-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
Top novinky
Reklama- Videa
- Podcasty
Nové podcasty
Reklama- Práce v oboru
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaThe bHLH-PAS Transcription Factor Dysfusion Regulates Tarsal Joint Formation in Response to Notch Activity during Leg Development
One of the keys of the evolutionary success of arthropods, the most diversified group of animals, is the acquisition of joints that allow the articulation of their appendages. Two main kinds of joints with different morphologies and evolutionary origin are found in the fly leg: the proximal or “true” joints that are motile due to muscular attachment and the distal joints that are immotile. A common event during joint formation is the activation of the Notch pathway at the presumptive joints along the leg proximo-distal axis. In this work we investigated how the same pathway, Notch, can control the formation of such homologous although different structures. We described that the transcription factor Dysfusion (Dys) is a Notch target required for distal joint development and that is sufficient to induce joint-like structures when ectopically expressed. Dys controls two important morphogenetic events that direct tarsal joint development such as programed cell death and epithelial cell shape. Moreover, we identified a regulatory DNA sequence that controls dys expression in the tarsal segment by direct binding of the transcriptional effector of the Notch pathway Su(H). Thus, Notch controls the development of proximal vs distal joints by the recruitment of specific downstream target genes such as dys.
Published in the journal: . PLoS Genet 10(10): e32767. doi:10.1371/journal.pgen.1004621
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1004621Summary
One of the keys of the evolutionary success of arthropods, the most diversified group of animals, is the acquisition of joints that allow the articulation of their appendages. Two main kinds of joints with different morphologies and evolutionary origin are found in the fly leg: the proximal or “true” joints that are motile due to muscular attachment and the distal joints that are immotile. A common event during joint formation is the activation of the Notch pathway at the presumptive joints along the leg proximo-distal axis. In this work we investigated how the same pathway, Notch, can control the formation of such homologous although different structures. We described that the transcription factor Dysfusion (Dys) is a Notch target required for distal joint development and that is sufficient to induce joint-like structures when ectopically expressed. Dys controls two important morphogenetic events that direct tarsal joint development such as programed cell death and epithelial cell shape. Moreover, we identified a regulatory DNA sequence that controls dys expression in the tarsal segment by direct binding of the transcriptional effector of the Notch pathway Su(H). Thus, Notch controls the development of proximal vs distal joints by the recruitment of specific downstream target genes such as dys.
Introduction
Throughout evolution, animal appendages have diversified to display very different morphologies, which are indicative of their diverse functionality such as locomotion, feeding or environment exploration. One of the keys of the evolutionary success of arthropods (from Greek árthron, “joint”, and pous i.e. “foot”), the most diversified group of animals, is the acquisition of joints that allow the articulation of their appendages. Appendages are external projections from the body wall, which require the formation of a proximo-distal axis (PD) that is specified de novo during development from previously established antero-posterior and dorso-ventral axes. In Drosophila, thoracic appendage primordia are specified during embryogenesis by the expression of the homeobox gene Distalless (Dll) [1]. During larval development, the PD axis is generated by the juxtaposition of cells that express two signaling molecules, wingless (wg) and decapentaplegic (dpp). They regulate the expression of the leg “gap genes” Dll, dachshund (dac) and homothorax (hth) dividing the leg into distal, medial and proximal domains, respectively (reviewed by [2]) [3]–[8]. Subsequently, the distal leg is further subdivided into more discrete domains of tarsal-specific gene expression in response to the Epidermal Growth Factor Receptor (EGFR) pathway activity [9], [10].
The Drosophila adult leg is divided in 10 segments (coxa, throchanter, femur, tibia, five tarsal segments and the claw or pretarsus), which are articulated due to the presence of joints in between them. A common event during joint formation is the positioning of Notch ligands Delta (Dl) and Serrate (Ser) by the combined action of the leg “gap genes” and PD tarsal genes in concentric rings at the proximal end of each segment [11]–[14]. Notch pathway activation adjacent and distal to the Dl and Ser expression domains involves the proteolytic cleavage and release of the intracellular fragment of Notch (NICD) that acts as a transcriptional co-activator with proteins of the CBF1-Suppressor of Hairless (Su(H)) family [15]. In the leg joints, Notch pathway activation, visualized by the expression of its target genes Enhancer-of-split mβ (E(spl)mβ) and big brain (bib), mediate leg segmentation and growth. Therefore, mutant clones for components of the Notch signaling pathway that span two leg segments are associated with joint and growth defects [12], [14], [16], [17].
While Notch activity is absolutely required for all joints, proximal and distal joints are functionally, morphologically and evolutionarily different [18]–[24]. Proximal joints, also known as “true joints”, such as the tibial/tarsal joint, are asymmetrical and motile due to muscular attachments. In contrast, distal joints, the ones that separate the tarsal segments, are radially symmetrical, not motile and depend on a different molecular mechanism for their development [20], [22]. Moreover, several genes are specifically expressed in the “true joints” such as the odd-skipped (odd) family members odd, drumstick (drm) and sister of odd and bowl (sob), and others are restricted to the tarsal ones, like deadpan (dpn) or tarsal-less (tal) [14], [25], [26].
Two morphogenetic processes, apoptosis and changes in cell shape, contribute to joint formation [20], [27]. Apoptosis in the legs involves the Jun kinase (JNK)-mediated activation of the pro-apoptotic gene reaper (rpr) in response to sharp boundaries of Dpp activity [20], [28]. The early expression of dpp in the leg disc in dorsal-anterior cells is later refined into a segmented pattern of incomplete rings with higher levels at the distal edge of each segment [20]. It has been suggested that a confrontation of cells with different levels of Dpp pathway activity at the distal end of tarsal segments triggers cell death via JNK activation at both sides of the activity discontinuity [20], [28]. Interestingly, the cell death-mediated mechanism is required for joint architecture only in the tarsal segments [20]. Changes in cell shape are in part mediated by the modulation of RhoGTPases activity [27]. RhoGTPases function as molecular switches (active GTP-bound state and inactive GDP-bound state) that regulate a variety of developmental processes such as cytoskeletal dynamics, cell migration, cell polarity, cell-cycle progression, vesicle trafficking and cytokinesis [29]. A subset of RhoGEFs and RhoGAPs, proteins that regulate Rho activity, are specifically expressed in all joints or restricted to tarsal joints, and the downregulation of some of them produce defects in leg joint formation [27].
An important and yet unresolved question is to understand how the same signaling pathway, Notch, could direct the formation of homologous but morphogenetically distinct structures, such as the different joints along the PD axis of the leg, which will require the deployment of different genetic programs for their formation [11], [12], [14], [20], [22]. In this work we characterize the expression, function and regulation of the Npas4/NXF vertebrate ortholog dysfusion (dys). dys encodes a basic-helix-loop-helix PAS containing (bHLH-PAS) transcription factor required for embryonic tracheal development [30]–[32]. Here we describe a novel and essential role for dys during leg joint formation. We show that dys is absolutely required for tarsal joint formation while it is dispensable for proximal joints. Dys regulates the expression of the pro-apoptotic genes rpr and head involution defective (hid) and the Rho GTPase regulators RhoGAP71E and RhoGEF2. dys expression at the presumptive tarsal joints is controlled by a dedicated cis-regulatory module (CRM), directly regulated by Notch through binding of Su(H). In summary our results provide a molecular explanation of how Notch can regulate the formation of different types of joints along the leg PD axis.
Results
dys is expressed in the presumptive tarsal joints
Proximal and distal leg joints are not only morphologically and functionally different, but also the mechanisms that sculpt them have evolved independently [18]–[22]. Since the Notch pathway is required for both types of joints, other factors should contribute to the differential development of distal versus proximal joints. To understand how the tarsal joints are genetically distinguished from the proximal joints, we have searched in the flylight database for CRMs exclusively active in rings in either tarsal or ‘true’ joints [33](see Materials and Methods). We identified two overlapping sequences within the dysfusion (dys) genomic locus that drove very similar or identical GFP reporter gene expression in concentric rings at the tarsomeres in third instar and prepupal leg imaginal discs (Figure 1A and Figure S1). We also detected GFP expression in distal rings in the antenna for these two CRMs (Figure S1). Using an available antibody for Dys [30], we confirmed that dys is expressed at the distal end of each tarsal segment in prepupal and third instar leg discs, with the exception of the boundary between the most distal tarsus and the pretarsus, which accounts for all the four tarsal joints (Figure 1B and 1C). We also detected an incomplete ring of expression at the distal tibia that was not reproduced by any of the two CRMs identified (Figure 1A and 1B).
Fig. 1. dys is expressed in the presumptive tarsal joints. 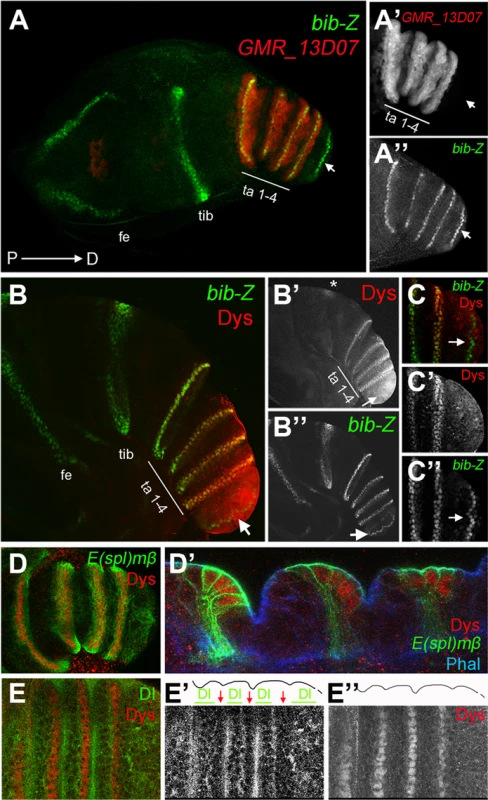
(A) Activity of the Janelia line GMR_13D07 that drives GFP expression (red) in the presumptive tarsal leg joints. Note that this line is not active in the most distal joint (arrow) or the proximal ones. bib-Z in green marks all the joints. Tarsal segments 1–4 (ta 1–4), tibia (tib) and femur (fe). (A′ and A″) Single channels for GMR_13D07 and bib-Z, respectively. Distal tip of prepupal leg discs is to the right in all the panels. (B and C) Dys antibody staining (red) and bib-Z (green) expression in a prepupa leg disc. (C) Close view of the tip of the leg. Single channels for Dys (B′ and C′) and bib-Z (B″ and C″). Note that dys is expressed in the presumptive tarsal joints 1 to 4 (ta 1–4) and in an incomplete ring in the tibial/tarsal joint (asterisk), and that Dys is not expressed in the tarsal/pretarsal joint (arrow). (D) Dys (red) co-localize with E(spl)mβ-CD2 (green) in the tarsal joints. (D′) Sagittal view of a prepupal leg disc epithelium stained with Phallodin (Phal) (blue), E(spl)mβ-CD2 (green) and Dys (red). The joints between tarsal segments 1/2, 2/3 and 3/4 are shown. (E) Dys (red) is localized distal and adjacent to Dl domains (green) in the tarsal segments 1 to 4. Single channels for Dl (E′) and Dys (E″) are shown. The row of dys-expressing cells is marked with red arrows, while Dl domains are indicated with green bars in E′. Outlined of the leg is drawn in E′ and E″. The specification of joints is controlled by the local activation of the Notch pathway [11], [12], [14]. Notch downstream targets and its ligands Dl and Ser are expressed in complementary concentric rings along the PD axis of the leg. The Notch targets bib and E(spl)mβ are restricted to the distal end of each segment (joint domain), while the Notch ligands, Dl and Ser, are restricted to proximal adjacent cells (inter-joint domain) [11], [12], [14]. To confirm that dys expression is restricted to the presumptive joints we compared its expression with that of bib, E(spl)mβ and Dl. We found that dys coexpress with both joint markers at the distal-most cells of tarsal segments 1 to 4, while its expression is distal to Dl localization (Figure 1B–1E). In this manner, dys expression is restricted to Notch-responsive cells of the tarsal region, suggesting a potential role for dys in the development of tarsal joints downstream of Notch.
dys is required for tarsal joint development and promotes epithelial fold formation
To analyze in detail the role of dys during leg morphogenesis we first used a combination of two null dys alelles, dys2/dys3, which produces some escapers (less than 1% of the flies) (see Materials and Methods). All dys2/dys3 adult flies display a complete absence of tarsal joint formation with a small shortening of the tarsal region (Figure 2A and 2B). Interestingly no defects were observed in most proximal joints (including the tibial/tarsal joint where a half-ring of dys expression is detected) or in the tarsus/pretarsus joint (Figure 2A, 2B and Figure S2A and S2B). In addition we detected the loss of the joint between the a5 and the arista in the antenna (Figure 2D and 2E). We also expressed a dys-RNAi construct that efficiently reduced Dys protein levels (Figure S2C). dys knockdown in the Dll domain (Dll-Gal4/UAS-dys-RNAi; distal tibia to claw) abolish joint formation in the tarsal segments without affecting the tibial/tarsal or tarsus/pretarsus joints just as described for dys2/dys3 mutant legs (Figure 2C).
