-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaInterpreting Meta-Analyses of Genome-Wide Association Studies
Meta-analysis is an increasingly popular tool for combining multiple genome-wide association studies in a single analysis to identify associations with small effect sizes. The effect sizes between studies in a meta-analysis may differ and these differences, or heterogeneity, can be caused by many factors. If heterogeneity is observed in the results of a meta-analysis, interpreting the cause of heterogeneity is important because the correct interpretation can lead to a better understanding of the disease and a more effective design of a replication study. However, interpreting heterogeneous results is difficult. The standard approach of examining the association p-values of the studies does not effectively predict if the effect exists in each study. In this paper, we propose a framework facilitating the interpretation of the results of a meta-analysis. Our framework is based on a new statistic representing the posterior probability that the effect exists in each study, which is estimated utilizing cross-study information. Simulations and application to the real data show that our framework can effectively segregate the studies predicted to have an effect, the studies predicted to not have an effect, and the ambiguous studies that are underpowered. In addition to helping interpretation, the new framework also allows us to develop a new association testing procedure taking into account the existence of effect.
Published in the journal: . PLoS Genet 8(3): e32767. doi:10.1371/journal.pgen.1002555
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1002555Summary
Meta-analysis is an increasingly popular tool for combining multiple genome-wide association studies in a single analysis to identify associations with small effect sizes. The effect sizes between studies in a meta-analysis may differ and these differences, or heterogeneity, can be caused by many factors. If heterogeneity is observed in the results of a meta-analysis, interpreting the cause of heterogeneity is important because the correct interpretation can lead to a better understanding of the disease and a more effective design of a replication study. However, interpreting heterogeneous results is difficult. The standard approach of examining the association p-values of the studies does not effectively predict if the effect exists in each study. In this paper, we propose a framework facilitating the interpretation of the results of a meta-analysis. Our framework is based on a new statistic representing the posterior probability that the effect exists in each study, which is estimated utilizing cross-study information. Simulations and application to the real data show that our framework can effectively segregate the studies predicted to have an effect, the studies predicted to not have an effect, and the ambiguous studies that are underpowered. In addition to helping interpretation, the new framework also allows us to develop a new association testing procedure taking into account the existence of effect.
Introduction
Meta-analysis is a tool for aggregating information from multiple independent studies [1]–[3]. In genome-wide association studies (GWASs) [4], the use of meta-analysis is becoming more and more popular because one can virtually collect tens of thousands of individuals that will provide power to identify associated variants with small effect sizes [5]–[7]. Several large scale meta-analyses have been performed for diseases including type 1 diabetes [8], type 2 diabetes [9]–[11], bipolar disorder [12], Crohns disease [13], and rheumatoid arthritis [14], and have identified associations not revealed in the individual studies.
In meta-analyses, the effect size between studies may differ and this difference, or heterogeneity, can be caused by many factors [15]–[18]. If the populations are different between studies, the genetic factors can cause heterogeneity [19], [20]. If the subjects are from different regions, the environmental factors can cause heterogeneity [21]. Even if the true effect size is invariant, the design factors can also cause a phenomenon that looks like heterogeneity, what is often called the statistical heterogeneity [22]. If the linkage disequilibrium structures are different between studies, the collected marker can show heterogeneity [23]. If the studies use different genotyping platforms, different imputation accuracies and different genotyping errors can cause heterogeneity [24].
In current meta-analyses of genome-wide association studies, heterogeneity is often observed in the results [9]–[11], [13], [17]. Interpreting the cause of such heterogeneity is important. If the heterogeneity is caused by either genetic or environmental factors, understanding the cause of heterogeneity can help our understanding of the disease mechanism. If the heterogeneity is statistical heterogeneity caused by the design factors, understanding the cause of heterogeneity is crucial in designing a replication study so that we can eliminate the design factors that can hinder the revelation of the true effect in the replication study.
However, interpreting heterogeneous results is difficult. One standard approach is to examine the association p-values of the studies. The inherent limitation of this approach is that a non-significant p-value is not evidence of the absence of an effect. Thus, a p-value does not provide the full information necessary for the interpretation whether or not there is an effect in the study. Another standard approach is to plot observed effect sizes and their confidence intervals of all studies [17], [25], [26]. This plot can be overly complicated when the number of studies is large and does not provide a single estimate that represents the existence of an effect in each study. The limitation of both approaches is that they use classical estimates that are calculated using only the data of each single study. That is, they utilize only within-study information.
In this paper, we propose a framework facilitating the interpretation of the results of a meta-analysis. Our framework is based on a new statistic termed the m-value which is the posterior probability that the effect exists in each study. Plotting the new statistic together with p-values in a two-dimensional space helps us distinguish between the studies predicted to have an effect, the studies predicted to not have an effect, and the ambiguous studies that are underpowered. We name this plot a P-M plot. In this framework, the outlier studies showing distinct characteristics from the other studies are easily identified, as we demonstrate using data from type 2 diabetes and Crohns disease meta-analyses [10], [13]. Our new statistic is fundamentally different from traditional estimates based on the data of single studies. We use all studies simultaneously to calculate the new statistic based on the assumption that the effect sizes are similar if the effect exists. Thus, we utilize cross-study information as well as within-study information.
In addition to helping interpretation, the new framework allows us to develop a new association testing procedure which takes into account the presence or absence of the effect. The new method called the binary effects model is a weighted sum of z-scores method [5] assigning a greater weight to the studies predicted to have an effect and a smaller weight to the studies predicted to not have an effect. Application to the Crohns disease data [13] shows that the new method gives more significant p-values than previous methods at certain loci already identified as associated.
The new method is available at http://genetics.cs.ucla.edu/meta.
Methods
Binary Effects Assumption
In our framework, we use a simplified model to describe heterogeneity among the studies which makes two assumptions. The first assumption is that effect is either present or absent in the studies. This assumption is different from the traditional assumption assuming normally distributed effect sizes [27]–[29]. Our assumption is inspired by the phenomenon that the effect sizes are sometimes observed to be much smaller in some studies than in the others. It is reported that different populations can cause such phenomenon [19], [20], [30], [31]. For example, the homozygosity for APOE 4 variant is known to confer fivefold smaller risk of Alzheimer disease in African Americans than in Asians [19], [30]. The HapK haplotype spanning the LTA4H gene is shown to confer threefold smaller risk of myocardial infraction in the populations of Europeans decent than in African Americans [31]. The HNF4A P2 promoter variants are shown to be associated with type 2 diabetes in Ashkenazi and the results have been replicated [20]. However, in the same study, the same variants did not show associations in four different cohorts of UK population suggesting a heterogeneous effect. Gene-environmental interactions can also cause such phenomenon. If a study lacks an environmental factor necessary for the interaction, the observed effect size can be much smaller in that study. It is generally agreed that the gene-environmental interactions exist in many diseases such as cardio vascular diseases [32], respiratory diseases [33], and mental disorders [34].
The second assumption is that if the effect exists, the effect sizes are similar between studies. We call these two assumptions together the binary effects assumption. While other types of heterogeneity structures are possible such as arbitrary effect sizes, for identifying which studies have an effect and which studies do not have an effect, we expect that this model will be appropriate.
M-Value
We propose a statistic called the m-value which is the posterior probability that the effect exists in each study of a meta-analysis. Suppose that we analyze studies together in a meta-analysis. Let () be the observed effect size of study and let be the estimated variance of . It is a common practice to consider the true variance. In the current GWASs, the distribution of is well approximated by a normal distribution due to the large sample sizes. Let denote the observed data.
If there is no effect in study ,where is the probability density function of a normal distribution whose mean is and the variance is . If there is effect in study ,where is the unknown true effect size.
Since we want a posterior probability, the Bayesian framework is a good fit. We assume that the prior for the effect size isA possible choice for in GWASs is 0.2 for small effect and 0.4 for large effect [35], [36].
Let be a random variable which has a value 1 if study has an effect and a value 0 if study does not have an effect. Let be the prior probability that each study will have an effect such thatThen we assume a beta prior on Through this paper, we use the uniform distribution prior ( and ), but other priors can also be chosen.
Let be the vector indicating the existence of effect in all studies. can have different values. Let be the set of those values.
