomics 9 374 384
10. OkaeHHiuraHNishidaYFunayamaRTanakaS 2012 Re-investigation and RNA sequencing-based identification of genes with placenta-specific imprinted expression. Hum Mol Genet 21 548 558
Štítky
Genetika Reprodukční medicína
Článek Physiological Notch Signaling Maintains Bone Homeostasis via RBPjk and Hey Upstream of NFATc1Článek Intronic -Regulatory Modules Mediate Tissue-Specific and Microbial Control of / TranscriptionČlánek Probing the Informational and Regulatory Plasticity of a Transcription Factor DNA–Binding DomainČlánek Repression of Germline RNAi Pathways in Somatic Cells by Retinoblastoma Pathway Chromatin ComplexesČlánek An Alu Element–Associated Hypermethylation Variant of the Gene Is Associated with Childhood ObesityČlánek Three Essential Ribonucleases—RNase Y, J1, and III—Control the Abundance of a Majority of mRNAsČlánek Genomic Tools for Evolution and Conservation in the Chimpanzee: Is a Genetically Distinct Population
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2012 Číslo 3
-
Všechny články tohoto čísla
- Comprehensive Research Synopsis and Systematic Meta-Analyses in Parkinson's Disease Genetics: The PDGene Database
- Genomic Analysis of the Hydrocarbon-Producing, Cellulolytic, Endophytic Fungus
- Networks of Neuronal Genes Affected by Common and Rare Variants in Autism Spectrum Disorders
- Akirin Links Twist-Regulated Transcription with the Brahma Chromatin Remodeling Complex during Embryogenesis
- Too Much Cleavage of Cyclin E Promotes Breast Tumorigenesis
- Imprinted Genes … and the Number Is?
- Genetic Architecture of Highly Complex Chemical Resistance Traits across Four Yeast Strains
- Exploring the Complexity of the HIV-1 Fitness Landscape
- MNS1 Is Essential for Spermiogenesis and Motile Ciliary Functions in Mice
- A Fundamental Regulatory Mechanism Operating through OmpR and DNA Topology Controls Expression of Pathogenicity Islands SPI-1 and SPI-2
- Evidence for Positive Selection on a Number of MicroRNA Regulatory Interactions during Recent Human Evolution
- Variation in Modifies Risk of Neonatal Intestinal Obstruction in Cystic Fibrosis
- PIF4–Mediated Activation of Expression Integrates Temperature into the Auxin Pathway in Regulating Hypocotyl Growth
- Critical Evaluation of Imprinted Gene Expression by RNA–Seq: A New Perspective
- A Meta-Analysis and Genome-Wide Association Study of Platelet Count and Mean Platelet Volume in African Americans
- Mouse Genetics Suggests Cell-Context Dependency for Myc-Regulated Metabolic Enzymes during Tumorigenesis
- Transcriptional Control in Cardiac Progenitors: Tbx1 Interacts with the BAF Chromatin Remodeling Complex and Regulates
- Synthetic Lethality of Cohesins with PARPs and Replication Fork Mediators
- APOBEC3G-Induced Hypermutation of Human Immunodeficiency Virus Type-1 Is Typically a Discrete “All or Nothing” Phenomenon
- Interpreting Meta-Analyses of Genome-Wide Association Studies
- Error-Prone ZW Pairing and No Evidence for Meiotic Sex Chromosome Inactivation in the Chicken Germ Line
- -Dependent Chemosensory Functions Contribute to Courtship Behavior in
- Diverse Forms of Splicing Are Part of an Evolving Autoregulatory Circuit
- Phenotypic Plasticity of the Drosophila Transcriptome
- Physiological Notch Signaling Maintains Bone Homeostasis via RBPjk and Hey Upstream of NFATc1
- Precocious Metamorphosis in the Juvenile Hormone–Deficient Mutant of the Silkworm,
- Igf1r Signaling Is Indispensable for Preimplantation Development and Is Activated via a Novel Function of E-cadherin
- Accurate Prediction of Inducible Transcription Factor Binding Intensities In Vivo
- Mitochondrial Oxidative Stress Alters a Pathway in Strongly Resembling That of Bile Acid Biosynthesis and Secretion in Vertebrates
- Mammalian Neurogenesis Requires Treacle-Plk1 for Precise Control of Spindle Orientation, Mitotic Progression, and Maintenance of Neural Progenitor Cells
- Tcf7 Is an Important Regulator of the Switch of Self-Renewal and Differentiation in a Multipotential Hematopoietic Cell Line
- REST–Mediated Recruitment of Polycomb Repressor Complexes in Mammalian Cells
- Intronic -Regulatory Modules Mediate Tissue-Specific and Microbial Control of / Transcription
- Age-Dependent Brain Gene Expression and Copy Number Anomalies in Autism Suggest Distinct Pathological Processes at Young Versus Mature Ages
- A Genome-Wide Association Study Identifies Variants Underlying the Shade Avoidance Response
- -by- Regulatory Divergence Causes the Asymmetric Lethal Effects of an Ancestral Hybrid Incompatibility Gene
- Genome-Wide Association and Functional Follow-Up Reveals New Loci for Kidney Function
- A Natural System of Chromosome Transfer in
- Cell Size and the Initiation of DNA Replication in Bacteria
- Probing the Informational and Regulatory Plasticity of a Transcription Factor DNA–Binding Domain
- Repression of Germline RNAi Pathways in Somatic Cells by Retinoblastoma Pathway Chromatin Complexes
- Temporal Transcriptional Profiling of Somatic and Germ Cells Reveals Biased Lineage Priming of Sexual Fate in the Fetal Mouse Gonad
- Rapid Analysis of Genome Rearrangements by Multiplex Ligation–Dependent Probe Amplification