Fig. 2. dys loss and gain of function phenotypes. 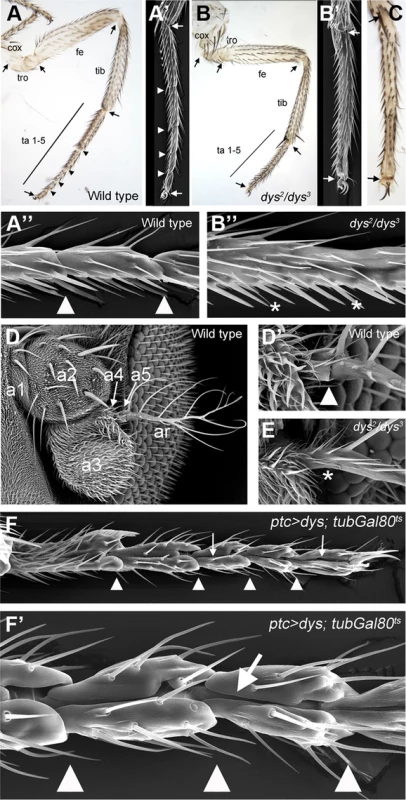
(A) Adult leg of a wild type female, where the distal joints (arrowheads) and the “true” joints (arrows) are pointed out. (A′ and A″) Scanning Electron Microscopy (SEM) imaging of wild type legs. Close view of wild type tarsal joints is shown in A″. (B) Adult leg of a dys2/dys3 mutant female. Note the complete absence of tarsal joints while the “true” joints are not affected (arrows). (B′ and B″) dys2/dys3 mutant leg, and a close view of the tarsal region (B″). Note the lack of tarsal joints (asterisks). (C) Dll-Gal4; UAS-dys-RNAi female leg without tarsal joints. As in the dys2/dys3 mutant leg, the tibial/tarsal and tarsal/pretarsal joints are not affected (arrows). (D) Wild type antenna, with antennal segments marked. (D′) Close view of the wild type a5/arista joint. (E) In a dys2/dys3 mutant antenna the a5/arista joint is missing (asterisk). (F and F′) Male adult leg of a ptc-Gal4; UAS-dys; tubGal80ts fly. Ectopic expression of dys along the PD axis of the leg induced the formation of a fold (arrows). Arrowheads indicate normal position of tarsal joints. To test whether dys is sufficient to induce joint-like structures in the leg, we ectopically expressed dys in an anterior row of cells along the PD axis of the leg disc using the patched (ptc)-Gal4 line. We restricted dys ectopic expression to mid-third instar stage using the Gal80ts technique (see Material and Methods). dys misexpression induces the formation of cuticle folds along the PD axis of the leg that resembles ectopic joint formation (Figure 2F). These joint-like structures are more evident in the tarsal region, although we also detected ectopic folds in more proximal domains such as the tibia or the femur (Figure S2D). These phenotypes are very similar to the ones described for ectopic Notch pathway activation in the leg [12], [14] (see Figure 3M). Although we can not conclude that ectopic dys is able to induce the complete joint architecture, which would include the ball-and-socket structure, we were able to observe a phenotype that recapitulates some major aspects of joint formation such as indentation of the cuticle and fold formation. Taken together, our results suggest that dys is necessary for tarsal joint formation and sufficient to induce joint-like structures.
Fig. 3. dys relationship with the Notch pathway. 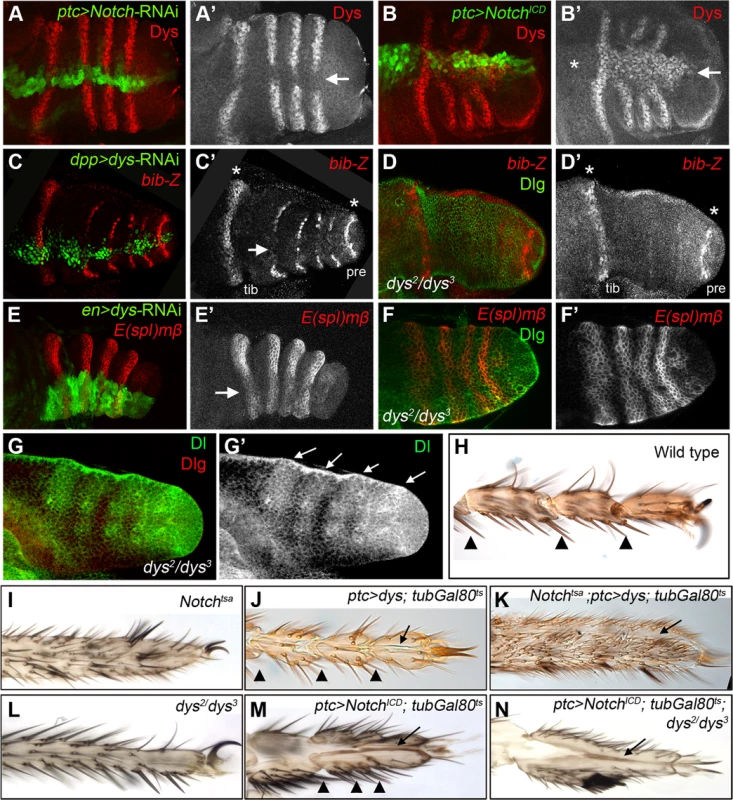
(A) ptc-Gal4, UAS-GFP; UAS-Notch-RNAi prepupal leg disc. Knockdown of Notch levels in the ptc domain (green and arrow in A′) downregulates dys expression (red and single channel in A′). (B) ptc-Gal4, UAS-GFP; UAS-NotchICD prepupal leg disc. Notch pathway activation in the ptc domain (green and arrow in B′) induced dys expression (red and single channel in B′) in the tarsal segments but not in more proximal ones (asterisk). (C) dpp-Gal4. UAS-GFP; UAS-dys-RNAi prepupal leg disc. Dys knockdown in the dpp domain (green and arrow in C) downregulates bib-Z expression in the tarsal segments (red and single channel in C′). Note that bib expression in the tibial/tarsal (tib) and the tarsal/pretarsal (pre) joints remain unaffected (asterisks). (D) dys2/dys3 mutant prepupal leg disc where bib-Z expression (red and single channel in D′) is downregulated in the tarsal segments while in the tibial/tarsal or the tarsal/pretarsal joints remain unaffected (asterisks). Discs-large (Dlg) is in green. (E) en-Gal4, UAS-GFP; UAS-dys-RNAi prepupal leg discs. Knockdown of Dys levels in the posterior compartment (green and arrow) slightly downregulated E(spl)mβ-CD2 expression in the tarsal segments (red and single channel in E′). (F) dys2/dys3 mutant prepupal leg disc. E(spl)mβ-CD2 (red and single channel in F′) is still active in the presumptive tarsal joints. Dlg is in green. (G) Dl expression pattern (green and arrows) remains unaffected in dys2/dys3 mutant prepupal leg. Dlg is in red and single channel for Dl is in (G′). (H–N) Distal adult legs of the following genotypes: (H) wild type, (I) Notchtsa,(J) ptc-Gal4; UAS-dys; tubGal80ts, (K) Notchtsa; ptc-Gal4, UAS-dys; tubGal80ts, (L) dys2/dys3, (M) ptc-Gal4; UAS-NotchICD; tubGal80ts and (N) ptc-Gal4; UAS-NotchICD; tubGal80ts; dys2/dys3. Normal tarsal joint formation is pointed out with arrowheads while ectopic folds along the PD axis are marked with arrows. Note the absence of tarsal joints in Notchtsa (I) and dys2/dys3 (L) mutant legs, and the ectopic folds after dys (J) or NotchICD (M) misexpression in a wild type background and in a Notchtsa (K) or dys2/dys3 (N) mutant background. dys is a target of the Notch pathway
dys expression is restricted to Notch responding cells in the presumptive tarsal leg joints, and its function is required for tarsal joint formation. To analyze the relationship between the Notch pathway and dys, we first tested if dys expression requires Notch activity. We found that Notch downregulation abolishes dys expression (ptc-Gal4/UAS-Notch-RNAi; Figure 3A), while Notch activation promotes dys expression (ptc-Gal4/UAS-NotchICD; Figure 3B). Interestingly, although Notch activation was driven within the ptc domain along the entire PD axis of the leg, dys ectopic expression is restricted to the tarsal segments, suggesting that dys is a downstream target of Notch exclusively in the tarsal region. Next, we tested whether dys is required for the expression of two Notch targets, bib and E(spl)mβ. When Dys function is reduced or eliminated using dys-RNAi or the dys2/dys3 allelic combination, respectively, we observed a strong downregulation of bib-lacZ expression in the tarsal segments, while E(spl)mβ-CD2 is only slightly downregulated or remains unaffected (Figure 3C–3F.). Interestingly, the expression of Dl is not altered in dys2/dys3 mutants (Figure 3G), indicating that neither Dl expression nor Notch activation require Dys function. In contrast, bib-lacZ expression does depend on Dys, although we do not know the basis of this regulation.
To study the functional relationships between the Notch pathway and dys, we first analyzed the ability of Dys to promote the formation of joint-like structures in a Notch mutant background. We induced dys ectopic expression in the ptc domain using the tubGal80ts technique in a hemizygous background for a Notch temperature-sensitive allele (Notchtsa) (see Material and Methods). Notchtsa mutants reared at 17°C (permissive temperature) and shifted to 29°C (restrictive temperature) at late third instar show a complete absence of tarsal joints (Figure 3I compared to 3H). As previously described, temporarily restricted dys misexpression in a wild type background induced cuticle invaginations in a joint-like fashion (Figure 3J). In Ntsa; ptc-Gal4/UAS-dys; tubGal80ts flies shifted to 29°C at late third instar we found a uniform and continuous tarsal cuticle with no joints, characteristic of Notch mutants, and a fold running along the PD axis (compare Figure 3I and 3J with 3K). In the corresponding leg discs, ectopic dys expression induced the formation of an epithelial fold along the PD axis, both in wild type and in Ntsa mutant backgrounds (compare Figure S2F and S2G with S2E). We also performed the reverse experiment, activating the Notch pathway in a dys null background. We found that in dys2/dys3 mutant legs, forced expression of NICD in the ptc domain still retains the capacity to make a fold even though endogenous tarsal joints are not formed (compare Figure 3L and 3M with 3N). This phenotype is very similar to that produced by the ectopic expression of NICD in a wild type background (Figure 3M). This result suggests that forced Notch pathway activation in the absence of dys could be inducing other effector genes that play key roles in the formation of joint-like structures different from the tarsal ones. All together, these results demonstrate that dys is downstream of Notch during tarsal joint formation.