Our goal is to estimate the m-value , the posterior probability that the effect exists in study . By the Bayes' theorem,(1)where is a subset of whose elements' th value is 1. Thus, we only need to know for each the posterior probability of ,consisting of the probability of given and the prior probability of .
The prior probability of iswhere is the number of 1's in and is the beta function.
And the probability of given is(2)where is the indices of 0 in and is the indices of 1 in . We can analytically work on the integration to obtainwherewhere is the inverse variance or precision. The summations are all with respect to .
is a scaling factor such thatThe details of the derivation is in Text S1 in Supporting Information S1. As a result, we can calculate for every and therefore obtain for each study .
MCMC
The drawback of the exact calculation of m-value is that we need to iterate over all which is exponential to . This is not problematic in most of the current meta-analyses of GWASs, but will be problematic in future studies if increases over several tens. Therefore, here we propose a simple Markov Chain Monte Carlo (MCMC) method to estimate m-value.
We propose the following Metropolis-Hastings algorithm [37].
-
Start from a random .
-
Choose a next .
-
If , move to . Otherwise, move to with probability .
-
Repeat from step 2.
The set of moves we use for choosing is . is a simple flipping move of between 0 and 1. is a move that shuffles the values of . This move is introduced to avoid being stuck on one mode in a special case that there are two modes which can happen when the observed direction of the effect is opposite in some studies. At each step, we randomly choose a move from this set assuming a uniform distribution. We allow burn-in and sample times. After sampling, samples gives us an approximation of the distribution over , which subsequently gives the approximations of m-values by the formula (1).
Interpretations and predictions
The m-value has a valid probabilistic interpretation that it is the posterior probability that the effect exists in each study under our binary effects model. If we are to choose studies predicted to have an effect and studies predicted to not have an effect, a threshold is needed. In this paper, we use the threshold of m-value for the former and m-value for the latter. Although this thresholding is arbitrary, the actual level of threshold is often not of importance because outlier studies showing different characteristics from the other studies usually stand out in the plotting framework described below.
Relationship to PPA
The m-value is closely related to the posterior probability of association (PPA) based on the Bayes factor (BF) [35] in the sense that the presence and absence of effects are essentially describing the same things as the alternative and null models in the association testing. There are two fundamental differences. First, in the usual PPA, the prior probability of association () is given by a point prior which is usually a very small value in GWAS reflecting the fact that the true associations are few. In our framework, we focus on interpreting meta-analysis results after we find associations using meta-analysis. Thus, reflects our belief on the effect conditioned on that the associations are already significant. For this reason, we need not use a very small value but instead choose to use a distribution prior. Second, the PPA is calculated for each study separately. However, the m-value is calculated using all studies simultaneously utilizing cross-study information. Thus, if the binary effects assumption approximates the truth, the m-value is more effective in predicting effects than the PPA or equivalently the BF, as we show by simulations in Results.
P-M Plot
We propose plotting the studies' p-values and m-values together in two dimensions. This plot, which we call the P-M plot, can help interpreting the results of a meta-analysis. Figure 1 shows that how to interpret such a plot. The right-most (pink) region is where the studies are predicted to have an effect. Often, a study can be in this region even if the p-value is not very significant. The left-most (light-blue) region is where the studies are predicted to not have an effect. This suggests that the sample size is large but the observed effect size is close to zero, suggesting a possibility that there exists no effect in that study. The middle (green) region is where the prediction is ambiguous. A study can be in this region because the study is underpowered due to a small sample size. If the sample size increases, the study will be drawn to either the left or the right side.
Fig. 1. A figure depicting the interpretations based on a P-M plot. 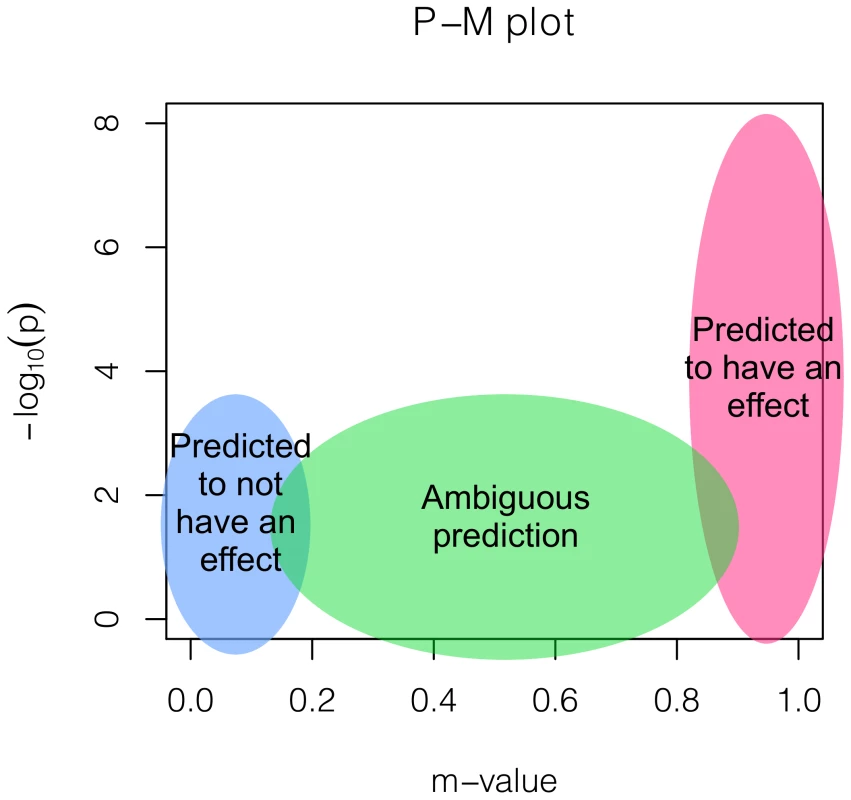
If the binary effects assumption does not hold, a study can sit in an unexpected region and a careful interpretation is necessary. For example, if the effects are significant but the effect sizes are in opposite direction in some studies, the studies can sit in the unusual top left region. However, such case will be rare and may be a result of the strand errors.
Binary Effects Model
We propose a new type of random effects model meta-analysis approach called the binary effects model. If the binary effects assumption holds, that is, if the effect is either present or absent in the studies, taking into account this pattern of heterogeneity in the association testing procedure can increase power compared to the general RE approach [23]. The binary effects model we propose is the weighted sum of z-scores method [5] where the m-values are incorporated into the weights. Intuitively, this is equivalent to assigning a greater weight to the studies predicted to have an effect and a smaller weight to the studies predicted to not have an effect.
Let be the z-score of study . The common form of the weighted sum of z-scores statistic for the fixed effects model isIn many cases, the weight approximates to the form where is the sample size and is the minor allele frequency [23]. When the minor allele frequency is similar between studies, the weight approximates to the popular form of [5].
The binary effects model statistic we propose isOur method is an empirical approach that uses estimated from the data as the prior weight for each study. Since the m-value is estimated using all studies, our approach can be thought of as gathering information from all studies and distributing back to each study in the form of weight. We choose this approach because of its simple formulation.
Since is not independent of , the statistic does not follow a normal distribution. Thus, the p-value is obtained using sampling which can be inefficient. We use two ideas to expedite the sampling. First, we propose an importance sampling procedure which is more efficient than the standard sampling. Second, we use an efficient approximation of m-value. See Text S2 and S3 in Supporting Information S1 for details.
Simulation Framework
In order to evaluate our methods, we use the following simulation approach. Assuming a minor allele frequency, a relative risk, and the number of individuals of cases and controls, a straightforward simulation approach is to sample alleles for cases and alleles for controls according to the expected minor allele frequencies in the cases and controls respectively [38]. However, since we perform extensive simulations assuming thousands of individuals, we use an approximation approach that samples the minor allele count from a normal distribution and rounds it to the nearest non-negative integer.