- Metabolic Profiling of a Mapping Population Exposes New Insights in the Regulation of Seed Metabolism and Seed, Fruit, and Plant Relations
- The Atypical Calpains: Evolutionary Analyses and Roles in Cellular Degeneration
- The Silkworm Coming of Age—Early
- Development of a Panel of Genome-Wide Ancestry Informative Markers to Study Admixture Throughout the Americas
- Balanced Codon Usage Optimizes Eukaryotic Translational Efficiency
- The Min System and Nucleoid Occlusion Are Not Required for Identifying the Division Site in but Ensure Its Efficient Utilization
- Neurobeachin, a Regulator of Synaptic Protein Targeting, Is Associated with Body Fat Mass and Feeding Behavior in Mice and Body-Mass Index in Humans
- Statistical Analysis of Readthrough Levels for Nonsense Mutations in Mammalian Cells Reveals a Major Determinant of Response to Gentamicin
- Gene Reactivation by 5-Aza-2′-Deoxycytidine–Induced Demethylation Requires SRCAP–Mediated H2A.Z Insertion to Establish Nucleosome Depleted Regions
- The miR-35-41 Family of MicroRNAs Regulates RNAi Sensitivity in
- Genetic Basis of Hidden Phenotypic Variation Revealed by Increased Translational Readthrough in Yeast
- An Alu Element–Associated Hypermethylation Variant of the Gene Is Associated with Childhood Obesity
- Modelling Human Regulatory Variation in Mouse: Finding the Function in Genome-Wide Association Studies and Whole-Genome Sequencing
- Novel Loci for Adiponectin Levels and Their Influence on Type 2 Diabetes and Metabolic Traits: A Multi-Ethnic Meta-Analysis of 45,891 Individuals
- Polycomb-Like 3 Promotes Polycomb Repressive Complex 2 Binding to CpG Islands and Embryonic Stem Cell Self-Renewal
- Insulin/IGF-1 and Hypoxia Signaling Act in Concert to Regulate Iron Homeostasis in
- EMF1 and PRC2 Cooperate to Repress Key Regulators of Arabidopsis Development
- Three Essential Ribonucleases—RNase Y, J1, and III—Control the Abundance of a Majority of mRNAs
- Contrasted Patterns of Molecular Evolution in Dominant and Recessive Self-Incompatibility Haplotypes in
- A Machine Learning Approach for Identifying Novel Cell Type–Specific Transcriptional Regulators of Myogenesis
- Genomic Tools for Evolution and Conservation in the Chimpanzee: Is a Genetically Distinct Population
- Nos2 Inactivation Promotes the Development of Medulloblastoma in Mice by Deregulation of Gap43–Dependent Granule Cell Precursor Migration
- Intracranial Aneurysm Risk Locus 5q23.2 Is Associated with Elevated Systolic Blood Pressure
- Heritability and Genetic Correlations Explained by Common SNPs for Metabolic Syndrome Traits
- A Genome-Wide Association Study of Nephrolithiasis in the Japanese Population Identifies Novel Susceptible Loci at 5q35.3, 7p14.3, and 13q14.1
- DNA Damage in Nijmegen Breakage Syndrome Cells Leads to PARP Hyperactivation and Increased Oxidative Stress
- DNA Resection at Chromosome Breaks Promotes Genome Stability by Constraining Non-Allelic Homologous Recombination
- Genetic Analysis of Floral Symmetry in Van Gogh's Sunflowers Reveals Independent Recruitment of Genes in the Asteraceae
- A Splice Site Variant in the Bovine Gene Compromises Growth and Regulation of the Inflammatory Response
- Promoter Nucleosome Organization Shapes the Evolution of Gene Expression
- The Nucleoside Diphosphate Kinase Gene Acts as Quantitative Trait Locus Promoting Non-Mendelian Inheritance
- The Ciliogenic Transcription Factor RFX3 Regulates Early Midline Distribution of Guidepost Neurons Required for Corpus Callosum Development
- Phosphorylation of the RNA–Binding Protein HOW by MAPK/ERK Enhances Its Dimerization and Activity
- A Genome-Wide Scan of Ashkenazi Jewish Crohn's Disease Suggests Novel Susceptibility Loci
- Parkinson's Disease–Associated Kinase PINK1 Regulates Miro Protein Level and Axonal Transport of Mitochondria
- LMW-E/CDK2 Deregulates Acinar Morphogenesis, Induces Tumorigenesis, and Associates with the Activated b-Raf-ERK1/2-mTOR Pathway in Breast Cancer Patients
- Mapping the Hsp90 Genetic Interaction Network in Reveals Environmental Contingency and Rewired Circuitry
- Autoregulation of the Noncoding RNA Gene
- The Human Pancreatic Islet Transcriptome: Expression of Candidate Genes for Type 1 Diabetes and the Impact of Pro-Inflammatory Cytokines
- Spo0A∼P Imposes a Temporal Gate for the Bimodal Expression of Competence in
- Antagonistic Regulation of Apoptosis and Differentiation by the Cut Transcription Factor Represents a Tumor-Suppressing Mechanism in
- A Downstream CpG Island Controls Transcript Initiation and Elongation and the Methylation State of the Imprinted Macro ncRNA Promoter
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- PIF4–Mediated Activation of Expression Integrates Temperature into the Auxin Pathway in Regulating Hypocotyl Growth
- Metabolic Profiling of a Mapping Population Exposes New Insights in the Regulation of Seed Metabolism and Seed, Fruit, and Plant Relations
- A Splice Site Variant in the Bovine Gene Compromises Growth and Regulation of the Inflammatory Response
- Comprehensive Research Synopsis and Systematic Meta-Analyses in Parkinson's Disease Genetics: The PDGene Database
Přihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