Dys regulate genes implicated in leg joint morphogenesis
Epithelial cells at the presumptive joints undergo apical constriction and form characteristic folds that prefigure the future joint [22], [23]. The formation of epithelial folds involve cells immediately distal to the E(spl)mβ-CD2 expression domain (Figure 4A). In contrast, in dys mutant prepupal legs these cells distal to the E(spl)mβ-CD2 domain fail to reproduce these shape changes compared to control legs (compare Figure 4B and 4A). Two processes help sculpt the joint structure, namely JNK-mediated apoptosis driven by the pro-apoptotic gene rpr [20] and cell shape changes mediated by the Rho-family of GTPases [27]. The expression of RhoGef2 and RhoGap71E is restricted to the tarsal segments where they are coexpressed with a single row of Dys-positive cells and are extended distally to a row of Dys-negative cells (Figure 4C and 4D). Downregulation of Dys levels with a dys-RNAi in the engrailed (en) or the hedgehog (hh) domains lead to compartment cell autonomous loss of RhoGap71E and RhoGef2 expression, respectively (Figure 4E and 4F). Next, we compared the expression of dys with the pro-apoptotic genes rpr and hid. We found that the expression of these two genes in the prepupal leg discs is restricted to the distal end of each tarsal segments, where they are coexpressed with dys (Figure 5A, S3A) [20], [34]. Interestingly, as previously described for RhoGap71E and RhoGef2, we observed that rpr and hid are also expressed in a row of cells distal to dys expression (Figure 5A and S3A). Next we investigated if the expression of rpr and hid depends on dys. The downregulation of Dys levels in anterior cells along the PD axis with a dpp-Gal4 line or in the posterior compartment with en-Gal4, strongly reduced or eliminated the expression of rpr and hid, respectively (Figure 5B and S3B). Furthermore, forced expression of dys in the posterior compartment for 24 hrs using the tubGal80ts technique is sufficient to promote cell autonomously rpr and hid expression in the inter-joint domain (Figure 5C and S3C). This ectopic activation of the pro-apoptotic genes rpr and hid is accompanied by an increase of cell death in larval and prepupal leg discs, as visualized by the presence of activated Caspase-3 (DCas-3) (Figure 5D and S3D). To test if the downregulation in the expression of the pro-apoptotic genes observed after reducing Dys levels is associated to a decrease in cell death, we compared the number of DCas-3 positive cells in wild type and dys mutant legs (see Material and Methods). To determine the precise location of the apoptotic cells during joint formation, we separately counted DCas-3 positive cells within the E(spl)mβ domain and at the gap between two E(spl)mβ domains (termed here as “fold” domain). In wild type prepupal legs we found a significant increase in the number of DCas-3 positive cells in the “fold” domain compared to the E(spl)mβ one, while in dys mutant legs such significant difference was not observed (Figure 5E–G). Interestingly, while the total number of apoptotic cells per segment was comparable between wild type (average = 13,8) and dys mutant legs (average = 14,8), the distribution of DCas-3-positive cells in the joint was altered. dys mutant legs have approximately the same number of apoptotic cells in the E(spl)mβ than in the “fold” domains (Figure 5E–G).
Fig. 4. dys is required for fold formation and for the expression of RhoGap71E and RhoGef2. 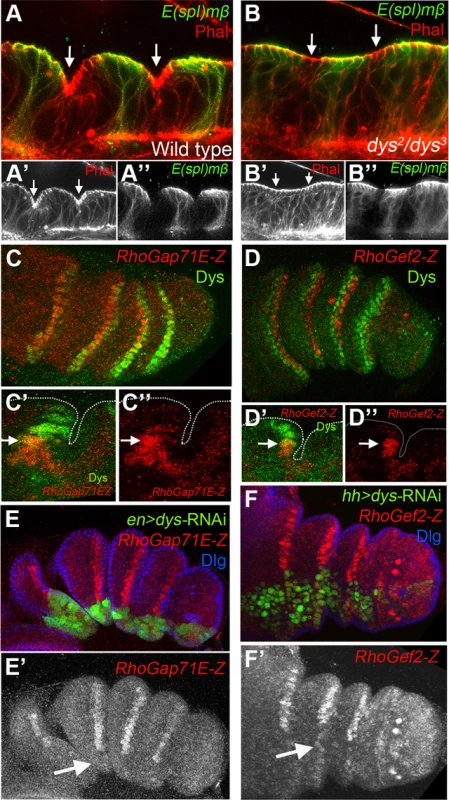
(A and B) Sagittal view of a wild type (A) and dys2/dys3 (B) prepupal leg disc epithelium, stained with Phallodin (Phal) (red) and E(spl)mβ-CD2 (green). The joints between tarsal segments 2/3 and 3/4 are shown in A (arrows), and the corresponding region in B (arrows). Note the apical constriction in cells immediately distal to the E(spl)mβ-CD2 expression domain in the wild type leg and its absence in dys2/dys3 mutant legs. Single channels for Phal (A′ and B′) and E(spl)mβ-CD2 (B′ and B″) are displayed below. (C and D) RhoGap71E-Z (C, red) and RhoGef2-Z (D, red) expression compared to Dys (green). (C′ and D′) Sagittal view of a single joint where the last dys expressing cell is marked by an arrow. Note that both genes, RhoGap71E-Z and RhoGef2-Z, are expressed in a single row of Dys-positive cells and in a row of adjacent Dys-negative cells. (C″ and D″) Single channels for RhoGap71E-Z and RhoGef2-Z. (E) en-Gal4, UAS-GFP; UAS-dys-RNAi (green) and (F) hh-Gal4, UAS-GFP; UAS-dys-RNAi (green) downregulates RhoGap71E-Z and RhoGef2-Z expression (red, arrows), respectively. Dlg is in blue. Single channels for RhoGap71E-Z (E′) and RhoGef2-Z (F′) are displayed below. Fig. 5. Dys regulates rpr expression and apoptosis. 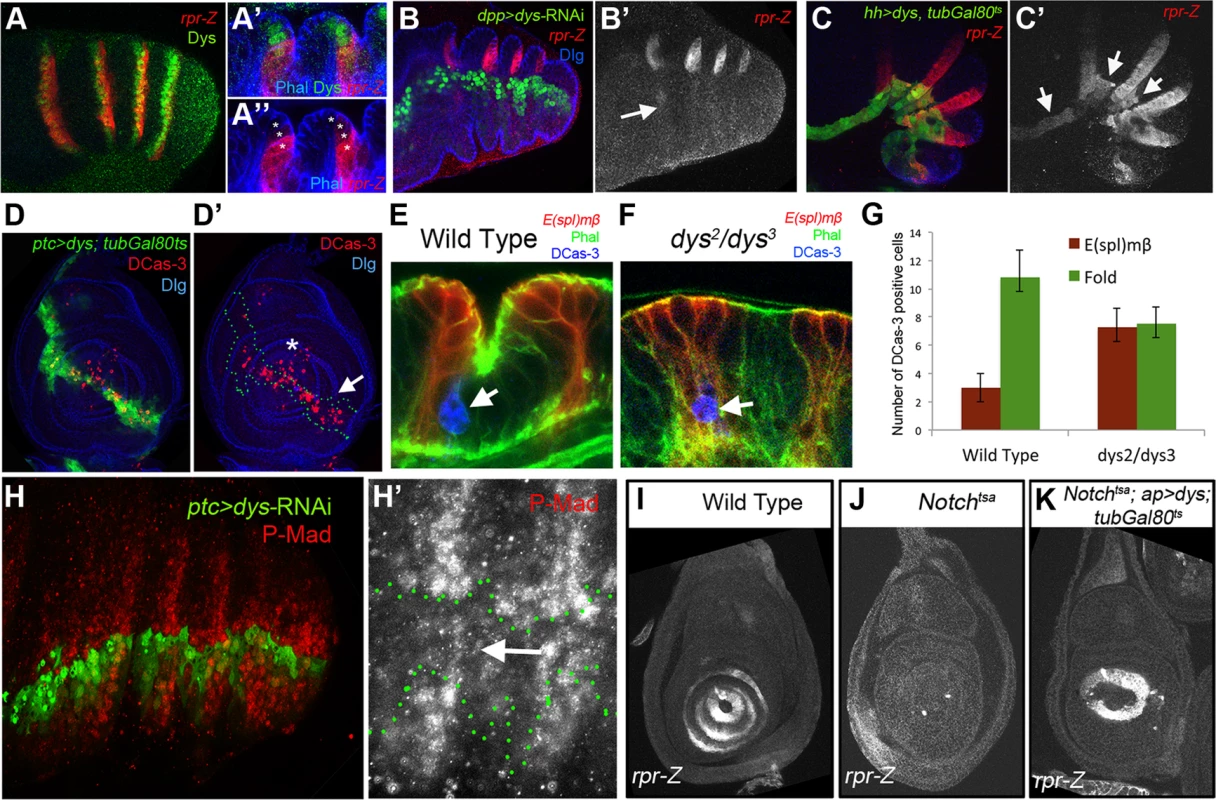
(A) rpr-Z (red) and dys (green) expression in prepupal leg. (A′ and A″) Close sagittal view of two tarsal joints (Phalloidin is in blue). Note that dys expression (green, asterisks) partially overlaps with rpr-Z (red), which extends a couple of cells distal to Dys. (B) dpp-Gal4, UAS-GFP; UAS-dys-RNAi (green) downregulates rpr-Z expression (red, and single channel in B′). rpr-Z downregulation is noted with an arrow in B′. (C) hh-Gal4, UAS-GFP; UAS-dys; tubGal80ts (green). Misexpression of dys for 24 hrs autonomously activates ectopic rpr-Z expression (red, and single channel in C′). Ectopic rpr-Z expression is pointed out with arrows. (D) ptc-Gal4, UAS-GFP; UAS-dys; tubGal80ts (green and green outline in D′). Misexpression of dys for 24 hrs in a third instar leg disc induces Caspase activity (DCas-3) cell autonomously (red, arrow). Dlg is in blue. (D′) Dlg and DCas-3 are shown. Asterisk marks the tip of the leg where endogenous high levels of DCas-3 are observed. (E–F) Close sagittal view of one tarsal joint in a wild type (E) and dys2/dys3 mutant prepupal legs where the domains of E(spl)mβ-CD2 (red) and the “fold” domain (between the two E(spl)mβ-CD2) are visible. Phal is in green and apoptotic cells are visualized by DCas-3 staining (blue, arrows). (G) Quantification of the number of apoptotic cells in the E(spl)mβ-CD2 and “fold” domains in wild type (n = 16 joints counted in 8 legs) and dys2/dys3 mutant prepupae legs (n = 20 joints in 10 legs). Error bars represent the standard error of the mean. The number of apoptotic cells in the E(spl)mβ-CD2 domain in dys2/dys3 is significantly increased in mutant prepupal legs compared to wild type legs (p<0.05). A decrease in the number of apoptotic cells in the “fold” domain in dys2/dys3 mutant prepupal legs can be observed, although is not statistical significant. (H) ptc-Gal4-UAS-GFP; UAS-dys-RNAi (green) prepupal leg. The activity of the Dpp pathway, visualized by P-Mad (red) is decreased after Dys knockdown in the ptc domain. (H′) A close up of the single P-Mad channel is shown with the ptc domain outlined in green. (I–K) rpr-Z expression in third instar leg imaginal discs of the following genotypes: (I) wild type, rpr-Z (J) Notchtsa; ap-Gal4, rpr-Z and (K) Notchtsa; ap-Gal4, rpr-Z/UAS-dys; tubGal80ts flies. (I) rpr-Z is expressed in rings in the presumptive tarsal joints. (J) rpr-Z fails to activate in Notchtsa larvae switched to 29°C for 24–48 hrs before dissection. (K) Notchtsa; ap-Gal4, rpr-Z/UAS-dys; tubGal80ts rescue rpr-Z expression after 24–48 hrs pulse of dys expression in the ap domain in a Notchtsa mutant background. It has been proposed that sharp discontinuities of Dpp activity trigger JNK-mediated apoptosis through the activity of rpr [20], [28]. Therefore we first compared P-Mad (a readout of Dpp signaling) levels with bib expression, which, as we have shown previously, is coexpressed with Dys in the tarsal joints. P-Mad in prepupal legs forms a dorsal ring-like pattern with higher levels coincident with bib-expressing cells (Figure S3E). Next we tested if the absence of rpr expression observed in dys-RNAi legs could be due to a failure in the generation of Dpp activity borders. ptc-Gal4; UAS-dys-RNAi prepupae legs show downregulation of P-Mad levels, suggesting that dys is required for the correct formation of sharp Dpp activity boundaries at the tarsal joints (Figure 5H).