Web Resources
The URL for methods presented herein is as follows:
http://genetics.cs.ucla.edu/meta
Results
Simulation of M-Values
We demonstrate a simple simulation example showing how m-value behaves depending on the presence and absence of the effect and the sample size. First, we make the following assumptions throughout all of the experiments in this paper. We assume that the minor allele frequency of the collected marker is 0.3. We assume that the equal number of cases and controls are collected and refers to the total number of individuals as sample size . We also assume a very small disease prevalence when we calculate the expected minor allele frequencies for cases and controls given a relative risk . For the details how the expected values are calculated, see Han and Eskin [23]. Note that these assumptions are not critical factors affecting our simulation results. In all experiments, the random effects model (RE) denotes the RE method of Han and Eskin [23]. We omit the results of the conventional RE method [15] because they are highly conservative [23]. Throughout this paper, we use the following priors for calculating m-values. We use for the prior of the effect size (). We use the uniform distribution prior, , for the prior of the existence of effect ().
In this simulation example, we assume four different types of studies. The first type is a large study having an effect ( and ). The second type is a small study having an effect ( and ). The third type is a large study not having an effect ( and ). The fourth type is a small study not having an effect ( and ). We generate two studies per each type, constructing a simulated meta-analysis set of total eight studies. We accept this simulation set only if none of eight studies' p-values exceeds the genome-wide threshold () but the meta-analysis p-value calculated by the RE approach exceeds the genome-wide threshold. Otherwise, we repeat. We construct 1,000 meta-analysis sets.
Given this simulated data, we plot the histogram of m-values for each type of studies separately in Figure 2. Figure 2A shows that almost all (99.9%) of large studies with an effect are concentrated on large m-values (), showing that the m-values effectively predict that the effect exists in the studies. Figure 2C shows that a large amount (78.6%) of large studies without an effect are concentrated on small m-values (). Figure 2B and 2D show that when the sample size is small, m-value tends to the mid-range regardless of the effect, suggesting that the studies are underpowered to determine the presence of an effect.
Fig. 2. Histograms of m-values of different types of studies. 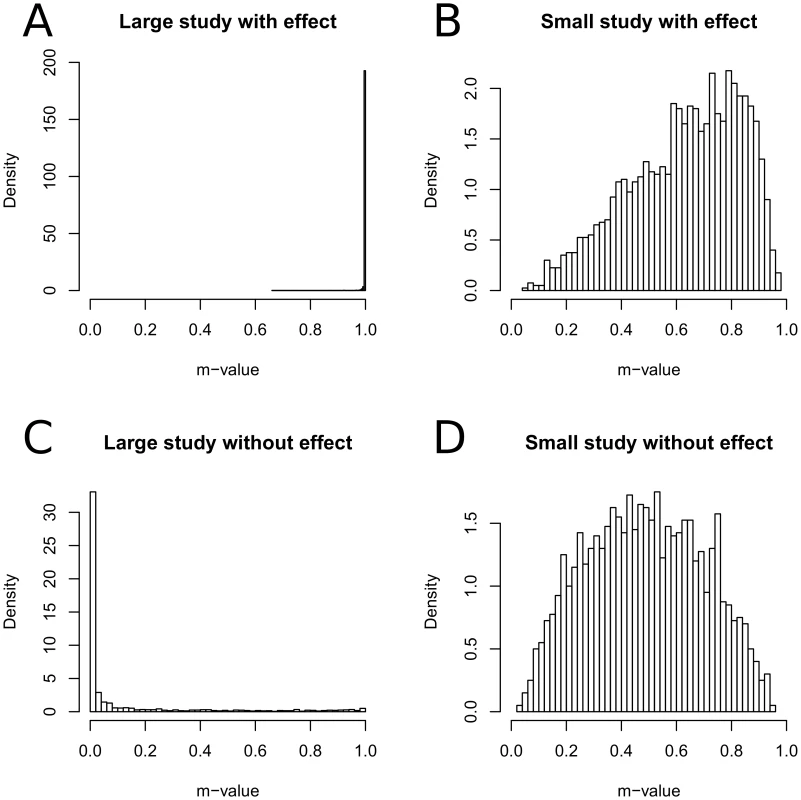
We assume four types of studies which is the all four combinations of large sample () and small sample (), and effect () and no effect (). We repeatedly simulate a meta-analysis of eight studies, two studies per each type, and calculate the m-values of the studies. Comparison of P-Value, M-Value, and BF
In this experiment, we compare the p-value, m-value, and BF by measuring how well they predict which studies have an effect and which studies do not have an effect. We assume a meta-analysis of 10 studies where the effect is either present () or not. We randomly pick the number of studies having an effect () from a uniform distribution ranging from 1 to 9, and randomly decide which studies have an effect. We randomly pick the sample size of each study from a uniform distribution between 500 and 2,000. Given the sample sizes and the effect sizes, we generate a meta-analysis study set. We accept the meta-analysis set only if none of the studies' p-values exceeds the genome-wide threshold () and the meta-analysis p-value exceeds the genome-wide threshold. We repeat until we construct 1,000 meta-analysis sets.
We examine each of 10,000 studies included in the simulated 1,000 meta-analysis sets. For each study, we calculate the p-value, m-value, and BF. We use the asymptotic BF of Wakefield [39] assuming the same prior distribution about the effect size as the m-value. Then we evaluate the performance of each statistic as follows. To evaluate the performance of m-value, we fix an arbitrary threshold so that we predict the studies having m-value to have an effect. Since we know the underlying truth if the effect exists or not in each study, we can measure what proportion of the studies actually having an effect is correctly predicted to have an effect (true prediction rate) and what proportion of the studies actually not having an effect is incorrectly predicted to have an effect (false prediction rate). Then we change the threshold to draw a curve between the true prediction rate and the false prediction rate, which is often called the receiver-operating-characteristic (ROC) curve. We do the same analysis for p-value and BF.
Figure 3A shows that m-value is superior to p-value and BF in predicting the studies having an effect. This is because m-value can utilize the cross-study information when the binary effects assumption holds. The performances of p-value and BF are almost identical.