Our results suggest that Dys is a downstream effector of Notch signaling during tarsal joint development, which activates the expression of the pro-apoptotic genes rpr and hid. To confirm these results, we tested if in the absence of Notch activity Dys is still able to induce the transcription of rpr. We found that the depletion of Notch function for 24 to 48 hrs before dissection, using the Notchtsa allele, was sufficient to completely abolish rpr-lacZ expression in the leg disc (compare Figure 5I with 5J). As expected, Notchtsa; ap-Gal4/UAS-dys; tubGal80ts larvae switched to 29°C, 24 to 48 hrs before dissection show ectopic rpr-lacZ expression in the ap domain of the leg imaginal discs (a ring in tarsal segments 4 and part of 5) while no rpr-lacZ expression is detected in more proximal tarsal rings (Figure 5K). These results confirm our previous observations that indicate that Dys is epistatic to Notch during tarsal joint development. In summary, these experiments suggest that dys is necessary for the correct expression of the pro-apoptotic genes rpr and hid and the RhoGTPase modulators RhoGap71E and RhoGef2, being both key events during tarsal joint morphogenesis.
Tango requirements during leg joint morphogenesis
Tango (Tgo) is a bHLH-PAS transcription factor that is able to form heterodimers with multiple bHLH-PAS proteins including Spineless (Ss), Trachealess (Trh) and Dys [32], [35], [36]. In the absence of any of its partners, Tgo localizes in the cytoplasm while in the presence of a partner bHLH-PAS protein, Tgo and its companion form a complex that is translocated into the nucleus where it is functional [36]. During embryonic tracheal development, Tgo dimerizes with Dys to activate the transcription of tracheal fusion target genes [32]. To investigate if the Dys function dependency on Tgo also exists for tarsal joint formation, we analyzed Tgo protein localization and Tgo loss of function phenotypes during joint development. In prepupal leg imaginal discs, Tgo protein shows nuclear localization specifically where dys is expressed, at the presumptive tarsal leg joints (Figure 6A). Moreover, a 24 hr pulse of ectopic expression of dys (ptc-Gal4/UAS-dys; tubGal80ts) induced the nuclear localization of Tgo, while in cells where we reduced Dys levels (en-Gal4; UAS-dys-RNAi) Tgo fails to accumulate in the nucleus of presumptive tarsal joint cells (Figure 6B and 6C).
Fig. 6. Tgo and Dys relationship during tarsal joint development. 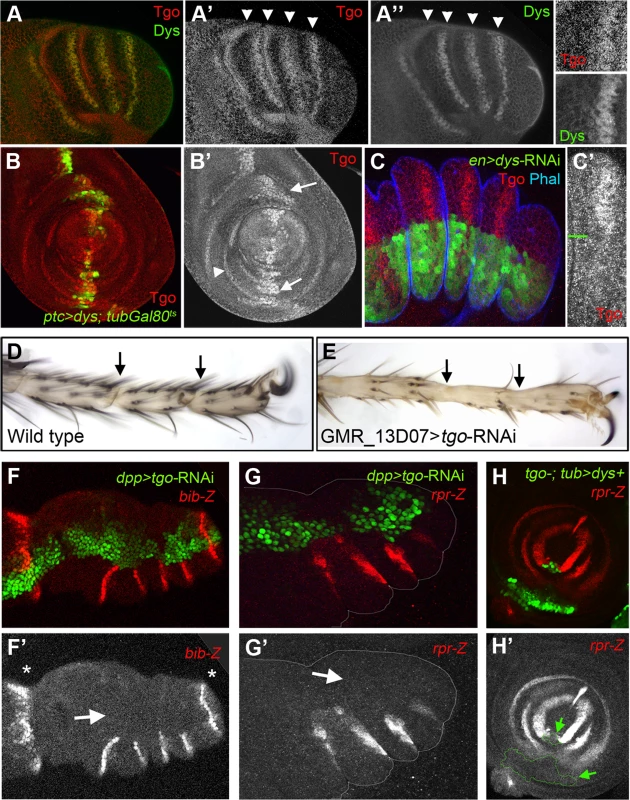
(A) Tgo (red) and Dys (green) co-localization in a prepupae leg. Note that Tgo nuclear protein localization coincides with that of Dys (arrowheads) in the presumptive tarsal joints. Single channels are displayed for Tgo (A′) and Dys (A″) and a close view of a single tarsal joint. (B) ptc-Gal4, UAS-GFP; UAS-dys; tubGal80ts (green) leg imaginal disc stained for Tgo (red and single channel in B″). After 24 hrs of dys misexpression, Tgo localizes to the nuclei in the ptc domain (green, arrows). tgo endogenous expression in tarsal rings is pointed out by arrowhead. (C) en-Gal4-UAS-GFP; UAS-dys-RNAi (green) prepupal leg disc stained for Tgo (red) and Phal (blue). After Dys knockdown, Tgo is no longer detected at the nucleus. (C′) A close view of a single tarsal segment is shown in which the border of compartment is marked with a green line. (D–E) View of the distal end of (D) wild type and (E) GMR_13D07-Gal4; UAS-tgo-RNAi adult legs. No tarsal joints are formed after Tgo knockdown in the presumptive tarsal joint domain (compare D with E; arrows indicate the position of the joints). (F–G) Knockdown of Tgo function in the dpp domain (green, dpp-Gal4, UAS-GFP; UAS-tgo-RNAi) strongly downregulates bib-Z (red in F) and rpr-Z (red in G) expression in prepupal leg discs. Note that no effect is observed in the tibial/tarsal or tarsal/pretarsal bib-Z expression (asterisks). Single channels for bib-Z (F′) and rpr-Z (G′) are displayed below. (H) tgo5 mutant clones that also expressed dys under the tub promoter are marked by GFP (green) loose rpr-lacZ expression (red). (H′) Single channel for rpr-Z is displayed below, and the clone is outlined (arrows). tgo loss of function phenotypes are characterized by fusions and deletions of tarsal segments without affecting proximal ones, phenotypes very similar to those of ss and trh null mutant flies [35], [37]. Since Tgo interacts with Ss to activate transcription, much of tgo tarsal leg defects were attributed to Ss phenotypes [35]. We decided to test if some of the tgo leg phenotypes described could be also due to Dys's inability to activate transcription in the absence of its partner. Tgo knockdown specifically at the presumptive tarsal joints using a tgo-RNAi driven by the GMR_13D07-Gal4 line (see Material and Methods) disrupted joint formation, a phenotype very similar to dys mutant legs (compare Figure 6D with 6E and Figure 2B). As described for dys loss-of-function conditions, we found strong bib-Z downregulation in the presumptive tarsal joints in dpp-Gal4; UAS-tgo-RNAi flies, while the proximal or the most distal ones remain unaffected (Figure 6F). The same effect was observed in the expression of rpr and RhoGap71E after reducing Tgo activity (Figure 6G). All these phenotypes could be due to Dys's inability to activate transcription in the absence of Tgo. To test this possibility we have generated ectopic expression clones of dys in cells that are also mutants for tgo (see Material and Methods). As previously described above, dys ectopic expression in the leg activates rpr in a cell autonomous manner. As expected for a Tgo-obligated transcriptional co-activator, dys ectopic expression clones mutant for tgo are not able to induce the expression of rpr (Figure 6H). All together these results demonstrate that Dys functions with Tgo and both together regulate the expression of their target genes such as rpr.
Binding of Su(H) to a dys 640 bp cis-regulatory module directly integrates Notch input to restrict dys expression to the joints
We have screened 11 DNA fragments derived from the Janelia Gal4 data base that cover the 5′ region and the introns of the dys genomic locus for expression in the leg imaginal disc [33]. Only two overlapping sequences, located between exons 2 and 3, GMR_13D07 and GMR_13B03, drive expression of a GFP reporter in a ring-like pattern that resembles dys expression in the leg (Figure 1A and S1). The overlapping sequence (640 bp long), cloned in a nuclear lacZ reporter vector (see Material and Methods), contains the information necessary to reproduce dys expression pattern in the tarsus (Figure 7B). We have previously shown that Notch acts upstream of dys and, in agreement with our genetic results, dys640-lacZ expression is disrupted in Notch knockdown prepupal leg discs (Figure S4A). Moreover, ectopic expression of NotchICD activates dys640-lacZ expression, although this activation is restricted to the tarsus, just as described for dys endogenous expression (Figure S5B).
Fig. 7. dys is a direct target of the Notch pathway. 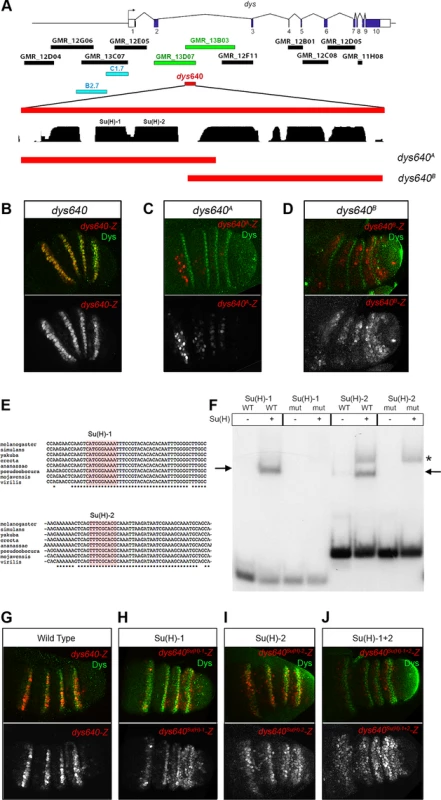
(A) Schematic representation of dys cis-regulatory region. Horizontal bars represent the DNA elements available in the Janelia database that maps around and within the dys gene. Blue bars correspond to the dys-CRMs identified by Jiang et al., 2010 [40] that drove reporter gene expression in fusion tracheal cells. Two Janelia lines (GMR_13D07 and GMR_13B03, green bars) drove reporter gene expression in the tarsal segments of the leg. The 640 bp overlapping sequence (dys640, red bar), DNA conservation between other Drosophilds and the two overlapping fragments (dys640A and dys640B) are also represented. (B–D) Prepupal leg discs stained for Dys (green) and for (B) dys640-Z, (C) dys640A-Z and (D) dys640B-Z. Note the perfect co-localization between dys640-Z and Dys (B). Single lacZ channels are displayed below. (E) DNA sequence of various Drosophilid species surrounding the two identified Su(H) binding sites (red shade) is shown. Asterisks mark perfect DNA conservation between species. Observe that both Su(H) sites are conserved (F) EMSA to assess binding of Su(H) to probes containing wild-type (WT) or mutated (mut) binding sites (see Material and Methods for sequences). Arrows indicate protein-DNA complexes, while asterisk indicate a non-specific band present in both wild type and mutant probe. (G–J) Prepupal leg discs stained for Dys (green) and for (G) dys640-Z, (H) dys640Su(H)-1-Z, (I) dys640Su(H)-2-Z and (J) dys640Su(H)-1+2-Z. All constructs have been inserted in the same genomic location and images were obtained keeping the confocal settings constant in the merge image. Single channels are displayed below, and for dys640Su(H)-1+2–Z the gain has been increase for visualization purposes. To gain further insights in the molecular regulation of dys, we divided the dys640 CRM in two overlapping fragments and studied their expression using the same lacZ reporter construct (Figure 7A). The dys640A-lacZ fragment included two putative Su(H) binding sites conserved among different Drosophila species, and it is expressed weakly in patches of cells that do not overlap with dys expression (Figure 7C and 7E). Fragment dys640B-lacZ does not include these Su(H) sites and resulted in a weak but consistently extended expression of lacZ in the tarsal inter-joint domain, with low levels in dys expressing cells (compare Figure 7D with 7B). The two candidate binding sites were also tested for their ability to bind Su(H) in an electrophoretic mobility shift assay (EMSA) (Figure 7E and 7F). Su(H) binds to both putative sites and this binding was abolished when the sites were mutated (Figure 7F).
To further assess the contribution of the identified binding sites to dys640-lacZ expression, we mutated each individual binding site and the combination of Su(H)-1 and 2 in vivo in a transgenesis reporter assay. Mutation of either of these sites in isolation, dys640Su(H)-1 and dys640Su(H)-2, significantly reduced but did not eliminate lacZ expression (Figure 7G–I). Interestingly we observed a slight derepression of lacZ signal in the inter-joint domain of all tarsal segments, being stronger in the fourth tarsal segment for the two constructs. The combined mutation of the two sites, dys640Su(H)-1+2-lacZ, resulted in an overall weaker expression than mutating each site separately and a derepression of lacZ in the inter-joints throughout the tarsal region and in the distal portion of the tibia (compare Figure 7J with 7G). Taken together, these results indicate that dys is a direct target of Notch through Su(H) binding to the dys640 CRM. We propose that in the absence of Notch signaling, as in the tarsal inter-joint region, Su(H) binds to dys640 CRM repressing dys expression. Conversely, Notch activation at the presumptive joint cells leads to loss of Su(H) repression, probably through a displacement of co-repressors and recruitment of co-activators, therefore converting Su(H) DNA-bound in a positive input to activate dys expression.
Discussion
In this work we analyzed the function of the bHLH-PAS transcription factor Dys during leg joint morphogenesis. Dys has been previously characterized in Drosophila as a transcription factor involved in embryonic tracheal fusion and, as its mammalian ortholog Npas4/Nxf, forms a heterodimer in vivo with Tgo (Arnt) [30]–[32], [38]. We have found that dys is expressed in the presumptive tarsal joints, where it is required for tarsal joint development. In these cells, Dys regulates the expression of the pro-apoptotic genes rpr and hid, and the expression of the RhoGTPases modulators, RhoGEf2 and RhoGap71E. When ectopically expressed, dys is able to induce some aspects of the morphogenetic program necessary for distal joint development such as fold formation and programmed cell death. As described for tracheal formation, this novel Dys function also depends on its obligated partner Tgo to activate the transcription of target genes. We also identified and characterized a dedicated dys CRM that regulates dys expression in the tarsal presumptive leg joints by Notch signaling through direct Su(H) binding. All these data place dys as a key player downstream of Notch, directing distal joint morphogenesis.
Role of dys in tarsal joint formation
In a search for genes expressed exclusively either in distal or proximal leg joints we identified the gene dys. dys is coexpressed with bib and E(spl)mβ and it is distal to the Notch ligand Dl in four rings at the presumptive tarsal joints and in an incomplete ring at the presumptive tibial/tarsal joint. Legs mutant for dys do not develop tarsal joints and have instead a smooth continuous cuticle, without defects in other proximal joints or in the most distal one, the tarsus/pretarsus joint. In dys mutant prepupal leg discs the characteristic apical constriction of cells and the subsequent fold at the presumptive joint are lost. Conversely, ectopic expression of dys is able to induce ectopic folds along the leg that resemble joint-like structures, a phenotype very similar to those observed after misexpression of the activated form of Notch [12], [14]. Several experiments suggest that dys is a Notch target that is indispensable for tarsal joint development: (1) Notch directly regulates dys expression (see below). (2) The expression of the Notch targets bib and rpr absolutely require dys function, even when the Notch pathway is still active. (3) Dys is able to induce the expression of rpr and the formation of joint-like cuticle invaginations in the absence of Notch signaling. All these results place dys genetically downstream of Notch in the development of tarsal joints.
dys is a new Notch-induced target in the distal leg
Our results show that dys expression at the presumptive tarsal segments is controlled by a dedicated CRM 640 bp long that integrates Notch signaling through direct Su(H) binding. This is, to our knowledge, the first characterized Notch direct target described for leg joint development. Interestingly, the mutation of the two identified Su(H) consensus sites, dys640-lacZSu(H)-1+2, resulted in lacZ derepression in the inter-joint domain of the tarsal segments although at lower levels compared to normal signal observed in the presumptive joints in dys640-lacZ control legs. These results are in favor of the “default repression” model in which Su(H) associates with co-repressors in the absence of Notch signaling to repress target gene transcription [39] (Figure 8A). In the event of Notch activation, co-repressors are displaced by NICD, so Su(H) binding could lead to target gene transcription through the recruitment of co-activators. Therefore, dys fulfills the two predictions of the model to occur in the absence of Su(H) binding: (1) target genes will be derepressed and (2) their expression will be reduced in their normal expression domains. These two predictions can be validated in the dys640Su(H)1+2 and dys640B gene reporter constructs, where Su(H) binding is compromised or lost. In both cases we observed a consistent derepression of lacZ expression in the inter-joint domain of the tarsal segments, although with weak levels. We also observed reduced lacZ expression at the presumptive joint domain in the dys CRM with the two Su(H) sites mutated or the dys640B reporter gene compared to the intact dys640. This characteristic is specially evident in the dys640B construct that lacks the two described Su(H) sites and probably others not identified in our analysis.
Fig. 8. dys molecular regulation and model for tarsal joint development. 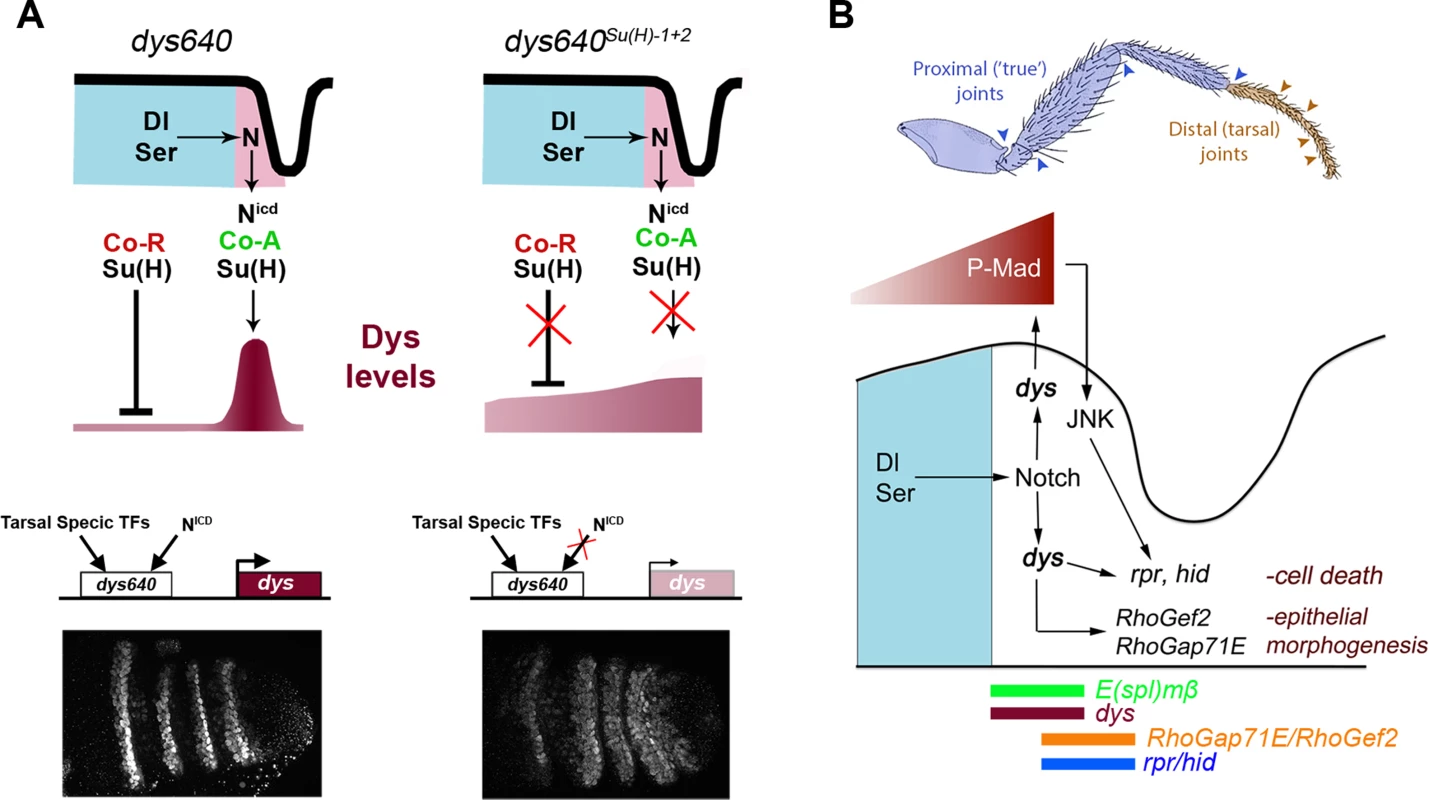
(A) Schematic representation of a tarsal joint of a dys640 wild type (left) and dys640Su(H)-1+2 where the two Su(H) binding sites are mutated (right). See text for description. (B) Model for tarsal joint formation. Blue and brown arrowheads mark proximal and distal joints, respectively. In a tarsal joint, Notch ligands Dl and Ser (blue) activate the Notch pathway in the distal adjacent cells. Notch, in turn, activates dys expression, which regulates the high levels of P-Mad observed in the joint. Dys also controls the expression of the pro-apoptotic genes rpr and hid and the RhoGTPases modulators RhoGef2 and RhoGap71E. The sharp boundary of Dpp signaling also regulates rpr expression via JNK pathway activation. The expression domains of E(spl)mβ, dys, rpr, hid, RhoGef2 and RhoGap71E are indicated. Enhancers are logic integrators of positive and negative inputs that drive precise temporal and spatial gene expression. In the case of dys, two previously identified CRMs drove reporter gene expression in all tracheal fusion cells in the embryo [40] (Figure 7A). For dys expression in the leg we have identified a different CRM that integrates Notch input and is only active in the tarsal presumptive joints. Although Notch pathway is active in rings along the PD axis of the leg, dys expression is restricted to the distal region, suggesting that other inputs are required to give dys its positional specificity along the PD axis. Thus, it is probable that tarsal transcription factors bind to dys640 CRM to restrict its activity (Figure 8A). Some candidates are bric à brac (bab), rotund (rn) or ss, which are expressed in the four tarsal segments where dys is specifically active [41]–[43].
Dys control of leg joint morphogenesis
Joint development involves a complex developmental program that leads to the formation of flexible structures connecting leg segments. At least two processes play key roles during tarsal joint development: epithelial morphogenesis controlled by the activity of RhoGTPases, and JNK-dependent programmed cell death [20], [27]. Several RhoGTPases modulators, RhoGaps and RhoGefs, which have a restricted expression pattern, are regulated by Notch and their downregulation affects joint formation [27]. Another important morphogenetic event is the formation of sharp Dpp signaling boundaries at the presumptive tarsal joints that triggers JNK-dependent localized cell death through rpr activation [20]. Interestingly P-Mad and rpr segmented expression depends on Notch activity, while ectopic formation of new Dpp signaling borders feedback to the system and activate downstream Notch targets such as E(spl)mβ [20]. Therefore, Notch controls tarsal joint morphogenesis in part through Dpp pathway and RhoGTPases regulation. Although a relationship between the Dpp pathway and the expression of RhoGTPases regulators has not been described, it is possible that these genes could also be regulated by sharp Dpp discontinuities, as is the case for rpr. Our results place dys directly downstream of Notch in the presumptive tarsal joints, regulating the Dpp pathway and the expression of rpr, hid, RhoGef2 and RhoGap71E (Figure 8B). One interesting and unresolved question is how Dys can control the expression of rpr, hid, RhoGap71E and RhoGef2 non-autonomously. We have described that these genes are expressed in a row of Dys-positive cells and in an additional Dys-negative distal row of cells (see Figure 4C, 4D, 5A, S3A and 8B). Interestingly, Dys downregulation blocks rpr, hid, RhoGap71E and RhoGef2 expression both in dys expressing and non-expressing cells, while dys misexpression activate rpr and hid exclusively in a cell autonomous manner. Our results suggest that Dys could regulate rpr in two ways. The first one would be a direct activation by Dys and the second one an indirect effect through P-Mad and JNK pathway regulation that also regulates rpr [20]. Therefore, sharp Dpp discontinuities lead to JNK-mediated rpr expression while Notch activation in the presumptive tarsal joints activates dys expression that also regulates rpr cell autonomously. The cross talk between the Notch and Dpp pathways might be important to ensure a robust activation of rpr expression at the tarsal joints, and dys could be a key player in the communication between these pathways (Figure 8B). A similar mechanism to that described for rpr could regulate the expression of hid, RhoGap71E and RhoGef2 in the presumptive tarsal joints.