Fig. 3. Comparison of prediction accuracies of p-value, m-value, and BF. 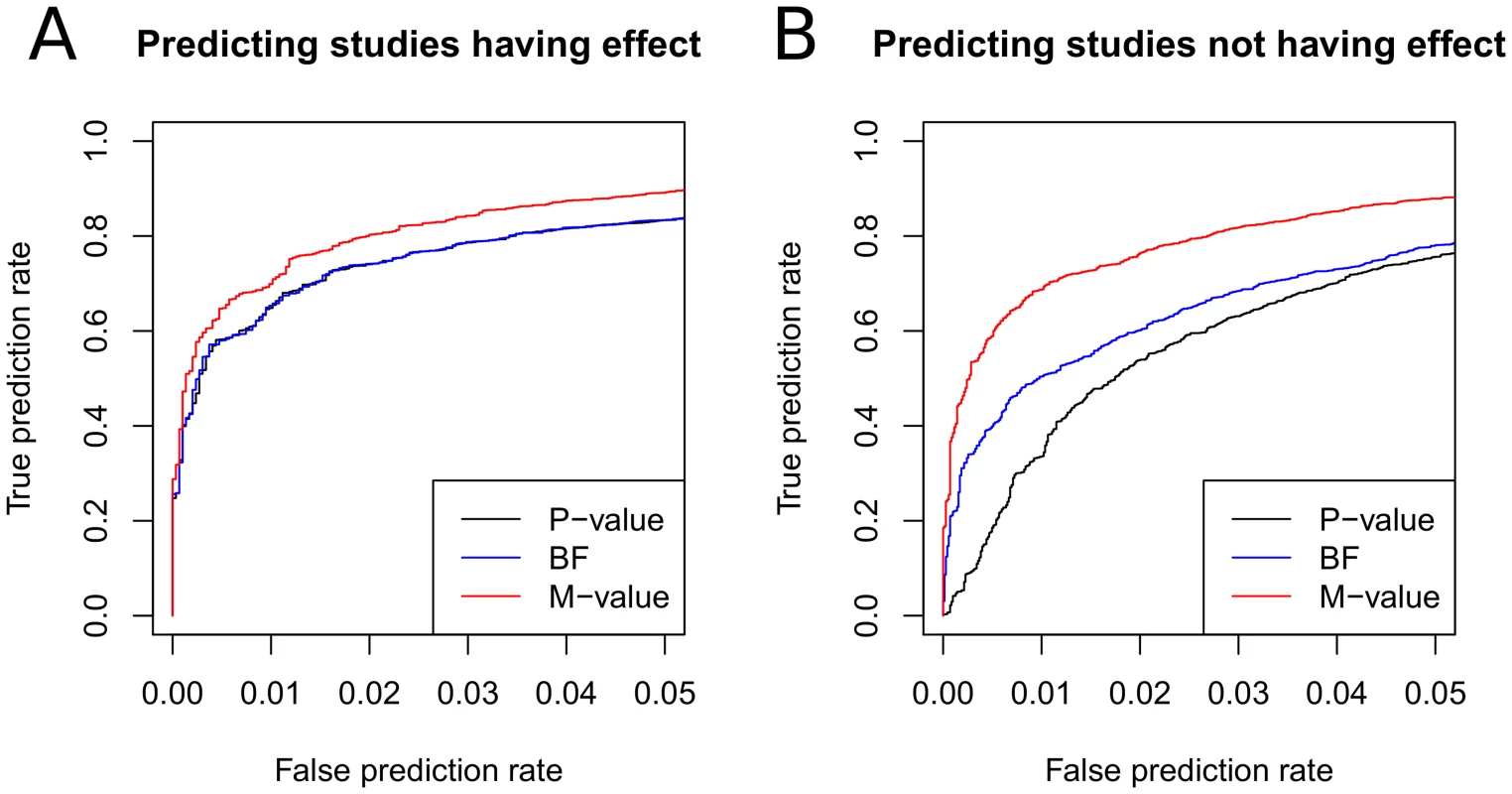
We simulate 1,000 meta-analysis of 10 studies with varying sample sizes where only a subset of the studies have an effect. Given 10,000 studies, we threshold each statistic to predict the studies having an effect and the studies not having an effect, and vary the threshold to draw the ROC curves. In A, true prediction rate is the proportion of the studies actually having an effect that are correctly predicted to have an effect and false prediction rate is the proportion of the studies actually not having an effect that are incorrectly predicted to have an effect. In B, true and false prediction rates are similarly defined but in the direction of predicting studies not having an effect. For BF, we use the asymptotic BF of Wakefield [39] with prior where . Next, we evaluate the performance of the statistics in predicting studies not having an effect. The experiment is exactly the same as the previous experiment except that, given a threshold , we predict the studies having m-value to not have an effect. We similarly draw the ROC curves for the three statistics. True and false prediction rates are defined similarly for the objective of predicting the studies not having an effect.
Figure 3B shows that the m-value is even more superior to the other statistics in this experiment than in the previous experiment. The p-value shows the most inferior performance. This is expected because p-value is designed for detecting the presence of an effect but not for detecting the absence of an effect. That is, a non-significant p-value is not evidence of the absence of an effect but can be the result of a small sample size. On the other hand, the BF testing for the absence of an effect is just the reciprocal of the BF testing for the presence of an effect. Thus, the same BF can be used for both purposes. Although the BF performs better than the p-value, the m-value is even more superior. The relative performance gain of the m-value compared to the BF is due to the cross-study information utilized.
P-M Plot: Type 2 Diabetes Data
We apply our P-M plot framework to the real data of the meta-analysis of type 2 Diabetes (T2D) of Scott et al. [10]. The meta-analysis consists of three different GWAS investigations, the Finland-United States Investigation on NIDDM Genetics (FUSION) [10], the Diabetes Genetics Initiative (DGI) [11], and the WTCCC [9], [40].
In their analysis, two SNPs are shown to have a heterogeneous effect, rs8050136 and rs9300039. Ioannidis et al. [17] provide an insightful explanation about the heterogeneity at rs8050136. The WTCCC/UKT2D groups identified evidence for T2D and body mass index (BMI) associations with a set of SNPs including rs8050136 in the FTO region [40]. On the other hand, in the DGI study, the SNP rs8050136 was not significant. The explanation that Ioannidis et al. suggest is that the observed association at rs8050136 (FTO) may be mediated by its association with obesity. In fact, DGI is the only study where the BMI is matched between cases and controls, and the T2D association appears to be mediated through a primary effect on adiposity [11]. Thus, although the truth is unknown, the explanation of Ioannidis et al. is reasonable. Compared to rs8050136, the cause of heterogeneity at rs9300039 is less understood. It is suggested that the heterogeneity might reflect the different tag polymorphisms used in the studies [17].
To gain insights on these studies, we apply our P-M plot. Figure 4A shows the forest plot, the plot showing only the p-values, and the P-M plot for rs8050136. In the P-M plot, DGI appears to be well separated from the other two studies, even though its m-value () is not below the threshold (). Thus, the P-M plot visualizes that DGI can have a different characteristic from the others. Such a separation is not clear in the plot showing only the p-values. In the plot showing only the p-values, DGI is close to FUSION since FUSION is also not very significant (). However, the m-value of FUSION is much greater () than that of DGI. This suggests that the effect is much more likely to exist in the FUSION study than in the DGI study.
Fig. 4. P-M plots of the type 2 diabetes meta-analysis results of Scott et al. [10]. ![P-M plots of the type 2 diabetes meta-analysis results of Scott <i>et al.</i> <em class="ref">[<b>10</b>]</em>.](https://pl-master.mdcdn.cz/media/cache/media_object_image_small/media/image/ac4f3e0d0e0ab66d68f6aff101f9145c.png)
Two associated loci showing high heterogeneity are plotted. The dashed horizontal line shows the genome-wide significance threshold. The dotted vertical lines show the prediction regions based on m-value. Figure 4B shows the plots for rs9300039. The P-M plot shows a different pattern from the P-M plot of rs8050136. In this P-M plot, every study has an m-value greater than 0.5. Thus, no study shows evidence of no effect. Comparing the plots of rs8050136 and rs9300039 gives an interesting observation. In the plot showing only the p-values, both SNPs show a specific pattern of p-values that a single study is considerably more significant than the other two. However, despite of this similarity in the pattern of p-values, the two SNPs' P-M plots look different enough that can lead us to different interpretations. This shows that our P-M plot can provide information that is not apparent in the analysis of only the p-values.
P-M Plot: Crohns Disease Data
We apply our plotting framework to the data of the recent meta-analysis of Crohns disease of Franke et al. [13]. This meta-analysis consists of six different GWAS comprising 6,333 cases and 15,056 controls, and even more samples in the replication stage. In this study, 39 associated loci are newly identified increasing the number of associated loci to 71. We apply our framework to six loci where a high level of heterogeneity is observed. Han and Eskin [23] showed that at these six loci, RE gave more significant p-values than the fixed effects model (FE).
Figure 5 shows the P-M plots of two loci. See Figure S1 for the plots of all six loci. The names of the studies follow the names used in Franke et al. [13]. At these two loci, rs3024505 and rs17293632, the m-value of WTCCC is close to the threshold for predicting no effect. A possible explanation is that the different marker sets could have caused the statistical heterogeneity at these loci. WTCCC [40] used the Affymetrix platform while others used the Illumina platform. Although we do not further investigate this hypothesis, it is true that the P-M plots visualize an interesting outlier behavior of WTCCC at these loci. Such an observation is not clear in both the forest plot and the plot showing only p-values. In the plot showing only p-values, studies having non-significant p-values are all clustered and WTCCC is only one of them. In the forest plot, WTCCC is not the only study showing a small effect size at both loci. For example, at rs3024505, NIDDKNJ shows a smaller effect size than WTCCC. However, the m-value of WTCCC is much smaller than NIDDKNJ's because of the large sample size. Such an interaction between the sample size and the prediction can also be inferred from the forest plot since the forest plot includes the confidence interval. However, it is difficult to numerically quantify the effect of sample size on the prediction by visually examining the forest plot.