It would be of interest to study if Dys controls the same subset of genes used for joint morphogenesis in other developmental contexts where it is expressed (e.g. embryonic tracheal fusion or leading edge cells) [30]. The role of Rho GTPases during tracheal development has been previously described [44], [45] but its implication in tracheal fusion and the possible role of Dys as a regulator of Rho GTPases in this process remains to be elucidated.
Another important observation is the discrepancy we observed between the expression of the pro-apoptotic genes and the number of DCas-3 positive cells. While in dys misexpression experiments we observed a positive correlation between the activation of rpr/hid and the increase of apoptotic cells, in dys mutant legs this relationship is more difficult to find. In wild type prepupal legs, the number of DCas-3 cells is significantly increased in cells distal to E(spl)mβ domain, while in dys mutants, this difference is not observed: there is a significant increase of apoptotic cells in the E(spl)mβ domain and a reduction, although not significant, in the “fold” domain. Therefore, our results suggest that in the absence of dys the apoptosis is not preferentially localized in any part of the joint, while in the presence of dys, this apoptosis is bent towards the cells distal to the E(spl)mβ domain. We propose that this imbalance of apoptosis is necessary for the correct formation of the tarsal joints.
Proximal vs Distal joint morphogenesis
An important question in developmental biology is how a given signaling pathway is able to direct different morphogenetic programs. For example, the Dpp and the Sonic Hedgehog (Shh) pathways pattern the fly wing and the ventral neural tube, respectively, in a morphogene concentration-dependent manner, activating different sets of target genes (reviewed in [46]). In the case of the Notch pathway, target genes are regulated cell autonomously and depending on the cellular context a different set of downstream genes is regulated by Notch [15]. In the leg disc, the Notch pathway is similarly activated in 10 concentric rings along the PD axis that prefigure the future joints [11], [12], [14], [23]. Although all joints are homologous structures, distal joints differ from proximal or “true joints” not only in the absence of muscular attachments but also in their morphology, evolutionary origin and in the morphogenetic program that sculpt them [18], [20], [22]. Moreover, while each proximal joint has a unique morphology, all distal joints display the same ball and socket organization. Therefore, if Notch is absolutely required for all joints, the question arises of how Notch can be directing the formation of two very different yet homologous structures like distal and proximal joints. One possible scenario is that Notch controls joint morphogenesis through the activation of different sets of downstream effectors along the PD axis of the leg. Several Notch downstream target genes have been characterized to be required in all joints, such as dAP-2 [16], while others are restricted to the proximal or “true joints”. Members of the odd-skipped family of zinc finger transcription factors are expressed in all joints of the leg except the tarsal joints and they might act redundantly to regulate the development of these proximal joints [21], [47]. Other genes are expressed exclusively in the tarsal joints like tal, dpn or dys. dpn, as dys, also encodes for a bHLH transcription factor expressed in the tarsal joints, although we did not find a phenotype in several dpn mutant allelic combinations or knocking down Dpn levels using a dpn-RNAi construct (data not shown) [26]. Interestingly, as it occurs in the embryonic trachea where Dys and another bHLH-PAS transcription factor, Trh, function in non-overlapping cells [30], in the leg these proteins are also present in distinct domains [37]. It would be interesting to study if the cross-regulation described in the trachea for Trh and Dys also exists in the leg.
Based on the expression and requirements for dys in the joints, we propose a model in which Notch directs the formation of the different joints by the PD-restricted activation of target genes such as odd in proximal joints and dys in the distal ones.
Materials and Methods
Drosophila strains
dys2 and dys3, as well as the UAS-dys strains are described in [30], [31] and GMR_13D07- and GMR_13B03-Gal4 and the rest of Janelia enhancer/GAL4 lines are described in the flylight database (http://flweb.janelia.org/cgi-bin/flew.cgi) [33] and are all publicly available at Bloomington Stock Center. dpn mutant alelles, dpn7, dpn6 and dpnDef3D5 were kindly provided by Antonio Baonza. The reporter lines bib-lacZ and E(spl)-mβ-CD2 [12] were used for assessing Notch pathway activation. To study the relation between dys and the Notch pathway, we used the UAS-NotchICD [48], and UAS-Notch-RNAi lines [49]. The Notch thermosensitive mutant allele (Notchtsa) allowed us to knockdown Notch activity when the flies are shifted to the restrictive temperature (29°C) [50]. For loss - and gain-of-function experiments, we employed the Gal4 lines ptc-Gal4, dpp-Gal4, Dll-Gal4MD212, ap-Gal4, hh-Gal4 and en-Gal4 and the tubGal80ts allele, all-available at Bloomington Stock Center. We used the reporter lines rpr-4kb-lacZ (rpr-lacZ) [51], hid-lacZW05014 [52], RhoGAP71E - and RhoGEF2-lacZ (Bloomington Stock Center). The lines dys-, tgo- and dpn-RNAi are available at the Vienna Drosophila Resource Center (VDRC). RNAi knockdown experiments were performed on a UAS-Dcr-2 background [53]. To generate tgo mutant clones we utilized the null allele tgo5 (Bloomington) and the MARCM technique, which allowed us to simultaneously eliminate tgo function and express dys cell autonomously. The detailed genotype is: yw hs-flp, tub-Gal4; UAS-dys; FRT 82B tubGal80/FRT 82B tgo5. Loss of function clones were created by heat-shocking the larvae for 1 hour at 37°C 48 to 72 hrs after egg laying.
Gain of function experiments
dys gain-of-function experiments were performed using the Gal4-tubGal80ts system, which allowed us temporal restriction of UAS-dys expression to mid-third instar stage. 24 hrs collection of hh-, en-, ap- or ptc-Gal4/UAS-dys; tubGal80ts flies were maintained at restrictive temperature (17°C) until mid-third instar stage, when the fly vials were shifted to the permissive temperature (29°C). Larva and prepupae were dissected between 24 to 48 hrs later.
Immunostaining
Larval and prepupal leg discs were fixed and stained following standard procedures. As primary antibodies we used rabbit and mouse anti-βGal, rabbit anti-Dys (a gift from L. Jiang and S.T. Crews), rabbit anti-DCas-3 (cleaved Drosophila Dcp-1, Cell Signaling Technology), rabbit anti-P-Mad (kindly provided by G. Morata). Mouse anti-Dl, anti-Dlg and anti-Tgo are from Developmental Studies Hybridoma Bank, University of Iowa. TRITC-phalloidin and Phalloidin-Atto 647N were used to stain F-actin (Sigma Aldrich), and secondary antibodies were coupled to Red-X, FITC and Cy5 fluorocromes (Alexa Fluor Dyes, Invitrogen).
To determine the levels of cell death in E(spl)-mβ and “fold” domains, we have performed Z-stack imaging of wild type (n = 8 prepupae leg discs) and dys2/dys3 mutants (n = 10 prepupae leg discs) and counted the number of D-Cas3 positive cells on each domain with the aid of the Fiji software. We selected for this analysis the joints between tarsal segments 2/3 and 3/4.
Cloning of dys CRM in a lacZ reporter vector
The 640 bp overlapping DNA sequence between the GMR_13D07 and GMR_13B03 lines as well as the different mutant conditions were cloned in the HLz attB plasmid vector, which expresses a nuclear lacZ reporter under the control of the cloned sequence [6].
The primers used were the following for each reporter line (restriction sites are underlined and restriction enzyme used is noted in brackets):
dys640:
Forward: 5′-cagtcctaggCCAAGCCGATGAGCCATTCCATACC-3′ (AvrII)
Reverse: 5′-cagtagatctCCACTCTGGAGCAAACCACACCGAA-3′ (BglII)
dys640A:
Forward: 5′-cagtcctaggCCAAGCCGATGAGCCATTCCATACC-3′ (AvrII)
Reverse: 5′-cagtagatctTTCTGCTGATTTTCTTCTTTAGGTT-3′ (BglII)
dys640B:
Forward: 5′-cagtcctaggCTCTCCATGGTTAAGCTCAGACTAA-3′ (AvrII)
Reverse: 5′-cagtagatctCCACTCTGGAGCAAACCACACCGAA-3′ (BglII)
Putative Su(H) binding sites were identified on the basis of a bioinformatics analysis combining data from the JASPAR CORE Insecta database (http://jaspar.genereg.net/) and the Target Explorer tool [54].
Mutagenesis of the Su(H) putative binding sites was performed using the QuikChange Site-Directed Mutagenesis Kit (Stratagene). We used the following primers: dys640Su(H)-1:
Forward: 5′-TCGATCCAAGAACCAAGTCcgagaccAATTTCCGTACACACACAA-3′
Reverse: 5′-TTGTGTGTGTACGGAAATTggtctcgGACTTGGTTCTTGGATCGA-3′ dys640Su(H)-2:
Forward: 5′-GGAGGAAGAAAAAACTCAGtggagacagCAAATTAAGATAATCG-3′ Reverse: 5′-CGATTATCTTAATTTGctgtctccaCTGAGTTTTTTCTTCCTCC-3′
dys640-lacZ reporter construct was inserted both in the 2R (51D) and 3R (86Fb) chromosomal locations. To allow proper comparison, all the dys640-lacZ versions (dys640-lacZ, dys640A-lacZ, dys640B-lacZ, dys640Su(H)-1-lacZ, dys640Su(H)-2-lacZ and dys640Su(H)-1+2-lacZ) were inserted in the same location. Confocal settings were kept constant when imaging wild type and mutant versions of dys640-lacZ, so lacZ expression levels are comparable between these conditions.
Electrophoretic mobility shift assay
An incomplete form of Su(H), that bears the DNA binding domain [55] was translated in vitro using the TNT T7 Quick master MiX kit (Promega) and tested for binding with a series of labeled dsDNA probes. 50 ng of each sense oligonucleotide were labeled following standard procedures with γ-32P ATP, and then hybridized with the complementary “cold” oligonucleotide. Wild type and mutant probes, where nucleotides at consensus Su(H) binding site were mutated, were generated for the two identified Su(H) sites. The designed oligonucleotides were:
Su(H)-1 WT
Forward: 5′-CCAAGTCATGGGAAAATTTCC-3′
Reverse: 5′-GGAAATTTTCCCATGACTTGG-3′
Su(H)-1 mut
Forward: 5′-CCAAGTCcgagaccAATTTCC-3′
Reverse: 5′-GGAAATTggtctcgGACTTGG-3′
Su(H)-2 WT
Forward:
5′-GGAGGAAGAAAAAACTCAGTTTCGCACGCAAATTAAGATAATCG-3′
Reverse:
5′-CGATTATCTTAATTTGCGTGCGAAACTGAGTTTTTTCTTCCTCC-3′
Su(H)-2 mut
Forward: 5′-GGAGGAAGAAAAAACTCAGtggagacagCAAATTAAGATAATCG-3′
Reverse: 5′-CGATTATCTTAATTTGctgtctccaCTGAGTTTTTTCTTCCTCC-3′ Mutated Su(H) sites are noted with lower case letters.
Scanning Electron Microscopy
Wild type and mutant adult flies were collected and their legs and heads dissected without any fixation and avoiding moisture prior to preparation for SEM. The preparation of the samples and Scanning Electron Microscopy was performed at the Microscopy Unit at Universidad Autónoma de Madrid.
Supporting Information
Zdroje
1. CohenSM (1990) Specification of limb development in the Drosophila embryo by positional cues from segmentation genes. Nature 343 : 173–177.
2. EstellaC, VoutevR, MannRS (2012) A dynamic network of morphogens and transcription factors patterns the fly leg. Curr Top Dev Biol 98 : 173–198.
3. Diaz-BenjumeaFJ, CohenB, CohenSM (1994) Cell interaction between compartments establishes the proximal-distal axis of Drosophila legs. Nature 372 : 175–179.
4. LecuitT, CohenSM (1997) Proximal-distal axis formation in the Drosophila leg. Nature 388 : 139–145.
5. CampbellG, WeaverT, TomlinsonA (1993) Axis specification in the developing Drosophila appendage: the role of wingless, decapentaplegic, and the homeobox gene aristaless. Cell 74 : 1113–1123.
6. EstellaC, McKayDJ, MannRS (2008) Molecular integration of wingless, decapentaplegic, and autoregulatory inputs into Distalless during Drosophila leg development. Dev Cell 14 : 86–96.
7. GiorgianniMW, MannRS (2011) Establishment of medial fates along the proximodistal axis of the Drosophila leg through direct activation of dachshund by Distalless. Dev Cell 20 : 455–468.
8. Abu-ShaarM, MannRS (1998) Generation of multiple antagonistic domains along the proximodistal axis during Drosophila leg development. Development 125 : 3821–3830.
9. GalindoMI, BishopSA, GreigS, CousoJP (2002) Leg patterning driven by proximal-distal interactions and EGFR signaling. Science 297 : 256–259.
10. CampbellG (2002) Distalization of the Drosophila leg by graded EGF-receptor activity. Nature 418 : 781–785.
11. BishopSA, KleinT, AriasAM, CousoJP (1999) Composite signalling from Serrate and Delta establishes leg segments in Drosophila through Notch. Development 126 : 2993–3003.
12. de CelisJF, TylerDM, de CelisJ, BraySJ (1998) Notch signalling mediates segmentation of the Drosophila leg. Development 125 : 4617–4626.
13. RauskolbC (2001) The establishment of segmentation in the Drosophila leg. Development 128 : 4511–4521.
14. RauskolbC, IrvineKD (1999) Notch-mediated segmentation and growth control of the Drosophila leg. Dev Biol 210 : 339–350.
15. BrayS, BernardF (2010) Notch Targets and Their Regulation. Curr Top Dev Biol 92 : 253–275.
16. KerberB, MongeI, MuellerM, MitchellPJ, CohenSM (2001) The AP-2 transcription factor is required for joint formation and cell survival in Drosophila leg development. 128 : 1231–1238.
17. MongeI, KrishnamurthyR, SimsD, HirthF, SpenglerM, et al. (2001) Drosophila transcription factor AP-2 in proboscis, leg and brain central complex development. Development 128 : 1239–1252.
18. Snodgrass R (1935) Principles of Insect Morphology. New York: McGraw-Hill. pp. 83–99.
19. CasaresF, MannRS (2001) The ground state of the ventral appendage in Drosophila. Science 293 : 1477–1480.
20. ManjonC, Sanchez-HerreroE, SuzanneM (2007) Sharp boundaries of Dpp signalling trigger local cell death required for Drosophila leg morphogenesis. Nat Cell Biol 9 : 57–63.
21. de Celis IbeasJM, BraySJ (2003) Bowl is required downstream of Notch for elaboration of distal limb patterning. Development 130 : 5943–5952.
22. MirthC, AkamM (2002) Joint development in the Drosophila leg: cell movements and cell populations. Dev Biol 246 : 391–406.
23. TajiriR, MisakiK, YonemuraS, HayashiS (2010) Dynamic shape changes of ECM-producing cells drive morphogenesis of ball-and-socket joints in the fly leg. Development 137 : 2055–2063.
24. TajiriR, MisakiK, YonemuraS, HayashiS (2011) Joint morphology in the insect leg: evolutionary history inferred from Notch loss-of-function phenotypes in Drosophila. Development 138 : 4621–4626.
25. PueyoJI, CousoJP (2011) Tarsal-less peptides control Notch signalling through the Shavenbaby transcription factor. Dev Biol 355 : 183–193.
26. BierE, VaessinH, Younger-ShepherdS, JanLY, JanYN (1992) deadpan, an essential pan-neural gene in Drosophila, encodes a helix-loop-helix protein similar to the hairy gene product. Genes Dev 6 : 2137–2151.
27. GreenbergL, HatiniV (2011) Systematic expression and loss-of-function analysis defines spatially restricted requirements for Drosophila RhoGEFs and RhoGAPs in leg morphogenesis. Mech Dev 128 : 5–17.
28. Adachi-YamadaT, O'ConnorMB (2002) Morphogenetic apoptosis: a mechanism for correcting discontinuities in morphogen gradients. Dev Biol 251 : 74–90.
29. Van AelstL, SymonsM (2002) Role of Rho family GTPases in epithelial morphogenesis. Genes Dev 16 : 1032–1054.
30. JiangL, CrewsST (2003) The Drosophila dysfusion basic helix-loop-helix (bHLH)-PAS gene controls tracheal fusion and levels of the trachealess bHLH-PAS protein. Mol Cell Biol 23 : 5625–5637.
31. JiangL, CrewsST (2006) Dysfusion transcriptional control of Drosophila tracheal migration, adhesion, and fusion. Mol Cell Biol 26 : 6547–6556.
32. JiangL, CrewsST (2007) Transcriptional specificity of Drosophila dysfusion and the control of tracheal fusion cell gene expression. J Biol Chem 282 : 28659–28668.
33. JoryA, EstellaC, GiorgianniMW, SlatteryM, LavertyTR, et al. (2012) A survey of 6,300 genomic fragments for cis-regulatory activity in the imaginal discs of Drosophila melanogaster. Cell Rep 2 : 1014–1024.
34. GuarnerA, ManjonC, EdwardsK, StellerH, SuzanneM, et al. (2014) The zinc finger homeodomain-2 gene of Drosophila controls Notch targets and regulates apoptosis in the tarsal segments. Dev Biol 385 : 350–365.
35. EmmonsRB, DuncanD, EstesPA, KiefelP, MosherJT, et al. (1999) The spineless-aristapedia and tango bHLH-PAS proteins interact to control antennal and tarsal development in Drosophila. Development 126 : 3937–3945.
36. WardMP, MosherJT, CrewsST (1998) Regulation of bHLH-PAS protein subcellular localization during Drosophila embryogenesis. Development 125 : 1599–1608.
37. TajiriR, TsujiT, UedaR, SaigoK, KojimaT (2007) Fate determination of Drosophila leg distal regions by trachealess and tango through repression and stimulation, respectively, of Bar homeobox gene expression in the future pretarsus and tarsus. Dev Biol 303 : 461–473.
38. OoeN, SaitoK, MikamiN, NakatukaI, KanekoH (2004) Identification of a novel basic helix-loop-helix-PAS factor, NXF, reveals a Sim2 competitive, positive regulatory role in dendritic-cytoskeleton modulator drebrin gene expression. Mol Cell Biol 24 : 608–616.
39. BaroloS, StoneT, BangAG, PosakonyJW (2002) Default repression and Notch signaling: Hairless acts as an adaptor to recruit the corepressors Groucho and dCtBP to Suppressor of Hairless. Genes Dev 16 : 1964–1976.
40. JiangL, PearsonJC, CrewsST (2010) Diverse modes of Drosophila tracheal fusion cell transcriptional regulation. Mech Dev 127 : 265–280.
41. CoudercJL, GodtD, ZollmanS, ChenJ, LiM, et al. (2002) The bric a brac locus consists of two paralogous genes encoding BTB/POZ domain proteins and acts as a homeotic and morphogenetic regulator of imaginal development in Drosophila. Development 129 : 2419–2433.
42. St PierreSE, GalindoMI, CousoJP, ThorS (2002) Control of Drosophila imaginal disc development by rotund and roughened eye: differentially expressed transcripts of the same gene encoding functionally distinct zinc finger proteins. Development 129 : 1273–1281.
43. DuncanDM, BurgessEA, DuncanI (1998) Control of distal antennal identity and tarsal development in Drosophila by spineless-aristapedia, a homolog of the mammalian dioxin receptor. Genes Dev 12 : 1290–1303.
44. LeeS, KolodziejPA (2002) The plakin Short Stop and the RhoA GTPase are required for E-cadherin-dependent apical surface remodeling during tracheal tube fusion. Development 129 : 1509–1520.
45. ChiharaT, KatoK, TaniguchiM, NgJ, HayashiS (2003) Rac promotes epithelial cell rearrangement during tracheal tubulogenesis in Drosophila. Development 130 : 1419–1428.
46. TabataT (2001) Genetics of morphogen gradients. Nat Rev Genet 2 : 620–630.
47. HaoI, GreenRB, DunaevskyO, LengyelJA, RauskolbC (2003) The odd-skipped family of zinc finger genes promotes Drosophila leg segmentation. Dev Biol 263 : 282–295.
48. de CelisJF, BrayS (1997) Feed-back mechanisms affecting Notch activation at the dorsoventral boundary in the Drosophila wing. Development 124 : 3241–3251.
49. PresenteA, ShawS, NyeJS, AndresAJ (2002) Transgene-mediated RNA interference defines a novel role for notch in chemosensory startle behavior. Genesis 34 : 165–169.
50. ShellenbargerDL, MohlerJD (1978) Temperature-sensitive periods and autonomy of pleiotropic effects of l(1)Nts1, a conditional notch lethal in Drosophila. Dev Biol 62 : 432–446.
51. JiangC, LamblinAF, StellerH, ThummelCS (2000) A steroid-triggered transcriptional hierarchy controls salivary gland cell death during Drosophila metamorphosis. Mol Cell 5 : 445–455.
52. GretherME, AbramsJM, AgapiteJ, WhiteK, StellerH (1995) The head involution defective gene of Drosophila melanogaster functions in programmed cell death. Genes Dev 9 : 1694–1708.
53. DietzlG, ChenD, SchnorrerF, SuKC, BarinovaY, et al. (2007) A genome-wide transgenic RNAi library for conditional gene inactivation in Drosophila. Nature 448 : 151–156.
54. SosinskyA, BoninCP, MannRS, HonigB (2003) Target Explorer: An automated tool for the identification of new target genes for a specified set of transcription factors. Nucleic Acids Res 31 : 3589–3592.
55. San-JuanBP, BaonzaA (2011) The bHLH factor deadpan is a direct target of Notch signaling and regulates neuroblast self-renewal in Drosophila. Dev Biol 352 : 70–82.