Fig. 5. P-M plots of the Crohns disease meta-analysis results of Franke et al. [13]. ![P-M plots of the Crohns disease meta-analysis results of Franke <i>et al.</i> <em class="ref">[<b>13</b>]</em>.](https://pl-master.mdcdn.cz/media/cache/media_object_image_small/media/image/4eeeb58798e01f58ca6cbe65d71ad1e3.png)
Two of six associated loci showing high heterogeneity are plotted. See Figure S1 for the plots of all six loci. The names of the studies follow Franke et al. [13]. The dashed horizontal line shows the genome-wide significance threshold. The dotted vertical lines show the prediction regions based on m-value. Binary Effects Model: False Positive Rate
We estimate the false positive rate of the new binary effects model (BE). Assuming the null hypothesis of no association, we construct 5 studies of sample size 1,000 to build a meta-analysis set. We calculate the meta-analysis p-value of BE using our importance sampling procedure with 10,000 samples. We also calculate the meta-analysis p-values of FE and RE. We build 100 million sets of meta-analysis and estimate the false positive rate as the proportion of the simulated sets whose p-value exceeds a threshold. We vary the threshold levels from 0.05 to . Table 1 shows that all methods including BE control the false positive rates accurately, at all threshold levels examined. When we increase the number of studies from 5 to 10, the results are essentially the same and the false positive rates are controlled (Data not shown).
Tab. 1. False positive rate of FE, RE, and BE at increasing significance thresholds. 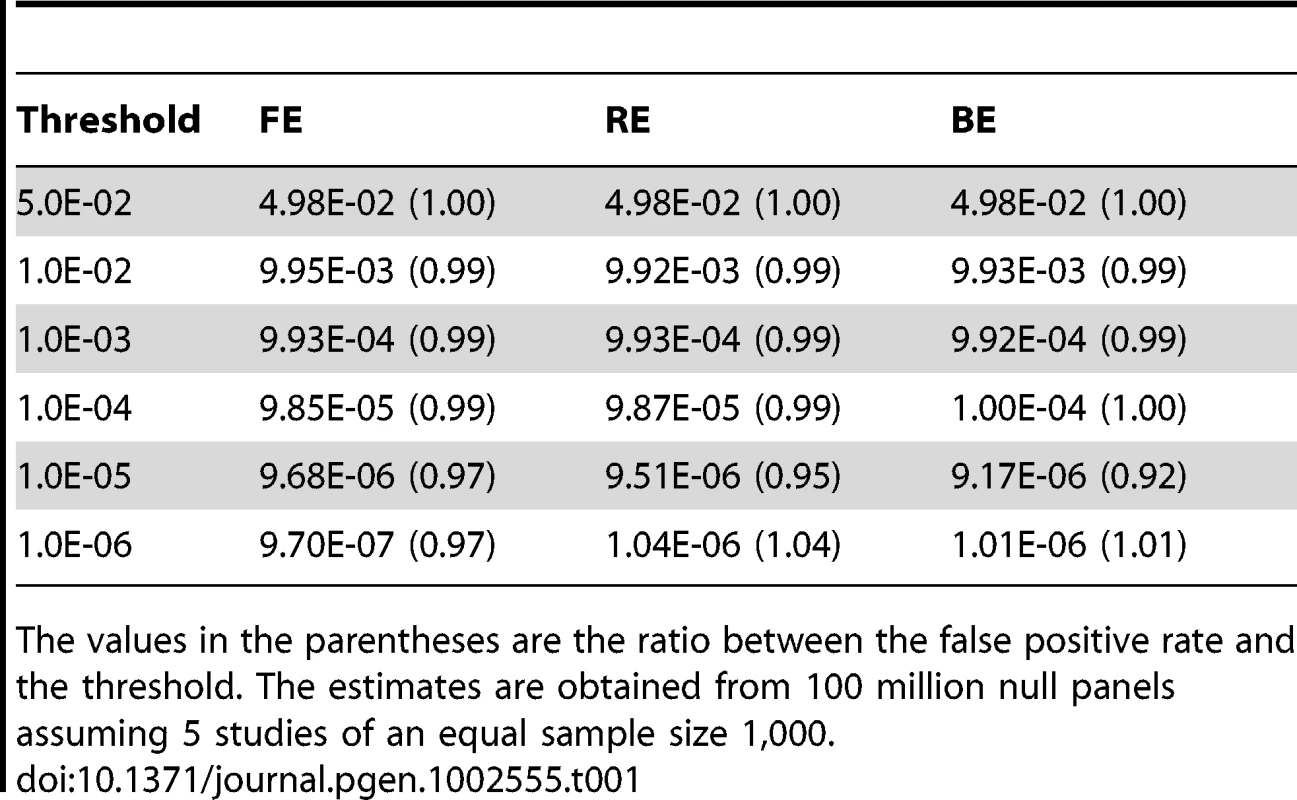
The values in the parentheses are the ratio between the false positive rate and the threshold. The estimates are obtained from 100 million null panels assuming 5 studies of an equal sample size 1,000. Binary Effects Model: Power
We compare the power of BE to the powers of FE and RE. Assuming a meta-analysis of five studies of an equal sample size 1,000, we construct 10,000 meta-analysis sets. The power of each method is estimated as the proportion of the meta-analysis sets whose meta-analysis p-value calculated by each method exceeds the genome-wide threshold ().
We measure power in two different situations. First, we assume a situation that the effect is either present or absent. We decrease the number of studies having an effect () from 5 to 2. We increase the relative risk as decreases, using for respectively, in order to show the relative performance between methods.
Figure 6 shows that except for the case that there is no heterogeneity (), BE is the most powerful among all methods. BE is more powerful than RE, even though both are a random effects model, possibly because it learns the fact that some studies do not have an effect from the data. When there is no heterogeneity (), FE achieves the highest power and BE achieves the lowest power.
Fig. 6. Power of FE, RE, and BE method when the number of studies having an effect varies. 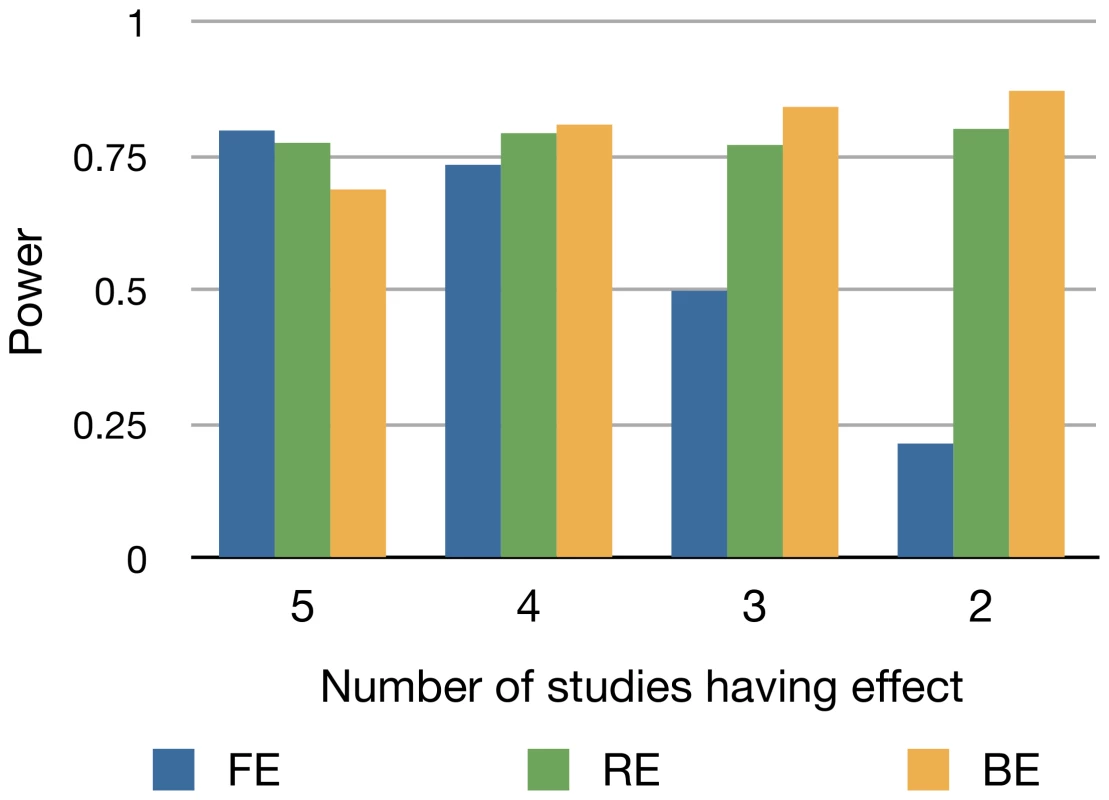
We assume 5 studies and gradually decrease the number of studies having an effect from 5 to 2. We assume an equal sample size of 1,000. We increase the odds ratio as the number of studies decreases to show the relative performance between methods. The power is estimated as the proportion of the simulated 10,000 meta-analysis sets whose meta-analysis p-value calculated by each method exceeds the genome-wide threshold (). Second, we assume a classical setting where the effect sizes follow a normal distribution. Assuming that the mean effect size of , we sample the log of effect size of each study from a normal distribution having the mean and the standard deviation where is the parameter we vary. As increases, the heterogeneity increases. We measure the power of each method varying from zero to one. Figure 7 shows that in this situation, BE is generally less powerful than RE. The power difference between BE and RE is the greatest when the heterogeneity is small. As the heterogeneity increases, BE shows a similar power to RE.
Fig. 7. Power of FE, RE, and BE method when the effect size varies between studies in the pattern following a normal distribution. 
The -axis denotes heterogeneity where we simulate the standard deviation of the effect size (log of relative risk) to be times the effect size. We assume the mean relative risk of 1.3 and assume 5 studies of an equal sample size 1,000. The power is estimated as the proportion of the simulated 10,000 meta-analysis sets whose meta-analysis p-value calculated by each method exceeds the genome-wide threshold (). Binary Effects Model: Real Data
We apply BE to the real data of Crohns disease of Franke et al. [13]. Han and Eskin [23] showed that out of 69 associated loci analyzed, RE gave more significant p-values than FE at six loci where high level of heterogeneity is observed. We calculate the p-values at these loci using BE and compare to the p-values of FE and RE.
Table 2 shows that at all six loci where RE gave more significant p-values than FE, BE gives even more significant p-values. The reason why BE gives more significant p-values can be explained by examining the P-M plots of these loci in Figure 5 and Figure S1. The P-M plots show that at these loci, some studies show high m-values and some studies show low m-values, suggesting a bimodal distribution of effect size. Thus, the situation is very similar to the case that the effect is either present or not, in which case BE achieves higher power than RE as shown in Figure 6.
Tab. 2. Application of FE, RE, and BE to the Crohns disease meta-analysis results of Franke et al. [13]. ![Application of FE, RE, and BE to the Crohns disease meta-analysis results of Franke <i>et al.</i> <em class="ref">[13]</em>.](https://pl-master.mdcdn.cz/media/cache/media_object_image_small/media/image/b5f35de9d1ad82df6781fc98a8910f7e.png)
The boldface denotes the top p-value among the three methods. Only six associated loci are presented that were shown to have more significant RE p-values than FE p-values [23]. Binary Effects Model: Accuracy of Importance Sampling
We measure how accurately the importance sampling procedure of BE estimates the p-value depending on the number of samples used. We calculate the BE p-value for the same dataset in 100 different runs to estimate the variance of the p-value estimate. Our criterion of interest is the ratio between the standard deviation of our estimate and the target p-value. For this, we use the 69 associated loci in the Crohns disease data of Franke et al. [13] that were previously analyzed in Han and Eskin [23]. We measure the ratio for each locus and average over all loci. We do this varying the number of samples from 1,000 to 1,000,000.
Table 3 shows that as the number of samples used for importance sampling increases, the accuracy increases. The pattern of accuracy increase is what we would usually expect in a sampling procedure; standard deviation is decreased approximately by the square root of the sample size increase. When the number of samples is 1,000, the ratio is roughly 0.5. A ratio of 0.5 is large, but can be enough for initial screening if we would apply an adaptive sampling that samples larger number of samples only for loci that are at least moderately significant (e.g. ).
Tab. 3. Accuracy of importance sampling depending on the number of samples. 
For each given number of samples used for the importance sampling, we measure the variance of the p-value estimate of BE by running the importance sampling 100 different times for the same dataset. We use the 69 associated loci from the Crohns disease data of Franke et al. [13]. The ratio between the standard deviation of the estimate and the target p-value is reported, which is averaged over 69 loci. Binary Effects Model: Computational Efficiency
We measure the computational efficiency of the importance sampling procedure of BE. In our software, we implemented an adaptive sampling procedure that samples smaller number first () and then larger number () for the loci that are at least moderately significant. In the machine equipped with Intel Xeon 1.68 GHz CPU, when we use 1,000 samples in the importance sampling, calculating BE p-values of 1,000 loci for the meta-analysis of 10 studies takes 100 seconds. Thus, to calculate BE p-values of one million loci assuming that 1,000 loci among them are moderately significant, it will take approximately 30 hours which is a feasible amount of time. If the number of samples is increased to achieve better accuracy, such as and , the procedure will still be efficient if one uses multiple computers or a cluster since the procedure is parallelizable.
Discussion
We introduce a framework facilitating the interpretation of meta-analysis results based on a new statistic representing the posterior probability that the effect exists in each study. Our framework utilizes cross-study information and is shown to help interpretations in the simulations and the real data. The new statistic also allows us to develop a new association testing procedure called the binary effects model.
In the current meta-analyses of genome-wide association studies, heterogeneity is often observed and our framework will be a useful tool for interpreting such results. We expect that our framework will be even more useful in the future meta-analyses. As the number of studies in a meta-analysis grows, the chance of heterogeneity will increase [6]. Also, a meta-analytic approach can often be applied to a broader area such as to multiple diseases with similar etiology, in which case the heterogeneity is more likely to occur. Moreover, the majority of the current meta-analyses only use the fixed effects model (FE). The use of a random effects model (RE) approach [23] such as the binary effects model presented herein will increase the number of identified associations showing heterogeneity, since an RE approach is more powerful than FE for detecting associations with heterogeneity.
One limitation of our approach is that although the new statistic can predict the studies having an effect and the studies not having an effect, it does not distinguish the true heterogeneity and the statistical heterogeneity [22]. Discriminating between the two can be very difficult based on the observed data and might often be possible only by external data such as the replication studies. In that sense, our method can help discriminating them because one can come up with a hypothesis based on m-values that the heterogeneity is caused by specific design factors and then control the factors in the replication stage. The heterogeneity will disappear in the replication stage if it was due to the design factors.
Similarly to other Bayesian approaches [35], [36], the prior choice in our method can have a non-negligible effect on the predictions. For the prior of the effect size , it is important to set a reasonable value based on the prior information about the effect size. See Stephens and Balding [35] for the general guideline for this choice. For the prior of the probability that the effect exists , we used the uniform distribution () in this paper. However, different priors can also be used for different situations. If one expects that most of the studies have an effect, an asymmetric prior such as can be used. If one is certain that the studies having an effect and the studies not having an effect are mixed, a bell-shape prior such as can be used. See Figure S2 for the plots of the possible choices of priors.