Štítky
Genetika Reprodukční medicína
Článek Oligoasthenoteratozoospermia and Infertility in Mice Deficient for miR-34b/c and miR-449 LociČlánek The Kinesin AtPSS1 Promotes Synapsis and is Required for Proper Crossover Distribution in MeiosisČlánek Payoffs, Not Tradeoffs, in the Adaptation of a Virus to Ostensibly Conflicting Selective PressuresČlánek Examination of Prokaryotic Multipartite Genome Evolution through Experimental Genome ReductionČlánek BMP-FGF Signaling Axis Mediates Wnt-Induced Epidermal Stratification in Developing Mammalian SkinČlánek Role of STN1 and DNA Polymerase α in Telomere Stability and Genome-Wide Replication in ArabidopsisČlánek RNA-Processing Protein TDP-43 Regulates FOXO-Dependent Protein Quality Control in Stress ResponseČlánek Integrating Functional Data to Prioritize Causal Variants in Statistical Fine-Mapping StudiesČlánek Salt-Induced Stabilization of EIN3/EIL1 Confers Salinity Tolerance by Deterring ROS Accumulation inČlánek Ethylene-Induced Inhibition of Root Growth Requires Abscisic Acid Function in Rice ( L.) SeedlingsČlánek Metabolic Respiration Induces AMPK- and Ire1p-Dependent Activation of the p38-Type HOG MAPK PathwayČlánek Signature Gene Expression Reveals Novel Clues to the Molecular Mechanisms of Dimorphic Transition inČlánek A Mouse Model Uncovers LKB1 as an UVB-Induced DNA Damage Sensor Mediating CDKN1A (p21) DegradationČlánek Dominant Sequences of Human Major Histocompatibility Complex Conserved Extended Haplotypes from to
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2014 Číslo 10
-
Všechny články tohoto čísla
- An Deletion Is Highly Associated with a Juvenile-Onset Inherited Polyneuropathy in Leonberger and Saint Bernard Dogs
- Licensing of Yeast Centrosome Duplication Requires Phosphoregulation of Sfi1
- Oligoasthenoteratozoospermia and Infertility in Mice Deficient for miR-34b/c and miR-449 Loci
- Basement Membrane and Cell Integrity of Self-Tissues in Maintaining Immunological Tolerance
- The Kinesin AtPSS1 Promotes Synapsis and is Required for Proper Crossover Distribution in Meiosis
- Germline Mutations in Are Associated with Familial Gastric Cancer
- POT1a and Components of CST Engage Telomerase and Regulate Its Activity in
- Controlling Meiotic Recombinational Repair – Specifying the Roles of ZMMs, Sgs1 and Mus81/Mms4 in Crossover Formation
- Payoffs, Not Tradeoffs, in the Adaptation of a Virus to Ostensibly Conflicting Selective Pressures
- FHIT Suppresses Epithelial-Mesenchymal Transition (EMT) and Metastasis in Lung Cancer through Modulation of MicroRNAs
- Genome-Wide Mapping of Yeast RNA Polymerase II Termination
- Examination of Prokaryotic Multipartite Genome Evolution through Experimental Genome Reduction
- White Cells Facilitate Opposite- and Same-Sex Mating of Opaque Cells in
- BMP-FGF Signaling Axis Mediates Wnt-Induced Epidermal Stratification in Developing Mammalian Skin
- Genome-Wide Association Study of CSF Levels of 59 Alzheimer's Disease Candidate Proteins: Significant Associations with Proteins Involved in Amyloid Processing and Inflammation
- COE Loss-of-Function Analysis Reveals a Genetic Program Underlying Maintenance and Regeneration of the Nervous System in Planarians
- Fat-Dachsous Signaling Coordinates Cartilage Differentiation and Polarity during Craniofacial Development
- Identification of Genes Important for Cutaneous Function Revealed by a Large Scale Reverse Genetic Screen in the Mouse
- Sensors at Centrosomes Reveal Determinants of Local Separase Activity
- Genes Integrate and Hedgehog Pathways in the Second Heart Field for Cardiac Septation
- Systematic Dissection of Coding Exons at Single Nucleotide Resolution Supports an Additional Role in Cell-Specific Transcriptional Regulation
- Recovery from an Acute Infection in Requires the GATA Transcription Factor ELT-2
- HIPPO Pathway Members Restrict SOX2 to the Inner Cell Mass Where It Promotes ICM Fates in the Mouse Blastocyst
- Role of and in Development of Abdominal Epithelia Breaks Posterior Prevalence Rule
- The Formation of Endoderm-Derived Taste Sensory Organs Requires a -Dependent Expansion of Embryonic Taste Bud Progenitor Cells
- Role of STN1 and DNA Polymerase α in Telomere Stability and Genome-Wide Replication in Arabidopsis
- Keratin 76 Is Required for Tight Junction Function and Maintenance of the Skin Barrier
- Encodes the Catalytic Subunit of N Alpha-Acetyltransferase that Regulates Development, Metabolism and Adult Lifespan
- Disruption of SUMO-Specific Protease 2 Induces Mitochondria Mediated Neurodegeneration
- Caudal Regulates the Spatiotemporal Dynamics of Pair-Rule Waves in
- It's All in Your Mind: Determining Germ Cell Fate by Neuronal IRE-1 in
- A Conserved Role for Homologs in Protecting Dopaminergic Neurons from Oxidative Stress
- The Master Activator of IncA/C Conjugative Plasmids Stimulates Genomic Islands and Multidrug Resistance Dissemination
- An AGEF-1/Arf GTPase/AP-1 Ensemble Antagonizes LET-23 EGFR Basolateral Localization and Signaling during Vulva Induction
- The Proteomic Landscape of the Suprachiasmatic Nucleus Clock Reveals Large-Scale Coordination of Key Biological Processes
- RNA-Processing Protein TDP-43 Regulates FOXO-Dependent Protein Quality Control in Stress Response
- A Complex Genetic Switch Involving Overlapping Divergent Promoters and DNA Looping Regulates Expression of Conjugation Genes of a Gram-positive Plasmid
- ZTF-8 Interacts with the 9-1-1 Complex and Is Required for DNA Damage Response and Double-Strand Break Repair in the Germline
- Integrating Functional Data to Prioritize Causal Variants in Statistical Fine-Mapping Studies
- Tpz1-Ccq1 and Tpz1-Poz1 Interactions within Fission Yeast Shelterin Modulate Ccq1 Thr93 Phosphorylation and Telomerase Recruitment
- Salt-Induced Stabilization of EIN3/EIL1 Confers Salinity Tolerance by Deterring ROS Accumulation in
- Telomeric (s) in spp. Encode Mediator Subunits That Regulate Distinct Virulence Traits
- Ethylene-Induced Inhibition of Root Growth Requires Abscisic Acid Function in Rice ( L.) Seedlings
- Ancient Expansion of the Hox Cluster in Lepidoptera Generated Four Homeobox Genes Implicated in Extra-Embryonic Tissue Formation
- Mechanism of Suppression of Chromosomal Instability by DNA Polymerase POLQ
- A Mutation in the Mouse Gene Leads to Impaired Hedgehog Signaling
- Keeping mtDNA in Shape between Generations
- Targeted Exon Capture and Sequencing in Sporadic Amyotrophic Lateral Sclerosis
- TIF-IA-Dependent Regulation of Ribosome Synthesis in Muscle Is Required to Maintain Systemic Insulin Signaling and Larval Growth
- At Short Telomeres Tel1 Directs Early Replication and Phosphorylates Rif1
- Evidence of a Bacterial Receptor for Lysozyme: Binding of Lysozyme to the Anti-σ Factor RsiV Controls Activation of the ECF σ Factor σ
- Hsp40s Specify Functions of Hsp104 and Hsp90 Protein Chaperone Machines
- Feeding State, Insulin and NPR-1 Modulate Chemoreceptor Gene Expression via Integration of Sensory and Circuit Inputs
- Functional Interaction between Ribosomal Protein L6 and RbgA during Ribosome Assembly
- Multiple Regulatory Systems Coordinate DNA Replication with Cell Growth in
- Fast Evolution from Precast Bricks: Genomics of Young Freshwater Populations of Threespine Stickleback
- Mmp1 Processing of the PDF Neuropeptide Regulates Circadian Structural Plasticity of Pacemaker Neurons
- The Nuclear Immune Receptor Is Required for -Dependent Constitutive Defense Activation in
- Genetic Modifiers of Neurofibromatosis Type 1-Associated Café-au-Lait Macule Count Identified Using Multi-platform Analysis
- Juvenile Hormone-Receptor Complex Acts on and to Promote Polyploidy and Vitellogenesis in the Migratory Locust
- Uncovering Enhancer Functions Using the α-Globin Locus
- The Analysis of Mutant Alleles of Different Strength Reveals Multiple Functions of Topoisomerase 2 in Regulation of Chromosome Structure
- Metabolic Respiration Induces AMPK- and Ire1p-Dependent Activation of the p38-Type HOG MAPK Pathway
- The Specification and Global Reprogramming of Histone Epigenetic Marks during Gamete Formation and Early Embryo Development in
- The DAF-16 FOXO Transcription Factor Regulates to Modulate Stress Resistance in , Linking Insulin/IGF-1 Signaling to Protein N-Terminal Acetylation
- Genetic Influences on Translation in Yeast
- Analysis of Mutants Defective in the Cdk8 Module of Mediator Reveal Links between Metabolism and Biofilm Formation
- Ribosomal Readthrough at a Short UGA Stop Codon Context Triggers Dual Localization of Metabolic Enzymes in Fungi and Animals
- Gene Duplication Restores the Viability of Δ and Δ Mutants
- Selection on a Variant Associated with Improved Viral Clearance Drives Local, Adaptive Pseudogenization of Interferon Lambda 4 ()
- Break-Induced Replication Requires DNA Damage-Induced Phosphorylation of Pif1 and Leads to Telomere Lengthening
- Dynamic Partnership between TFIIH, PGC-1α and SIRT1 Is Impaired in Trichothiodystrophy
- Signature Gene Expression Reveals Novel Clues to the Molecular Mechanisms of Dimorphic Transition in
- Mutations in Moderate or Severe Intellectual Disability
- Multifaceted Genome Control by Set1 Dependent and Independent of H3K4 Methylation and the Set1C/COMPASS Complex
- A Role for Taiman in Insect Metamorphosis
- The Small RNA Rli27 Regulates a Cell Wall Protein inside Eukaryotic Cells by Targeting a Long 5′-UTR Variant
- MMS Exposure Promotes Increased MtDNA Mutagenesis in the Presence of Replication-Defective Disease-Associated DNA Polymerase γ Variants
- Coexistence and Within-Host Evolution of Diversified Lineages of Hypermutable in Long-term Cystic Fibrosis Infections
- Comprehensive Mapping of the Flagellar Regulatory Network
- Topoisomerase II Is Required for the Proper Separation of Heterochromatic Regions during Female Meiosis
- A Splice Mutation in the Gene Causes High Glycogen Content and Low Meat Quality in Pig Skeletal Muscle
- KDM5 Interacts with Foxo to Modulate Cellular Levels of Oxidative Stress
- H2B Mono-ubiquitylation Facilitates Fork Stalling and Recovery during Replication Stress by Coordinating Rad53 Activation and Chromatin Assembly
- Copy Number Variation in the Horse Genome
- Unifying Genetic Canalization, Genetic Constraint, and Genotype-by-Environment Interaction: QTL by Genomic Background by Environment Interaction of Flowering Time in
- Spinster Homolog 2 () Deficiency Causes Early Onset Progressive Hearing Loss
- Genome-Wide Discovery of Drug-Dependent Human Liver Regulatory Elements
- Developmentally-Regulated Excision of the SPβ Prophage Reconstitutes a Gene Required for Spore Envelope Maturation in
- Protein Phosphatase 4 Promotes Chromosome Pairing and Synapsis, and Contributes to Maintaining Crossover Competence with Increasing Age
- The bHLH-PAS Transcription Factor Dysfusion Regulates Tarsal Joint Formation in Response to Notch Activity during Leg Development
- A Mouse Model Uncovers LKB1 as an UVB-Induced DNA Damage Sensor Mediating CDKN1A (p21) Degradation
- Notch3 Interactome Analysis Identified WWP2 as a Negative Regulator of Notch3 Signaling in Ovarian Cancer
- An Integrated Cell Purification and Genomics Strategy Reveals Multiple Regulators of Pancreas Development
- Dominant Sequences of Human Major Histocompatibility Complex Conserved Extended Haplotypes from to
- The Vesicle Protein SAM-4 Regulates the Processivity of Synaptic Vesicle Transport
- A Gain-of-Function Mutation in Impeded Bone Development through Increasing Expression in DA2B Mice
- Nephronophthisis-Associated Regulates Cell Cycle Progression, Apoptosis and Epithelial-to-Mesenchymal Transition
- Beclin 1 Is Required for Neuron Viability and Regulates Endosome Pathways via the UVRAG-VPS34 Complex
- The Not5 Subunit of the Ccr4-Not Complex Connects Transcription and Translation
- Abnormal Dosage of Ultraconserved Elements Is Highly Disfavored in Healthy Cells but Not Cancer Cells
- Genome-Wide Distribution of RNA-DNA Hybrids Identifies RNase H Targets in tRNA Genes, Retrotransposons and Mitochondria
- The Chromosomal Association of the Smc5/6 Complex Depends on Cohesion and Predicts the Level of Sister Chromatid Entanglement
- Cell-Autonomous Progeroid Changes in Conditional Mouse Models for Repair Endonuclease XPG Deficiency
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- The Master Activator of IncA/C Conjugative Plasmids Stimulates Genomic Islands and Multidrug Resistance Dissemination
- A Splice Mutation in the Gene Causes High Glycogen Content and Low Meat Quality in Pig Skeletal Muscle
- Keratin 76 Is Required for Tight Junction Function and Maintenance of the Skin Barrier
- A Role for Taiman in Insect Metamorphosis
Přihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