Supporting Information
Zdroje
1. CochranWG 1954 The combination of estimates from different experiments. Biometrics 10 101 129
2. MantelNHaenszelW 1959 Statistical aspects of the analysis of data from retrospective studies of disease. J Natl Cancer Inst 22 719 48
3. FleissJL 1993 The statistical basis of meta-analysis. Stat Methods Med Res 2 121 45
4. HardyJSingletonA 2009 Genomewide association studies and human disease. New England Journal of Medicine 360 1759 1768
5. de BakkerPIWFerreiraMARJiaXNealeBMRaychaudhuriS 2008 Practical aspects of imputation-driven meta-analysis of genome-wide association studies. Hum Mol Genet 17 R122 8
6. CantorRMLangeKSinsheimerJS 2010 Prioritizing gwas results: A review of statistical methods and recommendations for their application. Am J Hum Genet 86 6 22
7. ZegginiEIoannidisJPA 2009 Meta-analysis in genome-wide association studies. Pharmacoge - nomics 10 191 201
8. BarrettJCClaytonDGConcannonPAkolkarBCooperJD 2009 Genome-wide association study and meta-analysis find that over 40 loci affect risk of type 1 diabetes. Nat Genet 41
9. ZegginiEWeedonMNLindgrenCMFraylingTMElliottKS 2007 Replication of genome - wide association signals in uk samples reveals risk loci for type 2 diabetes. Science 316 1336 41
10. ScottLJMohlkeKLBonnycastleLLWillerCJLiY 2007 A genome-wide association study of type 2 diabetes in finns detects multiple susceptibility variants. Science 316 1341 5
11. SaxenaRVoightBFLyssenkoVBurttNPde BakkerPIW 2007 Genome-wide association analysis identifies loci for type 2 diabetes and triglyceride levels. Science 316 1331 6
12. ScottLJMugliaPKongXQGuanWFlickingerM 2009 Genome-wide association and meta-analysis of bipolar disorder in individuals of european ancestry. Proc Natl Acad Sci U S A 106 7501 6
13. FrankeAMcGovernDPBBarrettJCWangKRadford-SmithGL 2010 Genome-wide meta-analysis increases to 71 the number of confirmed crohn's disease susceptibility loci. Nat Genet 42 1118 25
14. StahlEARaychaudhuriSRemmersEFXieGEyreS 2010 Genome-wide association study meta-analysis identifies seven new rheumatoid arthritis risk loci. Nat Genet 42 508 14
15. DerSimonianRLairdN 1986 Meta-analysis in clinical trials. Control Clin Trials 7 177 88
16. HigginsJPTThompsonSG 2002 Quantifying heterogeneity in a meta-analysis. Stat Med 21 1539 58
17. IoannidisJPAPatsopoulosNAEvangelouE 2007 Uncertainty in heterogeneity estimates in meta-analyses. BMJ 335 914 6
18. FieldAP 2003 The problems in using fixed-effects models of meta-analysis on real-world data. Understanding Statistics
19. TangH 2006 Confronting ethnicity-specific disease risk. Nature genetics 38 13
20. BarrosoILuanJWheelerEWhittakerPWassonJ 2008 Population-specific risk of type 2 diabetes conferred by hnf4a p2 promoter variants: a lesson for replication studies. Diabetes 57 3161 5
21. Kim-CohenJCaspiATaylorAWilliamsBNewcombeR 2006 Maoa, maltreatment, and gene-environment interaction predicting children's mental health: new evidence and a meta-analysis. Mol Psychiatry 11 903 913
22. PereiraTVPatsopoulosNASalantiGIoannidisJPA 2009 Discovery properties of genome-wide association signals from cumulatively combined data sets. Am J Epidemiol 170 1197 206
23. HanBEskinE 2011 Random-effects model aimed at discovering associations in meta-analysis of genome-wide association studies. American journal of human genetics 88 586 598
24. ZaitlenNEskinE 2010 Imputation aware meta-analysis of genome-wide association studies. Genet Epidemiol 34 537 42
25. EvangelouEMaraganoreDMIoannidisJPA 2007 Meta-analysis in genome-wide association datasets: strategies and application in parkinson disease. PLoS ONE 2 e196 doi:10.1371/journal.pone.0000196
26. IoannidisJPA 2008 Interpretation of tests of heterogeneity and bias in meta-analysis. J Eval Clin Pract 14 951 7
27. HardyRJThompsonSG 1996 A likelihood approach to meta-analysis with random effects. Statistics in Medicine 15 619 629
28. BiggerstaffBJTweedieRL 1997 Incorporating variability in estimates of heterogeneity in the random effects model in meta-analysis. Stat Med 16 753 68
29. ThompsonSGSharpSJ 1999 Explaining heterogeneity in meta-analysis: a comparison of methods. Statistics in Medicine 18 2693 2708
30. FarrerLACupplesLAHainesJLHymanBKukullWA 1997 Effects of age, sex, and ethnicity on the association between apolipoprotein e genotype and alzheimer disease. a meta-analysis. apoe and alzheimer disease meta analysis consortium. JAMA 278 1349 56
31. HelgadottirAManolescuAHelgasonAThorleifssonGThorsteinsdottirU 2006 A variant of the gene encoding leukotriene a4 hydrolase confers ethnicity-specific risk of myocardial infarction. Nat Genet 38 68 74
32. CorellaDOrdovasJM 2005 Single nucleotide polymorphisms that inuence lipid metabolism: Interaction with dietary factors. Annu Rev Nutr 25 341 90
33. KleebergerSRPedenD 2005 Gene-environment interactions in asthma and other respiratory diseases. Annu Rev Med 56 383 400
34. CaspiAMoffittTE 2006 Gene-environment interactions in psychiatry: joining forces with neu-roscience. Nat Rev Neurosci 7 583 90
35. StephensMBaldingDJ 2009 Bayesian statistical methods for genetic association studies. Nat Rev Genet 10 681 90
36. MarchiniJHowieBMyersSMcVeanGDonnellyP 2007 A new multipoint method for genome-wide association studies by imputation of genotypes. Nat Genet 39 906 13
37. WassermanL 2004 All of statistics Springer New York
38. HanBKangHMEskinE 2009 Rapid and accurate multiple testing correction and power estimation for millions of correlated markers. PLoS Genet 5 e1000456 doi:10.1371/journal.pgen.1000456
39. WakefieldJ 2009 Bayes factors for genome-wide association studies: comparison with p-values. Genet Epidemiol 33 79 86
40. ConsortiumWTCC 2007 Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447 661 78
Štítky
Genetika Reprodukční medicína
Článek Physiological Notch Signaling Maintains Bone Homeostasis via RBPjk and Hey Upstream of NFATc1Článek Intronic -Regulatory Modules Mediate Tissue-Specific and Microbial Control of / TranscriptionČlánek Probing the Informational and Regulatory Plasticity of a Transcription Factor DNA–Binding DomainČlánek Repression of Germline RNAi Pathways in Somatic Cells by Retinoblastoma Pathway Chromatin ComplexesČlánek An Alu Element–Associated Hypermethylation Variant of the Gene Is Associated with Childhood ObesityČlánek Three Essential Ribonucleases—RNase Y, J1, and III—Control the Abundance of a Majority of mRNAsČlánek Genomic Tools for Evolution and Conservation in the Chimpanzee: Is a Genetically Distinct Population
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2012 Číslo 3
-
Všechny články tohoto čísla
- Comprehensive Research Synopsis and Systematic Meta-Analyses in Parkinson's Disease Genetics: The PDGene Database
- Genomic Analysis of the Hydrocarbon-Producing, Cellulolytic, Endophytic Fungus
- Networks of Neuronal Genes Affected by Common and Rare Variants in Autism Spectrum Disorders
- Akirin Links Twist-Regulated Transcription with the Brahma Chromatin Remodeling Complex during Embryogenesis
- Too Much Cleavage of Cyclin E Promotes Breast Tumorigenesis
- Imprinted Genes … and the Number Is?
- Genetic Architecture of Highly Complex Chemical Resistance Traits across Four Yeast Strains
- Exploring the Complexity of the HIV-1 Fitness Landscape
- MNS1 Is Essential for Spermiogenesis and Motile Ciliary Functions in Mice
- A Fundamental Regulatory Mechanism Operating through OmpR and DNA Topology Controls Expression of Pathogenicity Islands SPI-1 and SPI-2
- Evidence for Positive Selection on a Number of MicroRNA Regulatory Interactions during Recent Human Evolution
- Variation in Modifies Risk of Neonatal Intestinal Obstruction in Cystic Fibrosis
- PIF4–Mediated Activation of Expression Integrates Temperature into the Auxin Pathway in Regulating Hypocotyl Growth
- Critical Evaluation of Imprinted Gene Expression by RNA–Seq: A New Perspective
- A Meta-Analysis and Genome-Wide Association Study of Platelet Count and Mean Platelet Volume in African Americans
- Mouse Genetics Suggests Cell-Context Dependency for Myc-Regulated Metabolic Enzymes during Tumorigenesis
- Transcriptional Control in Cardiac Progenitors: Tbx1 Interacts with the BAF Chromatin Remodeling Complex and Regulates
- Synthetic Lethality of Cohesins with PARPs and Replication Fork Mediators
- APOBEC3G-Induced Hypermutation of Human Immunodeficiency Virus Type-1 Is Typically a Discrete “All or Nothing” Phenomenon
- Interpreting Meta-Analyses of Genome-Wide Association Studies
- Error-Prone ZW Pairing and No Evidence for Meiotic Sex Chromosome Inactivation in the Chicken Germ Line
- -Dependent Chemosensory Functions Contribute to Courtship Behavior in
- Diverse Forms of Splicing Are Part of an Evolving Autoregulatory Circuit
- Phenotypic Plasticity of the Drosophila Transcriptome
- Physiological Notch Signaling Maintains Bone Homeostasis via RBPjk and Hey Upstream of NFATc1
- Precocious Metamorphosis in the Juvenile Hormone–Deficient Mutant of the Silkworm,
- Igf1r Signaling Is Indispensable for Preimplantation Development and Is Activated via a Novel Function of E-cadherin
- Accurate Prediction of Inducible Transcription Factor Binding Intensities In Vivo
- Mitochondrial Oxidative Stress Alters a Pathway in Strongly Resembling That of Bile Acid Biosynthesis and Secretion in Vertebrates
- Mammalian Neurogenesis Requires Treacle-Plk1 for Precise Control of Spindle Orientation, Mitotic Progression, and Maintenance of Neural Progenitor Cells
- Tcf7 Is an Important Regulator of the Switch of Self-Renewal and Differentiation in a Multipotential Hematopoietic Cell Line
- REST–Mediated Recruitment of Polycomb Repressor Complexes in Mammalian Cells
- Intronic -Regulatory Modules Mediate Tissue-Specific and Microbial Control of / Transcription
- Age-Dependent Brain Gene Expression and Copy Number Anomalies in Autism Suggest Distinct Pathological Processes at Young Versus Mature Ages
- A Genome-Wide Association Study Identifies Variants Underlying the Shade Avoidance Response
- -by- Regulatory Divergence Causes the Asymmetric Lethal Effects of an Ancestral Hybrid Incompatibility Gene
- Genome-Wide Association and Functional Follow-Up Reveals New Loci for Kidney Function
- A Natural System of Chromosome Transfer in
- Cell Size and the Initiation of DNA Replication in Bacteria
- Probing the Informational and Regulatory Plasticity of a Transcription Factor DNA–Binding Domain
- Repression of Germline RNAi Pathways in Somatic Cells by Retinoblastoma Pathway Chromatin Complexes
- Temporal Transcriptional Profiling of Somatic and Germ Cells Reveals Biased Lineage Priming of Sexual Fate in the Fetal Mouse Gonad
- Rapid Analysis of Genome Rearrangements by Multiplex Ligation–Dependent Probe Amplification
- Metabolic Profiling of a Mapping Population Exposes New Insights in the Regulation of Seed Metabolism and Seed, Fruit, and Plant Relations
- The Atypical Calpains: Evolutionary Analyses and Roles in Cellular Degeneration
- The Silkworm Coming of Age—Early
- Development of a Panel of Genome-Wide Ancestry Informative Markers to Study Admixture Throughout the Americas
- Balanced Codon Usage Optimizes Eukaryotic Translational Efficiency
- The Min System and Nucleoid Occlusion Are Not Required for Identifying the Division Site in but Ensure Its Efficient Utilization
- Neurobeachin, a Regulator of Synaptic Protein Targeting, Is Associated with Body Fat Mass and Feeding Behavior in Mice and Body-Mass Index in Humans
- Statistical Analysis of Readthrough Levels for Nonsense Mutations in Mammalian Cells Reveals a Major Determinant of Response to Gentamicin
- Gene Reactivation by 5-Aza-2′-Deoxycytidine–Induced Demethylation Requires SRCAP–Mediated H2A.Z Insertion to Establish Nucleosome Depleted Regions
- The miR-35-41 Family of MicroRNAs Regulates RNAi Sensitivity in
- Genetic Basis of Hidden Phenotypic Variation Revealed by Increased Translational Readthrough in Yeast
- An Alu Element–Associated Hypermethylation Variant of the Gene Is Associated with Childhood Obesity
- Modelling Human Regulatory Variation in Mouse: Finding the Function in Genome-Wide Association Studies and Whole-Genome Sequencing
- Novel Loci for Adiponectin Levels and Their Influence on Type 2 Diabetes and Metabolic Traits: A Multi-Ethnic Meta-Analysis of 45,891 Individuals
- Polycomb-Like 3 Promotes Polycomb Repressive Complex 2 Binding to CpG Islands and Embryonic Stem Cell Self-Renewal
- Insulin/IGF-1 and Hypoxia Signaling Act in Concert to Regulate Iron Homeostasis in
- EMF1 and PRC2 Cooperate to Repress Key Regulators of Arabidopsis Development
- Three Essential Ribonucleases—RNase Y, J1, and III—Control the Abundance of a Majority of mRNAs
- Contrasted Patterns of Molecular Evolution in Dominant and Recessive Self-Incompatibility Haplotypes in
- A Machine Learning Approach for Identifying Novel Cell Type–Specific Transcriptional Regulators of Myogenesis
- Genomic Tools for Evolution and Conservation in the Chimpanzee: Is a Genetically Distinct Population
- Nos2 Inactivation Promotes the Development of Medulloblastoma in Mice by Deregulation of Gap43–Dependent Granule Cell Precursor Migration
- Intracranial Aneurysm Risk Locus 5q23.2 Is Associated with Elevated Systolic Blood Pressure
- Heritability and Genetic Correlations Explained by Common SNPs for Metabolic Syndrome Traits
- A Genome-Wide Association Study of Nephrolithiasis in the Japanese Population Identifies Novel Susceptible Loci at 5q35.3, 7p14.3, and 13q14.1
- DNA Damage in Nijmegen Breakage Syndrome Cells Leads to PARP Hyperactivation and Increased Oxidative Stress
- DNA Resection at Chromosome Breaks Promotes Genome Stability by Constraining Non-Allelic Homologous Recombination
- Genetic Analysis of Floral Symmetry in Van Gogh's Sunflowers Reveals Independent Recruitment of Genes in the Asteraceae
- A Splice Site Variant in the Bovine Gene Compromises Growth and Regulation of the Inflammatory Response
- Promoter Nucleosome Organization Shapes the Evolution of Gene Expression
- The Nucleoside Diphosphate Kinase Gene Acts as Quantitative Trait Locus Promoting Non-Mendelian Inheritance
- The Ciliogenic Transcription Factor RFX3 Regulates Early Midline Distribution of Guidepost Neurons Required for Corpus Callosum Development
- Phosphorylation of the RNA–Binding Protein HOW by MAPK/ERK Enhances Its Dimerization and Activity
- A Genome-Wide Scan of Ashkenazi Jewish Crohn's Disease Suggests Novel Susceptibility Loci
- Parkinson's Disease–Associated Kinase PINK1 Regulates Miro Protein Level and Axonal Transport of Mitochondria
- LMW-E/CDK2 Deregulates Acinar Morphogenesis, Induces Tumorigenesis, and Associates with the Activated b-Raf-ERK1/2-mTOR Pathway in Breast Cancer Patients
- Mapping the Hsp90 Genetic Interaction Network in Reveals Environmental Contingency and Rewired Circuitry
- Autoregulation of the Noncoding RNA Gene
- The Human Pancreatic Islet Transcriptome: Expression of Candidate Genes for Type 1 Diabetes and the Impact of Pro-Inflammatory Cytokines
- Spo0A∼P Imposes a Temporal Gate for the Bimodal Expression of Competence in
- Antagonistic Regulation of Apoptosis and Differentiation by the Cut Transcription Factor Represents a Tumor-Suppressing Mechanism in
- A Downstream CpG Island Controls Transcript Initiation and Elongation and the Methylation State of the Imprinted Macro ncRNA Promoter
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- PIF4–Mediated Activation of Expression Integrates Temperature into the Auxin Pathway in Regulating Hypocotyl Growth
- Metabolic Profiling of a Mapping Population Exposes New Insights in the Regulation of Seed Metabolism and Seed, Fruit, and Plant Relations
- A Splice Site Variant in the Bovine Gene Compromises Growth and Regulation of the Inflammatory Response
- Comprehensive Research Synopsis and Systematic Meta-Analyses in Parkinson's Disease Genetics: The PDGene Database
Přihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



